|
Nibrin
Nibrin, also known as NBN or NBS1, is a protein which in humans is encoded by the ''NBN'' gene. Function Nibrin is a protein associated with the repair of double strand breaks (DSBs) which pose serious damage to a genome. It is a 754 amino acid protein identified as a member of the NBS1/hMre11/RAD50(N/M/R, more commonly referred to as MRN) double strand DNA break repair complex. This complex recognizes DNA damage and rapidly relocates to DSB sites and forms nuclear foci. It also has a role in regulation of N/M/R (MRN) protein complex activity which includes end-processing of both physiological and mutagenic DNA double strand breaks (DSBs). Cellular response to DSBs Cellular response is performed by damage sensors, effectors of lesion repair and signal transduction. The central role is carried out by ataxia telangiectasia mutated (ATM) by activating the DSB signaling cascade, phosphorylating downstream substrates such as histone H2AX and NBS1. NBS1 relocates to DSB sit ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Rad50
DNA repair protein RAD50, also known as RAD50, is a protein that in humans is encoded by the ''RAD50'' gene. Function The protein encoded by this gene is highly similar to ''Saccharomyces cerevisiae'' Rad50, a protein involved in DNA double-strand break repair. This protein forms a complex with MRE11 and NBS1 (also known as Xrs2 in yeast). This MRN complex (MRX complex in yeast) binds to broken DNA ends and displays numerous enzymatic activities that are required for double-strand break repair by nonhomologous end-joining or homologous recombination. Gene knockout studies of the mouse homolog of Rad50 suggest it is essential for cell growth and viability. Two alternatively spliced transcript variants of Rad50, which encode distinct proteins, have been reported. Structure Rad50 is a member of the structural maintenance of chromosomes (SMC) family of proteins. Like other SMC proteins, Rad50 contains a long internal coiled-coil domain that folds back on itself, bringing t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
H2AX
H2A histone family member X (usually abbreviated as H2AX) is a type of histone protein from the H2A family encoded by the ''H2AFX'' gene. An important phosphorylated form is γH2AX (S139), which forms when double-strand breaks appear. In humans and other eukaryotes, the DNA is wrapped around histone octamers, consisting of core histones H2A, H2B, H3 and H4, to form chromatin. H2AX contributes to nucleosome-formation, chromatin-remodeling and DNA repair, and is also used ''in vitro'' as an assay for double-strand breaks in dsDNA. Formation of γH2AX H2AX becomes phosphorylated on serine 139, then called γH2AX, as a reaction on DNA double-strand breaks (DSB). The kinases of the PI3-family (Ataxia telangiectasia mutated, ATR and DNA-PKcs) are responsible for this phosphorylation, especially ATM. The modification can happen accidentally during replication fork collapse or in the response to ionizing radiation but also during controlled physiological processes such as V(D)J re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRE11A
Double-strand break repair protein MRE11 (Meiotic recombination 11) is an enzyme that in humans is encoded by the ''MRE11'' gene. The gene has been designated ''MRE11A'' to distinguish it from the pseudogene ''MRE11B'' that is nowadays named ''MRE11P1''. Function This gene encodes a nuclear protein involved in homologous recombination, telomere length maintenance, and DNA double-strand break repair. By itself, the protein has 3' to 5' exonuclease activity and endonuclease activity. The protein forms a complex with the RAD50 homolog; this complex is required for nonhomologous joining of DNA ends and possesses increased single-stranded DNA endonuclease and 3' to 5' exonuclease activities. In conjunction with a DNA ligase, this protein promotes the joining of noncomplementary ends in vitro using short homologies near the ends of the DNA fragments. This gene has a pseudogene on chromosome 3. Alternative splicing of this gene results in two transcript variants encoding different is ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BRCT Domain
