RNA-Polymerase on:
[Wikipedia]
[Google]
[Amazon]
 In
In
 Control of the process of
Control of the process of
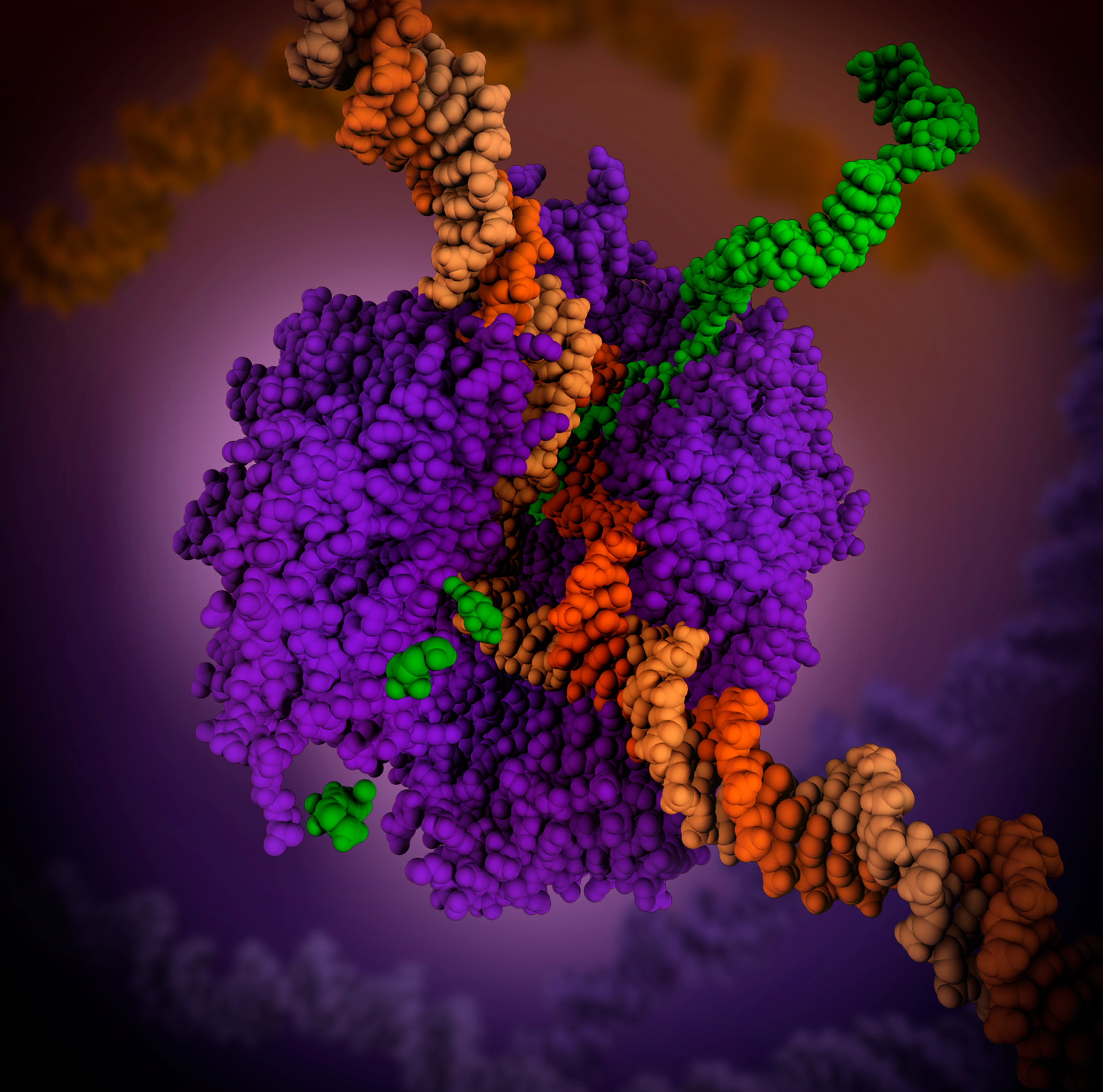
 The 17-bp transcriptional complex has an 8-bp DNA-RNA hybrid, that is, 8 base-pairs involve the RNA transcript bound to the DNA template strand. As transcription progresses, ribonucleotides are added to the 3′ end of the RNA transcript and the RNAP complex moves along the DNA. The characteristic elongation rates in prokaryotes and eukaryotes are about 10–100 nts/sec.
Aspartyl ( asp) residues in the RNAP will hold on to Mg2+ ions, which will, in turn, coordinate the phosphates of the ribonucleotides. The first Mg2+ will hold on to the α-phosphate of the NTP to be added. This allows the nucleophilic attack of the 3′-OH from the RNA transcript, adding another NTP to the chain. The second Mg2+ will hold on to the pyrophosphate of the NTP. The overall reaction equation is:
:(NMP)n + NTP → (NMP)n+1 + PPi
The 17-bp transcriptional complex has an 8-bp DNA-RNA hybrid, that is, 8 base-pairs involve the RNA transcript bound to the DNA template strand. As transcription progresses, ribonucleotides are added to the 3′ end of the RNA transcript and the RNAP complex moves along the DNA. The characteristic elongation rates in prokaryotes and eukaryotes are about 10–100 nts/sec.
Aspartyl ( asp) residues in the RNAP will hold on to Mg2+ ions, which will, in turn, coordinate the phosphates of the ribonucleotides. The first Mg2+ will hold on to the α-phosphate of the NTP to be added. This allows the nucleophilic attack of the 3′-OH from the RNA transcript, adding another NTP to the chain. The second Mg2+ will hold on to the pyrophosphate of the NTP. The overall reaction equation is:
:(NMP)n + NTP → (NMP)n+1 + PPi
DNAi
– DNA Interactive, including information and Flash clips on RNA Polymerase. * *
RNA Polymerase – Synthesis RNA from DNA Template
(
3D macromolecular structures of RNA Polymerase from the EM Data Bank(EMDB)
{{Authority control Gene expression RNA Enzymes EC 2.7.7
 In
In molecular biology
Molecular biology is a branch of biology that seeks to understand the molecule, molecular basis of biological activity in and between Cell (biology), cells, including biomolecule, biomolecular synthesis, modification, mechanisms, and interactio ...
, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different mol ...
that catalyzes the chemical reactions that synthesize RNA
Ribonucleic acid (RNA) is a polymeric molecule that is essential for most biological functions, either by performing the function itself (non-coding RNA) or by forming a template for the production of proteins (messenger RNA). RNA and deoxyrib ...
from a DNA
Deoxyribonucleic acid (; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a double helix. The polymer carries genetic instructions for the development, functioning, growth and reproduction of al ...
template.
Using the enzyme helicase
Helicases are a class of enzymes that are vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic double helix, separating the two hybridized ...
, RNAP locally opens the double-stranded DNA so that one strand of the exposed nucleotide
Nucleotides are Organic compound, organic molecules composed of a nitrogenous base, a pentose sugar and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both o ...
s can be used as a template for the synthesis of RNA, a process called transcription
Transcription refers to the process of converting sounds (voice, music etc.) into letters or musical notes, or producing a copy of something in another medium, including:
Genetics
* Transcription (biology), the copying of DNA into RNA, often th ...
. A transcription factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription (genetics), transcription of genetics, genetic information from DNA to messenger RNA, by binding t ...
and its associated transcription mediator complex
Mediator is a multiprotein complex that functions as a transcriptional coactivator in all eukaryotes. It was discovered in 1990 in the lab of Roger D. Kornberg, recipient of the 2006 Nobel Prize in Chemistry. Mediator complexes interact with tr ...
must be attached to a DNA binding site
DNA binding sites are a type of binding site found in DNA where other molecules may bind. DNA binding sites are distinct from other binding sites in that (1) they are part of a DNA sequence (e.g. a genome) and (2) they are bound by DNA-binding ...
called a promoter region
In genetics, a promoter is a sequence of DNA to which proteins bind to initiate transcription of a single RNA transcript from the DNA downstream of the promoter. The RNA transcript may encode a protein (mRNA), or can have a function in and of it ...
before RNAP can initiate the DNA unwinding at that position. RNAP not only initiates RNA transcription, it also guides the nucleotides into position, facilitates attachment and elongation, has intrinsic proofreading and replacement capabilities, and termination recognition capability. In eukaryote
The eukaryotes ( ) constitute the Domain (biology), domain of Eukaryota or Eukarya, organisms whose Cell (biology), cells have a membrane-bound cell nucleus, nucleus. All animals, plants, Fungus, fungi, seaweeds, and many unicellular organisms ...
s, RNAP can build chains as long as 2.4 million nucleotides.
RNAP produces RNA that, functionally, is either for protein coding, i.e. messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
(mRNA); or non-coding
Non-coding DNA (ncDNA) sequences are components of an organism's DNA that do not encode protein sequences. Some non-coding DNA is transcribed into functional non-coding RNA molecules (e.g. transfer RNA, microRNA, piRNA, ribosomal RNA, and regula ...
(so-called "RNA genes"). Examples of four functional types of RNA genes are:
; Transfer RNA
Transfer ribonucleic acid (tRNA), formerly referred to as soluble ribonucleic acid (sRNA), is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes). In a cell, it provides the physical link between the gene ...
(tRNA): Transfers specific amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although over 500 amino acids exist in nature, by far the most important are the 22 α-amino acids incorporated into proteins. Only these 22 a ...
s to growing polypeptide
Peptides are short chains of amino acids linked by peptide bonds. A polypeptide is a longer, continuous, unbranched peptide chain. Polypeptides that have a molecular mass of 10,000 Da or more are called proteins. Chains of fewer than twenty ...
chains at the ribosomal site of protein synthesis
Protein biosynthesis, or protein synthesis, is a core biological process, occurring inside cells, balancing the loss of cellular proteins (via degradation or export) through the production of new proteins. Proteins perform a number of critica ...
during translation
Translation is the communication of the semantics, meaning of a #Source and target languages, source-language text by means of an Dynamic and formal equivalence, equivalent #Source and target languages, target-language text. The English la ...
;
; Ribosomal RNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal ...
(rRNA): Incorporates into ribosomes;
; Micro RNA
Micro ribonucleic acid (microRNA, miRNA, μRNA) are small, single-stranded, non-coding RNA molecules containing 21–23 nucleotides. Found in plants, animals, and even some viruses, miRNAs are involved in RNA silencing and post-transcri ...
(miRNA): Regulates gene activity; and, RNA silencing
; Catalytic RNA
Ribozymes (ribonucleic acid enzymes) are RNA molecules that have the ability to catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozymes demonst ...
(ribozyme
Ribozymes (ribonucleic acid enzymes) are RNA molecules that have the ability to Catalysis, catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozy ...
): Functions as an enzymatically active RNA molecule.
RNA polymerase is essential to life, and is found in all living organism
An organism is any life, living thing that functions as an individual. Such a definition raises more problems than it solves, not least because the concept of an individual is also difficult. Many criteria, few of them widely accepted, have be ...
s and many virus
A virus is a submicroscopic infectious agent that replicates only inside the living Cell (biology), cells of an organism. Viruses infect all life forms, from animals and plants to microorganisms, including bacteria and archaea. Viruses are ...
es. Depending on the organism, a RNA polymerase can be a protein complex
A protein complex or multiprotein complex is a group of two or more associated polypeptide chains. Protein complexes are distinct from multidomain enzymes, in which multiple active site, catalytic domains are found in a single polypeptide chain.
...
(multi-subunit RNAP) or only consist of one subunit (single-subunit RNAP, ssRNAP), each representing an independent lineage. The former is found in bacteria
Bacteria (; : bacterium) are ubiquitous, mostly free-living organisms often consisting of one Cell (biology), biological cell. They constitute a large domain (biology), domain of Prokaryote, prokaryotic microorganisms. Typically a few micr ...
, archaea
Archaea ( ) is a Domain (biology), domain of organisms. Traditionally, Archaea only included its Prokaryote, prokaryotic members, but this has since been found to be paraphyletic, as eukaryotes are known to have evolved from archaea. Even thou ...
, and eukaryote
The eukaryotes ( ) constitute the Domain (biology), domain of Eukaryota or Eukarya, organisms whose Cell (biology), cells have a membrane-bound cell nucleus, nucleus. All animals, plants, Fungus, fungi, seaweeds, and many unicellular organisms ...
s alike, sharing a similar core structure and mechanism. The latter is found in phage
A bacteriophage (), also known informally as a phage (), is a virus that infects and replicates within bacteria. The term is derived . Bacteriophages are composed of proteins that encapsulate a DNA or RNA genome, and may have structures tha ...
s as well as eukaryotic chloroplast
A chloroplast () is a type of membrane-bound organelle, organelle known as a plastid that conducts photosynthesis mostly in plant cell, plant and algae, algal cells. Chloroplasts have a high concentration of chlorophyll pigments which captur ...
s and mitochondria
A mitochondrion () is an organelle found in the cells of most eukaryotes, such as animals, plants and fungi. Mitochondria have a double membrane structure and use aerobic respiration to generate adenosine triphosphate (ATP), which is us ...
, and is related to modern DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create t ...
s. Eukaryotic and archaeal RNAPs have more subunits than bacterial ones do, and are controlled differently.
Bacteria and archaea only have one RNA polymerase. Eukaryotes have multiple types of nuclear RNAP, each responsible for synthesis of a distinct subset of RNA:
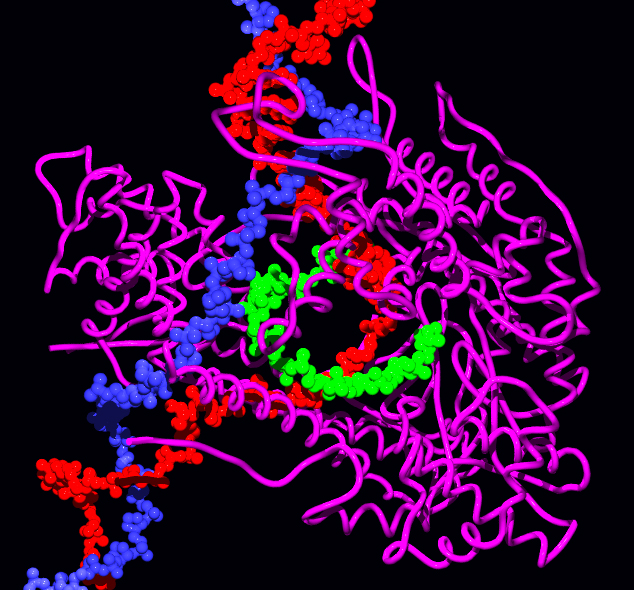
Structure
The 2006Nobel Prize in Chemistry
The Nobel Prize in Chemistry () is awarded annually by the Royal Swedish Academy of Sciences to scientists in the various fields of chemistry. It is one of the five Nobel Prizes established by the will of Alfred Nobel in 1895, awarded for outst ...
was awarded to Roger D. Kornberg for creating detailed molecular images of RNA polymerase during various stages of the transcription process.
In most prokaryote
A prokaryote (; less commonly spelled procaryote) is a unicellular organism, single-celled organism whose cell (biology), cell lacks a cell nucleus, nucleus and other membrane-bound organelles. The word ''prokaryote'' comes from the Ancient Gree ...
s, a single RNA polymerase species transcribes all types of RNA. RNA polymerase "core" from E. coli
''Escherichia coli'' ( )Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. is a gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escherichia'' that is commonly foun ...
consists of five subunits: two alpha (α) subunits of 36 kDa
The dalton or unified atomic mass unit (symbols: Da or u, respectively) is a unit of mass defined as of the mass of an unbound neutral atom of carbon-12 in its nuclear and electronic ground state and at rest. It is a non-SI unit accepted f ...
, a beta (β) subunit of 150 kDa, a beta prime subunit (β′) of 155 kDa, and a small omega (ω) subunit. A sigma (σ) factor binds to the core, forming the holoenzyme. After transcription starts, the factor can unbind and let the core enzyme proceed with its work. The core RNA polymerase complex forms a "crab claw" or "clamp-jaw" structure with an internal channel running along the full length. Eukaryotic and archaeal RNA polymerases have a similar core structure and work in a similar manner, although they have many extra subunits.
All RNAPs contain metal cofactors, in particular zinc
Zinc is a chemical element; it has symbol Zn and atomic number 30. It is a slightly brittle metal at room temperature and has a shiny-greyish appearance when oxidation is removed. It is the first element in group 12 (IIB) of the periodic tabl ...
and magnesium
Magnesium is a chemical element; it has Symbol (chemistry), symbol Mg and atomic number 12. It is a shiny gray metal having a low density, low melting point and high chemical reactivity. Like the other alkaline earth metals (group 2 ...
cations which aid in the transcription process.
Function
 Control of the process of
Control of the process of gene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protei ...
transcription affects patterns of gene expression
Gene expression is the process (including its Regulation of gene expression, regulation) by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, proteins or non-coding RNA, ...
and, thereby, allows a cell
Cell most often refers to:
* Cell (biology), the functional basic unit of life
* Cellphone, a phone connected to a cellular network
* Clandestine cell, a penetration-resistant form of a secret or outlawed organization
* Electrochemical cell, a de ...
to adapt to a changing environment, perform specialized roles within an organism, and maintain basic metabolic processes necessary for survival. Therefore, it is hardly surprising that the activity of RNAP is long, complex, and highly regulated. In ''Escherichia coli'' bacteria, more than 100 transcription factor
In molecular biology, a transcription factor (TF) (or sequence-specific DNA-binding factor) is a protein that controls the rate of transcription (genetics), transcription of genetics, genetic information from DNA to messenger RNA, by binding t ...
s have been identified, which modify the activity of RNAP.
RNAP can initiate transcription at specific DNA sequences known as promoters. It then produces an RNA chain, which is complementary
Complement may refer to:
The arts
* Complement (music), an interval that, when added to another, spans an octave
** Aggregate complementation, the separation of pitch-class collections into complementary sets
* Complementary color, in the visu ...
to the template DNA strand. The process of adding nucleotide
Nucleotides are Organic compound, organic molecules composed of a nitrogenous base, a pentose sugar and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both o ...
s to the RNA strand is known as elongation; in eukaryotes, RNAP can build chains as long as 2.4 million nucleotides
Nucleotides are Organic compound, organic molecules composed of a nitrogenous base, a pentose sugar and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both o ...
(the full length of the dystrophin
Dystrophin is a rod-shaped cytoplasmic protein, and a vital part of a protein complex that connects the cytoskeleton of a muscle fiber to the surrounding extracellular matrix through the cell membrane. This complex is variously known as the costa ...
gene). RNAP will preferentially release its RNA transcript at specific DNA sequences encoded at the end of genes, which are known as terminators.
Products of RNAP include:
* Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein.
mRNA is created during the ...
(mRNA)—template for the synthesis of proteins by ribosome
Ribosomes () are molecular machine, macromolecular machines, found within all cell (biology), cells, that perform Translation (biology), biological protein synthesis (messenger RNA translation). Ribosomes link amino acids together in the order s ...
s.
* Non-coding RNA
A non-coding RNA (ncRNA) is a functional RNA molecule that is not Translation (genetics), translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally imp ...
or "RNA genes"—a broad class of genes that encode RNA that is not translated into protein. The most prominent examples of RNA genes are transfer RNA
Transfer ribonucleic acid (tRNA), formerly referred to as soluble ribonucleic acid (sRNA), is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes). In a cell, it provides the physical link between the gene ...
(tRNA) and ribosomal RNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal ...
(rRNA), both of which are involved in the process of translation
Translation is the communication of the semantics, meaning of a #Source and target languages, source-language text by means of an Dynamic and formal equivalence, equivalent #Source and target languages, target-language text. The English la ...
. However, since the late 1990s, many new RNA genes have been found, and thus RNA genes may play a much more significant role than previously thought.
** Transfer RNA
Transfer ribonucleic acid (tRNA), formerly referred to as soluble ribonucleic acid (sRNA), is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes). In a cell, it provides the physical link between the gene ...
(tRNA)—transfers specific amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although over 500 amino acids exist in nature, by far the most important are the 22 α-amino acids incorporated into proteins. Only these 22 a ...
s to growing polypeptide
Peptides are short chains of amino acids linked by peptide bonds. A polypeptide is a longer, continuous, unbranched peptide chain. Polypeptides that have a molecular mass of 10,000 Da or more are called proteins. Chains of fewer than twenty ...
chains at the ribosomal site of protein synthesis during translation
Translation is the communication of the semantics, meaning of a #Source and target languages, source-language text by means of an Dynamic and formal equivalence, equivalent #Source and target languages, target-language text. The English la ...
** Ribosomal RNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal ...
(rRNA)—a component of ribosomes
** Micro RNA
Micro ribonucleic acid (microRNA, miRNA, μRNA) are small, single-stranded, non-coding RNA molecules containing 21–23 nucleotides. Found in plants, animals, and even some viruses, miRNAs are involved in RNA silencing and post-transcri ...
—regulates gene activity
** Catalytic RNA (Ribozyme
Ribozymes (ribonucleic acid enzymes) are RNA molecules that have the ability to Catalysis, catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozy ...
)—enzymatically active RNA molecules
RNAP accomplishes ''de novo'' synthesis. It is able to do this because specific interactions with the initiating nucleotide hold RNAP rigidly in place, facilitating chemical attack on the incoming nucleotide. Such specific interactions explain why RNAP prefers to start transcripts with ATP (followed by GTP, UTP, and then CTP). In contrast to DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create t ...
, RNAP includes helicase
Helicases are a class of enzymes that are vital to all organisms. Their main function is to unpack an organism's genetic material. Helicases are motor proteins that move directionally along a nucleic double helix, separating the two hybridized ...
activity, therefore no separate enzyme is needed to unwind DNA.
Action
Initiation
RNA polymerase binding in bacteria involves thesigma factor
A sigma factor (σ factor or specificity factor) is a protein needed for initiation of Transcription (biology), transcription in bacteria. It is a bacterial transcription initiation factor that enables specific binding of RNA polymerase (RNAP) to g ...
recognizing the core promoter region containing the −35 and −10 elements (located ''before'' the beginning of sequence to be transcribed) and also, at some promoters, the α subunit C-terminal domain recognizing promoter upstream elements. There are multiple interchangeable sigma factors, each of which recognizes a distinct set of promoters. For example, in ''E. coli'', σ70 is expressed under normal conditions and recognizes promoters for genes required under normal conditions ("housekeeping gene
In molecular biology, housekeeping genes are typically constitutive genes that are required for the maintenance of basic cellular function, and are expressed in all cells of an organism under normal and patho-physiological conditions. Although ...
s"), while σ32 recognizes promoters for genes required at high temperatures (" heat-shock genes"). In archaea and eukaryotes, the functions of the bacterial general transcription factor sigma are performed by multiple general transcription factor
General transcription factors (GTFs), also known as basal transcriptional factors, are a class of protein transcription factors that bind to specific sites ( promoter) on DNA to activate transcription of genetic information from DNA to messenger ...
s that work together. The RNA polymerase-promoter closed complex is usually referred to as the "transcription preinitiation complex
The preinitiation complex (abbreviated PIC) is a complex of approximately 100 proteins that is necessary for the transcription (genetics), transcription of protein-coding genes in eukaryotes and archaea. The preinitiation complex positions RNA po ...
."
After binding to the DNA, the RNA polymerase switches from a closed complex to an open complex. This change involves the separation of the DNA strands to form an unwound section of DNA of approximately 13 bp, referred to as the "transcription bubble
A transcription bubble is a molecular structure formed during the initialization of DNA transcription, when a limited portion of the DNA double helix is unwound, providing enough space for RNA polymerase (RNAP) to bind to the template strand and ...
". Supercoiling plays an important part in polymerase activity because of the unwinding and rewinding of DNA. Because regions of DNA in front of RNAP are unwound, there are compensatory positive supercoils. Regions behind RNAP are rewound and negative supercoils are present.
Promoter escape
RNA polymerase then starts to synthesize the initial DNA-RNA heteroduplex, with ribonucleotides base-paired to the template DNA strand according to Watson-Crick base-pairing interactions. As noted above, RNA polymerase makes contacts with the promoter region. However these stabilizing contacts inhibit the enzyme's ability to access DNA further downstream and thus the synthesis of the full-length product. In order to continue RNA synthesis, RNA polymerase must escape the promoter. It must maintain promoter contacts while unwinding more downstream DNA for synthesis, "scrunching" more downstream DNA into the initiation complex. During the promoter escape transition, RNA polymerase is considered a "stressed intermediate." Thermodynamically the stress accumulates from the DNA-unwinding and DNA-compaction activities. Once the DNA-RNA heteroduplex is long enough (~10 bp), RNA polymerase releases its upstream contacts and effectively achieves the promoter escape transition into the elongation phase. The heteroduplex at the active center stabilizes the elongation complex. However, promoter escape is not the only outcome. RNA polymerase can also relieve the stress by releasing its downstream contacts, arresting transcription. The paused transcribing complex has two options: (1) release the nascent transcript and begin anew at the promoter or (2) reestablish a new 3′-OH on the nascent transcript at the active site via RNA polymerase's catalytic activity and recommence DNA scrunching to achieve promoter escape.Abortive initiation Abortive initiation, also known as abortive transcription, is an early process of genetic transcription in which RNA polymerase binds to a DNA promoter and enters into cycles of synthesis of short mRNA transcripts which are released before the tra ...
, the unproductive cycling of RNA polymerase before the promoter escape transition, results in short RNA fragments of around 9 bp in a process known as abortive transcription. The extent of abortive initiation depends on the presence of transcription factors and the strength of the promoter contacts.
Elongation
 The 17-bp transcriptional complex has an 8-bp DNA-RNA hybrid, that is, 8 base-pairs involve the RNA transcript bound to the DNA template strand. As transcription progresses, ribonucleotides are added to the 3′ end of the RNA transcript and the RNAP complex moves along the DNA. The characteristic elongation rates in prokaryotes and eukaryotes are about 10–100 nts/sec.
Aspartyl ( asp) residues in the RNAP will hold on to Mg2+ ions, which will, in turn, coordinate the phosphates of the ribonucleotides. The first Mg2+ will hold on to the α-phosphate of the NTP to be added. This allows the nucleophilic attack of the 3′-OH from the RNA transcript, adding another NTP to the chain. The second Mg2+ will hold on to the pyrophosphate of the NTP. The overall reaction equation is:
:(NMP)n + NTP → (NMP)n+1 + PPi
The 17-bp transcriptional complex has an 8-bp DNA-RNA hybrid, that is, 8 base-pairs involve the RNA transcript bound to the DNA template strand. As transcription progresses, ribonucleotides are added to the 3′ end of the RNA transcript and the RNAP complex moves along the DNA. The characteristic elongation rates in prokaryotes and eukaryotes are about 10–100 nts/sec.
Aspartyl ( asp) residues in the RNAP will hold on to Mg2+ ions, which will, in turn, coordinate the phosphates of the ribonucleotides. The first Mg2+ will hold on to the α-phosphate of the NTP to be added. This allows the nucleophilic attack of the 3′-OH from the RNA transcript, adding another NTP to the chain. The second Mg2+ will hold on to the pyrophosphate of the NTP. The overall reaction equation is:
:(NMP)n + NTP → (NMP)n+1 + PPi
Fidelity
Unlike the proofreading mechanisms ofDNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create t ...
those of RNAP have only recently been investigated. Proofreading begins with separation of the mis-incorporated nucleotide from the DNA template. This pauses transcription. The polymerase then backtracks by one position and cleaves the dinucleotide that contains the mismatched nucleotide. In the RNA polymerase this occurs at the same active site used for polymerization and is therefore markedly different from the DNA polymerase where proofreading occurs at a distinct nuclease active site.
The overall error rate is around 10−4 to 10−6.
Termination
In bacteria, termination of RNA transcription can be rho-dependent or rho-independent. The former relies on therho factor
A ρ factor (Rho factor) is a bacterial protein involved in the termination of transcription.
* Rho factor binds to the transcription terminator pause site, an exposed region of single stranded RNA (a stretch of 72 nucleotides) after the open ...
, which destabilizes the DNA-RNA heteroduplex and causes RNA release. The latter, also known as intrinsic termination
Intrinsic, or rho-independent termination, is a process to signal the end of transcription and release the newly constructed RNA molecule. In bacteria such as ''E. coli'', transcription is terminated either by a rho-dependent process or rho-indep ...
, relies on a palindromic region of DNA. Transcribing the region causes the formation of a "hairpin" structure from the RNA transcription looping and binding upon itself. This hairpin structure is often rich in G-C base-pairs, making it more stable than the DNA-RNA hybrid itself. As a result, the 8 bp DNA-RNA hybrid in the transcription complex shifts to a 4 bp hybrid. These last 4 base pairs are weak A-U base pairs, and the entire RNA transcript will fall off the DNA.
Transcription termination in eukaryotes is less well understood than in bacteria, but involves cleavage of the new transcript followed by template-independent addition of adenines at its new 3′ end, in a process called polyadenylation
Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In euka ...
.
Other organisms
Given that DNA and RNA polymerases both carry out template-dependent nucleotide polymerization, it might be expected that the two types of enzymes would be structurally related. However, x-ray crystallographic studies of both types of enzymes reveal that, other than containing a critical Mg2+ ion at the catalytic site, they are virtually unrelated to each other; indeed template-dependent nucleotide polymerizing enzymes seem to have arisen independently twice during the early evolution of cells. One lineage led to the modern DNA polymerases and reverse transcriptases, as well as to a few single-subunit RNA polymerases (ssRNAP) from phages and organelles. The other multi-subunit RNAP lineage formed all of the modern cellular RNA polymerases.Bacteria
Inbacteria
Bacteria (; : bacterium) are ubiquitous, mostly free-living organisms often consisting of one Cell (biology), biological cell. They constitute a large domain (biology), domain of Prokaryote, prokaryotic microorganisms. Typically a few micr ...
, the same enzyme catalyzes the synthesis of mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
and non-coding RNA (ncRNA).
RNAP is a large molecule. The core enzyme has five subunits (~400 kDa
The dalton or unified atomic mass unit (symbols: Da or u, respectively) is a unit of mass defined as of the mass of an unbound neutral atom of carbon-12 in its nuclear and electronic ground state and at rest. It is a non-SI unit accepted f ...
):
; β′: The β′ subunit is the largest subunit, and is encoded by the rpoC gene. The β′ subunit contains part of the active center responsible for RNA synthesis and contains some of the determinants for non-sequence-specific interactions with DNA and nascent RNA. It is split into two subunits in Cyanobacteria and chloroplasts.
; β: The β subunit is the second-largest subunit, and is encoded by the rpoB gene. The β subunit contains the rest of the active center responsible for RNA synthesis and contains the rest of the determinants for non-sequence-specific interactions with DNA and nascent RNA.
; α (αI and αII): Two copies of the α subunit, being the third-largest subunit, are present in a molecule of RNAP: αI and αII (one and two). Each α subunit contains two domains: αNTD (N-terminal domain) and αCTD (C-terminal domain). αNTD contains determinants for assembly of RNAP. αCTD (C-terminal domain) contains determinants for interaction with promoter DNA, making non-sequence-non-specific interactions at most promoters and sequence-specific interactions at upstream-element-containing promoters, and contains determinants for interactions with regulatory factors.
; ω: The ω subunit is the smallest subunit. The ω subunit facilitates assembly of RNAP and stabilizes assembled RNAP.
In order to bind promoters, RNAP core associates with the transcription initiation factor sigma
Sigma ( ; uppercase Σ, lowercase σ, lowercase in word-final position ς; ) is the eighteenth letter of the Greek alphabet. In the system of Greek numerals, it has a value of 200. In general mathematics, uppercase Σ is used as an operator ...
(σ) to form RNA polymerase holoenzyme. Sigma reduces the affinity of RNAP for nonspecific DNA while increasing specificity for promoters, allowing transcription to initiate at correct sites. The complete holoenzyme therefore has 6 subunits: β′βαI and αIIωσ (~450 kDa).
Eukaryotes
Eukaryote
The eukaryotes ( ) constitute the Domain (biology), domain of Eukaryota or Eukarya, organisms whose Cell (biology), cells have a membrane-bound cell nucleus, nucleus. All animals, plants, Fungus, fungi, seaweeds, and many unicellular organisms ...
s have multiple types of nuclear RNAP, each responsible for synthesis of a distinct subset of RNA. All are structurally and mechanistically related to each other and to bacterial RNAP:
Eukaryotic chloroplast
A chloroplast () is a type of membrane-bound organelle, organelle known as a plastid that conducts photosynthesis mostly in plant cell, plant and algae, algal cells. Chloroplasts have a high concentration of chlorophyll pigments which captur ...
s contain a multi-subunit RNAP ("PEP, plastid-encoded polymerase"). Due to its bacterial origin, the organization of PEP resembles that of current bacterial RNA polymerases: It is encoded by the RPOA, RPOB, RPOC1 and RPOC2 genes on the plastome, which as proteins form the core subunits of PEP, respectively named α, β, β′ and β″. Similar to the RNA polymerase in ''E. coli'', PEP requires the presence of sigma
Sigma ( ; uppercase Σ, lowercase σ, lowercase in word-final position ς; ) is the eighteenth letter of the Greek alphabet. In the system of Greek numerals, it has a value of 200. In general mathematics, uppercase Σ is used as an operator ...
(σ) factors for the recognition of its promoters, containing the -10 and -35 motifs. Despite the many commonalities between plant organellar and bacterial RNA polymerases and their structure, PEP additionally requires the association of a number of nuclear encoded proteins, termed PAPs (PEP-associated proteins), which form essential components that are closely associated with the PEP complex in plants. Initially, a group consisting of 10 PAPs was identified through biochemical methods, which was later extended to 12 PAPs.
Chloroplast also contain a second, structurally and mechanistically unrelated, single-subunit RNAP ("nucleus-encoded polymerase, NEP"). Eukaryotic mitochondria
A mitochondrion () is an organelle found in the cells of most eukaryotes, such as animals, plants and fungi. Mitochondria have a double membrane structure and use aerobic respiration to generate adenosine triphosphate (ATP), which is us ...
use POLRMT
DNA-directed RNA polymerase, mitochondrial is an enzyme that in humans is encoded by the ''POLRMT'' gene.
Function
This gene encodes a mitochondrial DNA-directed RNA polymerase. The gene product is responsible for mitochondrial gene expressio ...
(human), a nucleus-encoded single-subunit RNAP. Such phage-like polymerases are referred to as RpoT in plants.
Archaea
Archaea
Archaea ( ) is a Domain (biology), domain of organisms. Traditionally, Archaea only included its Prokaryote, prokaryotic members, but this has since been found to be paraphyletic, as eukaryotes are known to have evolved from archaea. Even thou ...
have a single type of RNAP, responsible for the synthesis of all RNA. Archaeal RNAP is structurally and mechanistically similar to bacterial RNAP and eukaryotic nuclear RNAP I-V, and is especially closely structurally and mechanistically related to eukaryotic nuclear RNAP II.
The history of the discovery of the archaeal RNA polymerase is quite recent. The first analysis of the RNAP of an archaeon was performed in 1971, when the RNAP from the extreme halophile
A halophile (from the Greek word for 'salt-loving') is an extremophile that thrives in high salt
In common usage, salt is a mineral composed primarily of sodium chloride (NaCl). When used in food, especially in granulated form, it is more ...
''Halobacterium cutirubrum
''Halobacterium salinarum'', formerly known as ''Halobacterium cutirubrum'' or ''Halobacterium halobium'', is an extremely halophilic marine obligate aerobic archaeon. Despite its name, this is not a bacterium, but a member of the domain Archaea. ...
'' was isolated and purified. Crystal structures of RNAPs from ''Sulfolobus solfataricus
''Saccharolobus solfataricus'' is a species of thermophilic archaeon. It was transferred from the genus ''Sulfolobus'' to the new genus ''Saccharolobus'' with the description of ''Saccharolobus caldissimus'' in 2018.
It was first discovered an ...
'' and ''Sulfolobus shibatae
''Saccharolobus shibatae'' is an archaeal species belongs to the phylum Thermoproteota. ''Saccharolobus shibatae'' was described for the first time as ''Sulfolobus shibatae'' in 1990, after being isolated from geothermal pools in Beppu, Ōita, B ...
'' set the total number of identified archaeal subunits at thirteen.
Archaea has the subunit corresponding to Eukaryotic Rpb1 split into two. There is no homolog to eukaryotic Rpb9 (POLR2I
DNA-directed RNA polymerase II subunit RPB9 is an enzyme that in humans is encoded by the ''POLR2I'' gene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucle ...
) in the ''S. shibatae'' complex, although TFS (TFIIS homolog) has been proposed as one based on similarity. There is an additional subunit dubbed Rpo13; together with Rpo5 it occupies a space filled by an insertion found in bacterial β′ subunits (1,377–1,420 in ''Taq''). An earlier, lower-resolution study on ''S. solfataricus'' structure did not find Rpo13 and only assigned the space to Rpo5/Rpb5. Rpo3 is notable in that it's an iron–sulfur protein
Iron–sulfur proteins are proteins characterized by the presence of iron–sulfur clusters containing sulfide-linked di-, tri-, and tetrairon centers in variable oxidation states. Iron–sulfur clusters are found in a variety of metalloproteins ...
. RNAP I/III subunit AC40 found in some eukaryotes share similar sequences, but does not bind iron. This domain, in either case, serves a structural function.
Archaeal RNAP subunit previously used an "RpoX" nomenclature where each subunit is assigned a letter in a way unrelated to any other systems. In 2009, a new nomenclature based on Eukaryotic Pol II subunit "Rpb" numbering was proposed.
Viruses

Orthopoxvirus
''Orthopoxvirus'' is a genus of viruses in the family ''Poxviridae'' and subfamily ''Chordopoxvirinae''. Vertebrates, including mammals and humans, and arthropods serve as natural hosts. There are 12 species in this genus. Diseases associated wi ...
es and some other nucleocytoplasmic large DNA viruses
''Nucleocytoviricota'' is a phylum of viruses. Members of the phylum are also known as the nucleocytoplasmic large DNA viruses (NCLDV), which serves as the basis of the name of the phylum with the suffix - for virus phylum. These viruses are refe ...
synthesize RNA using a virally encoded multi-subunit RNAP. They are most similar to eukaryotic RNAPs, with some subunits minified or removed. Exactly which RNAP they are most similar to is a topic of debate. Most other viruses that synthesize RNA use unrelated mechanics.
Many viruses use a single-subunit DNA-dependent RNAP (ssRNAP) that is structurally and mechanistically related to the single-subunit RNAP of eukaryotic chloroplasts (RpoT) and mitochondria (POLRMT
DNA-directed RNA polymerase, mitochondrial is an enzyme that in humans is encoded by the ''POLRMT'' gene.