BRCA1 C Terminus (BRCT) domain is a family of evolutionarily related proteins. It is named after the C-terminal domain of BRCA1, a DNA-repair protein that serves as a marker of breast cancer susceptibility. The BRCT domain is found predominantly in proteins involved in cell cycle checkpoint functions responsive to DNA damage, for example as found in the breast cancer DNA-repair protein BRCA1. The domain is an approximately 100 amino acid tandem repeat, which appears to act as a phospho-protein binding domain. Examples Human proteins containing this domain include: * BARD1; BRCA1 * CTDP1; TDT or DNTT * ECT2 * LIG4 * MCPH1; MDC1 * NBN * PARP1; PARP4; PAXIP1; PES1 * REV1; RFC1; TOPBP1 DNA topoisomerase 2-binding protein 1 (TOPBP1) is a scaffold protein that in humans is encoded by the ''TOPBP1'' gene. TOPBP1 was first identified as a protein binding partner of DNA TOP2B, topoisomerase-IIβ by a Two-hybrid screening, yeast 2- ...; TP53BP1; XRCC1 References ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Microhomology-mediated End Joining
Microhomology-mediated end joining (MMEJ), also known as alternative nonhomologous end-joining (Alt-NHEJ) is one of the pathways for repairing double-strand breaks in DNA. As reviewed by McVey and Lee, the foremost distinguishing property of MMEJ is the use of microhomologous sequences during the alignment of broken ends before joining, thereby resulting in deletions flanking the original break. MMEJ is frequently associated with chromosome abnormalities such as deletions, translocations, inversions and other complex rearrangements. There are multiple pathways for repairing double strand breaks, mainly non-homologous end joining (NHEJ), homologous recombination (HR), and MMEJ. NHEJ directly joins both ends of the double strand break and is relatively accurate, although small (usually less than a few nucleotides) insertions or deletions sometimes occur. HR is highly accurate and uses the sister chromatid as a template for accurate repair of the DSB. MMEJ is distinguished from thes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ataxia Telangiectasia Mutated
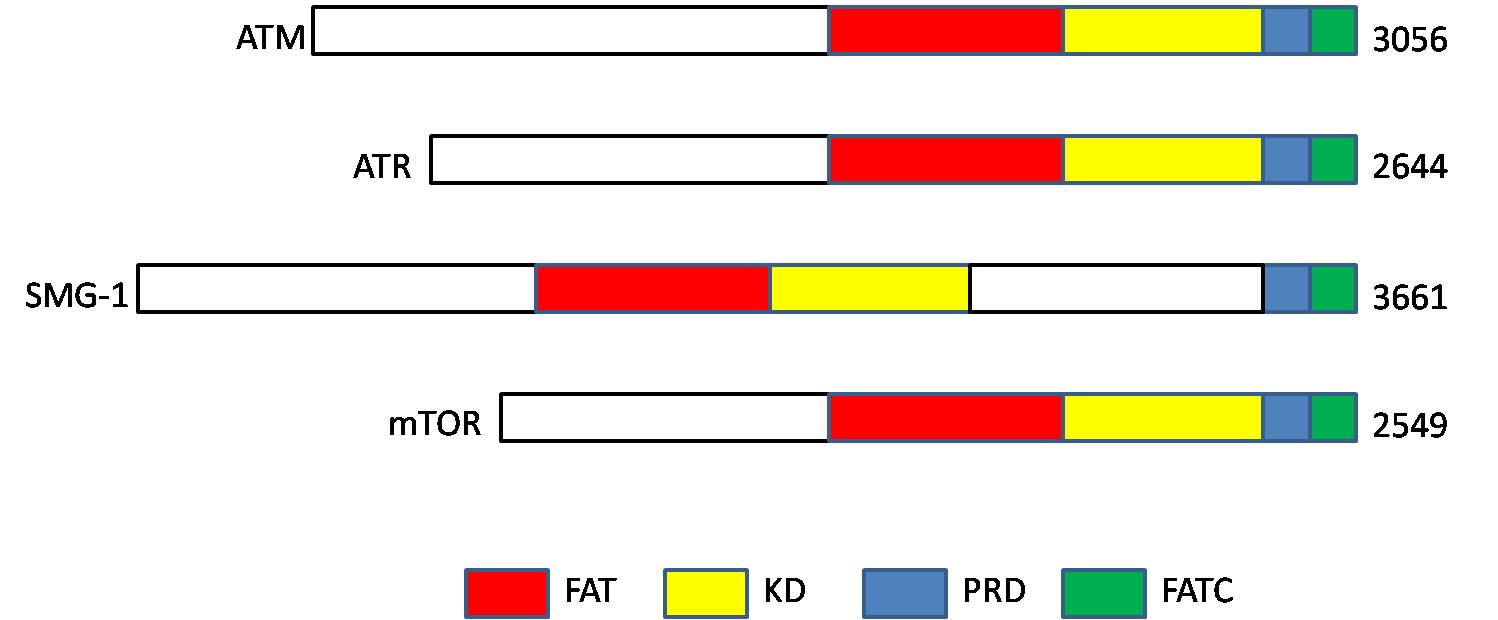
ATM serine/threonine kinase or Ataxia-telangiectasia mutated, symbol ATM, is a serine/threonine protein kinase that is recruited and activated by DNA repair#Double-strand breaks, DNA double-strand breaks (Canonical pathway, canonical pathway), oxidative stress, topoisomerase cleavage complexes, splicing intermediates, R-loops and in some cases by DNA repair, single-strand DNA breaks. It phosphorylates several key proteins that initiate activation of the DNA damage cell cycle checkpoint, checkpoint, leading to cell cycle arrest, DNA repair or apoptosis. Several of these targets, including p53, CHK2, BRCA1, NBS1 and H2AX are tumor suppressors. In 1995, the gene was discovered by Yosef Shiloh who named its product ATM since he found that its mutations are responsible for the disorder ataxia–telangiectasia#Cause, ataxia–telangiectasia. In 1998, the Shiloh and Michael B. Kastan, Kastan laboratories independently showed that ATM is a protein kinase whose activity is enhanced by DNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mre11-Rad50-Nbs1
The MRN complex (MRX complex in yeast) is a protein complex consisting of MRE11A, Mre11, Rad50 and Nbs1 (also known as Nibrin in humans and as Xrs2 in yeast). In eukaryotes, the MRN/X complex plays an important role in the initial processing of DNA repair, double-strand DNA breaks prior to repair by homologous recombination or non-homologous end joining. The MRN complex binds avidly to double-strand breaks both in vitro and in vivo and may serve to tether broken ends prior to repair by non-homologous end joining or to initiate DNA end resection prior to repair by homologous recombination. The MRN complex also participates in activating the checkpoint kinase Ataxia telangiectasia mutated, ATM in response to DNA damage. Production of short single-strand oligonucleotides by Mre11 endonuclease activity has been implicated in ATM activation by the MRN complex. Evolutionary ancestry and biologic function The MRN complex has been mainly studied in eukaryotes. However, recent work shows ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Homology (biology)
In biology, homology is similarity in anatomical structures or genes between organisms of different taxa due to shared ancestry, ''regardless'' of current functional differences. Evolutionary biology explains homologous structures as retained heredity from a common descent, common ancestor after having been subjected to adaptation (biology), adaptive modifications for different purposes as the result of natural selection. The term was first applied to biology in a non-evolutionary context by the anatomist Richard Owen in 1843. Homology was later explained by Charles Darwin's theory of evolution in 1859, but had been observed before this from Aristotle's biology onwards, and it was explicitly analysed by Pierre Belon in 1555. A common example of homologous structures is the forelimbs of vertebrates, where the bat wing development, wings of bats and origin of avian flight, birds, the arms of primates, the front flipper (anatomy), flippers of whales, and the forelegs of quadrupedalis ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Arabidopsis Thaliana