Function
This gene encodes a mitochondrial DNA-directed RNA polymerase. The gene product is responsible for mitochondrial gene expressio ...
) and, more distantly, to DNA polymerase
A DNA polymerase is a member of a family of enzymes that catalyze the synthesis of DNA molecules from nucleoside triphosphates, the molecular precursors of DNA. These enzymes are essential for DNA replication and usually work in groups to create t ...
s and reverse transcriptase
A reverse transcriptase (RT) is an enzyme used to convert RNA genome to DNA, a process termed reverse transcription. Reverse transcriptases are used by viruses such as HIV and hepatitis B to replicate their genomes, by retrotransposon mobi ...
s. Perhaps the most widely studied such single-subunit RNAP is bacteriophage
A bacteriophage (), also known informally as a phage (), is a virus that infects and replicates within bacteria. The term is derived . Bacteriophages are composed of proteins that Capsid, encapsulate a DNA or RNA genome, and may have structu ...
T7 RNA polymerase
T7 RNA Polymerase is an RNA polymerase from the T7 bacteriophage that catalyzes the formation of RNA from DNA in the 5'→ 3' direction.
Activity
T7 polymerase is extremely promoter-specific and transcribes only DNA downstream of a T7 promo ...
. ssRNAPs cannot proofread.
B. subtilis
''Bacillus subtilis'' (), known also as the hay bacillus or grass bacillus, is a gram-positive, catalase-positive bacterium, found in soil and the gastrointestinal tract of ruminants, humans and marine sponges. As a member of the genus ''Bacill ...
prophage
A prophage is a bacteriophage (often shortened to "phage") genome that is integrated into the circular bacterial chromosome or exists as an extrachromosomal plasmid within the bacterial cell (biology), cell. Integration of prophages into the bacte ...
SPβ uses YonO, a homolog of the β+β′ subunits of msRNAPs to form a monomeric (both barrels on the same chain) RNAP distinct from the usual "right hand" ssRNAP. It probably diverged very long ago from the canonical five-unit msRNAP, before the time of the last universal common ancestor
The last universal common ancestor (LUCA) is the hypothesized common ancestral cell from which the three domains of life, the Bacteria, the Archaea, and the Eukarya originated. The cell had a lipid bilayer; it possessed the genetic code a ...
.
Other viruses use an RNA-dependent RNAP (an RNAP that employs RNA as a template instead of DNA). This occurs in negative strand RNA viruses
Negative-strand RNA viruses (−ssRNA viruses) are a group of related viruses that have negative-sense, single-stranded genomes made of ribonucleic acid (RNA). They have genomes that act as complementary strands from which messenger RNA (mRNA) i ...
and dsRNA viruses, both of which exist for a portion of their life cycle as double-stranded RNA. However, some positive strand RNA viruses, such as polio
Poliomyelitis ( ), commonly shortened to polio, is an infectious disease caused by the poliovirus. Approximately 75% of cases are asymptomatic; mild symptoms which can occur include sore throat and fever; in a proportion of cases more severe ...
virus, also contain RNA-dependent RNAP.
History
RNAP was discovered independently by Sam Weiss, Audrey Stevens, andJerard Hurwitz
Jerard Hurwitz (November 20, 1928 – January 24, 2019) was an American biochemist who co-discovered RNA polymerase in 1960 along with Sam Weiss, Audrey Stevens, and James Bonner. He most recently worked at the Sloan-Kettering Institute in New Y ...
in 1960. By this time, one half of the 1959 Nobel Prize
The Nobel Prizes ( ; ; ) are awards administered by the Nobel Foundation and granted in accordance with the principle of "for the greatest benefit to humankind". The prizes were first awarded in 1901, marking the fifth anniversary of Alfred N ...
in Medicine had been awarded to Severo Ochoa
Severo Ochoa de Albornoz (; 24 September 1905 – 1 November 1993) was a Spanish physician and biochemist, and winner of the 1959 Nobel Prize in Physiology or Medicine together with Arthur Kornberg for their discovery of "the mechanisms in the ...
for the discovery of what was believed to be RNAP, but instead turned out to be polynucleotide phosphorylase
Polynucleotide Phosphorylase (PNPase) is a bifunctional enzyme with a phosphorolytic 3' to 5' exoribonuclease activity and a 3'-terminal oligonucleotide polymerase activity. That is, it dismantles the RNA chain starting at the 3' end and working ...
.
Purification
RNA polymerase can be isolated in the following ways: * By a phosphocellulose column. * By glycerol gradient centrifugation. * By a DNA column. * By anion chromatography
Ion chromatography (or ion-exchange chromatography) is a form of chromatography that separates ions and ionizable polar molecules based on their affinity to the ion exchanger. It works on almost any kind of Charge (chemistry), charged molecule ...
column.
And also combinations of the above techniques.
See also
* Alpha-amanitin *Primase
DNA primase is an enzyme involved in the replication of DNA and is a type of RNA polymerase. Primase catalyzes the synthesis of a short RNA (or DNA in some
living organisms) segment called a primer complementary to a ssDNA (single-stranded ...
References
External links
DNAi
– DNA Interactive, including information and Flash clips on RNA Polymerase. * *
RNA Polymerase – Synthesis RNA from DNA Template
(
Wayback Machine
The Wayback Machine is a digital archive of the World Wide Web founded by Internet Archive, an American nonprofit organization based in San Francisco, California. Launched for public access in 2001, the service allows users to go "back in ...
copy)
3D macromolecular structures of RNA Polymerase from the EM Data Bank(EMDB)
{{Authority control Gene expression RNA Enzymes EC 2.7.7