''Arabidopsis thaliana'', the thale cress, mouse-ear cress or arabidopsis, is a small plant from the mustard family (Brassicaceae), native to Eurasia and Africa. Commonly found along the shoulders of roads and in disturbed land, it is generally considered a weed. A winter annual with a relatively short lifecycle, ''A. thaliana'' is a popular model organism in plant biology and genetics. For a complex multicellular eukaryote, ''A. thaliana'' has a relatively small genome of around 135 Base pair#Length measurements, megabase pairs. It was the first plant to have its genome sequenced, and is an important tool for understanding the molecular biology of many plant traits, including flower development and phototropism, light sensing. Description ''Arabidopsis thaliana'' is an annual plant, annual (rarely biennial plant, biennial) plant, usually growing to 20–25 cm tall. The leaf, leaves form a rosette at the base of the plant, with a few leaves also on the flowering Plant ste ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Zygosity
Zygosity (the noun, zygote, is from the Greek "yoked," from "yoke") () is the degree to which both copies of a chromosome or gene have the same genetic sequence. In other words, it is the degree of similarity of the alleles in an organism. Most eukaryotes have two matching sets of chromosomes; that is, they are diploid. Diploid organisms have the same locus (genetics), loci on each of their two sets of homologous chromosomes except that the sequences at these loci may differ between the two chromosomes in a matching pair and that a few chromosomes may be mismatched as part of a chromosomal Sex-determination system#Chromosomal determination, sex-determination system. If both alleles of a diploid organism are the same, the organism is #Homozygous, homozygous at that locus. If they are different, the organism is #Heterozygous, heterozygous at that locus. If one allele is missing, it is #Hemizygous, hemizygous, and, if both alleles are missing, it is #Nullizygous, nullizygous. The ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Meiosis
Meiosis () is a special type of cell division of germ cells in sexually-reproducing organisms that produces the gametes, the sperm or egg cells. It involves two rounds of division that ultimately result in four cells, each with only one copy of each chromosome (haploid). Additionally, prior to the division, genetic material from the paternal and maternal copies of each chromosome is crossed over, creating new combinations of code on each chromosome. Later on, during fertilisation, the haploid cells produced by meiosis from a male and a female will fuse to create a zygote, a cell with two copies of each chromosome. Errors in meiosis resulting in aneuploidy (an abnormal number of chromosomes) are the leading known cause of miscarriage and the most frequent genetic cause of developmental disabilities. In meiosis, DNA replication is followed by two rounds of cell division to produce four daughter cells, each with half the number of chromosomes as the original parent cell. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Somatic Cell
In cellular biology, a somatic cell (), or vegetal cell, is any biological cell forming the body of a multicellular organism other than a gamete, germ cell, gametocyte or undifferentiated stem cell. Somatic cells compose the body of an organism and divide through mitosis. In contrast, gametes derive from meiosis within the germ cells of the germline and they fuse during sexual reproduction. Stem cells also can divide through mitosis, but are different from somatic in that they differentiate into diverse specialized cell types. In mammals, somatic cells make up all the internal organs, skin, bones, blood and connective tissue, while mammalian germ cells give rise to spermatozoa and ova which fuse during fertilization to produce a cell called a zygote, which divides and differentiates into the cells of an embryo. There are approximately 220 types of somatic cell in the human body. Theoretically, these cells are not germ cells (the source of gametes); they transmit their mut ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |