|
Double-strand Break Repair Model
A double-strand break repair model refers to the various models of pathways that cells undertake to repair double-strand breaks (DSB). DSB repair is an important cellular process, as the accumulation of unrepaired DSB could lead to chromosomal rearrangements, tumorigenesis or even cell death. In human cells, there are two main DSB repair mechanisms: Homologous recombination (HR) and non-homologous end joining (NHEJ). HR can be seen as a more accurate site specific form of repair. It requires much more larger and intricate protein complexes. These complexes that involve proteins such as RAD51 (searches for homology and mediates strand invasion) and BRCA2 (the well studied RAD51 localizer) are critical in support of DNA replication and the recovery of stalled or broken replication forks. NHEJ modifies and ligates the damaged ends regardless of homology. In terms of DSB repair pathway choice, most mammalian cells appear to favor NHEJ rather than HR. This is because the employment of H ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Double-strand Break Repair Pathway Models
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" (or "Watson–Crick–Franklin") base pairs (guanine–cytosine and adenine–thymine) allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding proteins ca ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
R-loop
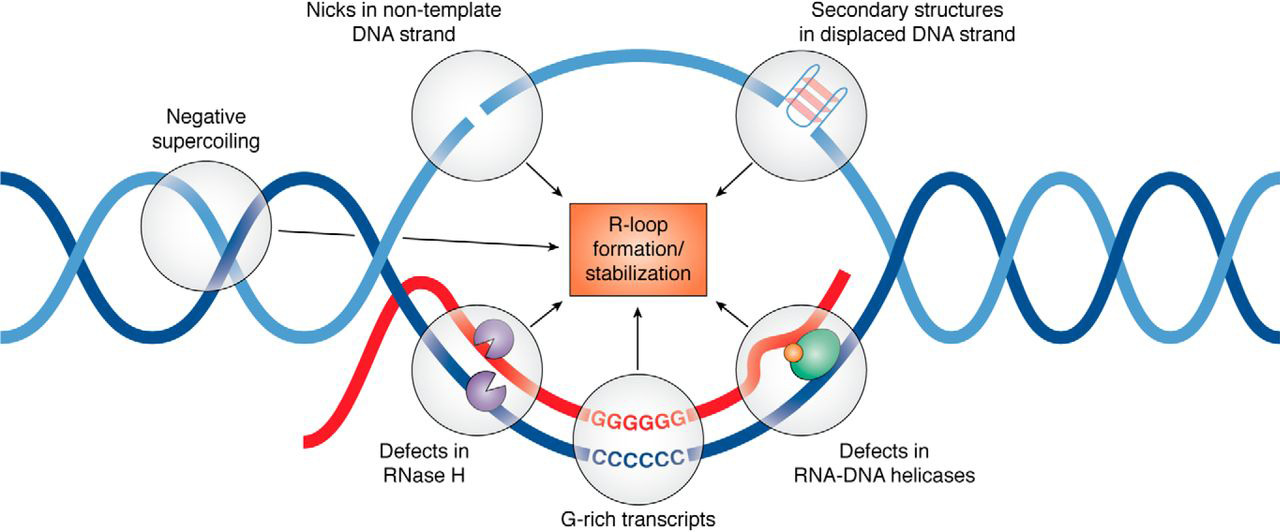
An R-loop is a three-stranded nucleic acid structure, composed of a DNA:RNA hybrid and the associated non-template single-stranded DNA. R-loops may be formed in a variety of circumstances and may be tolerated or cleared by cellular components. The term "R-loop" was given to reflect the similarity of these structures to D-loops; the "R" in this case represents the involvement of an RNA moiety (chemistry), moiety. In the laboratory, R-loops can be created by transcription of DNA sequences (for example those that have a high GC content) that favor annealing of the RNA behind the progressing RNA polymerase. At least 100bp of DNA:RNA hybrid is required to form a stable R-loop structure. R-loops may also be created by the Nucleic acid hybridization#Hybridization, hybridization of mature mRNA with double-stranded DNA under conditions favoring the formation of a DNA-RNA hybrid; in this case, the intron regions (which have been Splicing (genetics), spliced out of the mRNA) form single-stra ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ERCC1
DNA excision repair protein ERCC-1 is a protein that in humans is encoded by the ''ERCC1'' gene. Together with ERCC4, ERCC1 forms the ERCC1-XPF enzyme complex that participates in DNA repair and DNA recombination. Many aspects of these two gene products are described together here because they are partners during DNA repair. The ERCC1-XPF nuclease is an essential activity in the pathway of DNA nucleotide excision repair (NER). The ERCC1-XPF nuclease also functions in pathways to repair double-strand breaks in DNA, and in the repair of “crosslink” damage that harmfully links the two DNA strands. Cells with disabling mutations in ''ERCC1'' are more sensitive than normal to particular DNA damaging agents, including ultraviolet (UV) radiation and to chemicals that cause crosslinking between DNA strands. Genetically engineered mice with disabling mutations in ERCC1 have defects in DNA repair, accompanied by metabolic stress-induced changes in physiology that result in prematur ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Somatic Cell
In cellular biology, a somatic cell (), or vegetal cell, is any biological cell forming the body of a multicellular organism other than a gamete, germ cell, gametocyte or undifferentiated stem cell. Somatic cells compose the body of an organism and divide through mitosis. In contrast, gametes derive from meiosis within the germ cells of the germline and they fuse during sexual reproduction. Stem cells also can divide through mitosis, but are different from somatic in that they differentiate into diverse specialized cell types. In mammals, somatic cells make up all the internal organs, skin, bones, blood and connective tissue, while mammalian germ cells give rise to spermatozoa and ova which fuse during fertilization to produce a cell called a zygote, which divides and differentiates into the cells of an embryo. There are approximately 220 types of somatic cell in the human body. Theoretically, these cells are not germ cells (the source of gametes); they transmit their mut ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Synthesis-dependent Strand Annealing
Synthesis-dependent strand annealing (SDSA) is a major mechanism of homology-directed repair of DNA double-strand breaks (DSBs). Although many of the features of SDSA were first suggested in 1976, the double-Holliday junction model proposed in 1983 was favored by many researchers. In 1994, studies of double-strand gap repair in ''Drosophila'' were found to be incompatible with the double-Holliday junction model, leading researchers to propose a model they called synthesis-dependent strand annealing. Subsequent studies of meiotic recombination in ''S. cerevisiae'' found that non-crossover products appear earlier than double-Holliday junctions or crossover products, challenging the previous notion that both crossover and non-crossover products are produced by double-Holliday junctions and leading the authors to propose that non-crossover products are generated through SDSA. In the accompanying Figure, the first step labeled “5’ to 3’ resection” shows the formation of a 3’ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Holliday Junction
A Holliday junction is a branched nucleic acid structure that contains four double-stranded arms joined. These arms may adopt one of several conformations depending on buffer salt concentrations and the sequence of nucleobases closest to the junction. The structure is named after Robin Holliday, the molecular biologist who proposed its existence in 1964. In biology, Holliday junctions are a key intermediate in many types of genetic recombination, as well as in double-strand break repair. These junctions usually have a symmetrical sequence and are thus mobile, meaning that the four individual arms may slide through the junction in a specific pattern that largely preserves base pairing. Additionally, four-arm junctions similar to Holliday junctions appear in some functional RNA molecules. Immobile Holliday junctions, with asymmetrical sequences that lock the strands in a specific position, were artificially created by scientists to study their structure as a model for natura ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Replication Protein A
Replication protein A (RPA) is the major protein that binds to single-stranded DNA (ssDNA) in eukaryotic cells. In vitro, RPA shows a much higher affinity for ssDNA than RNA or double-stranded DNA. RPA is required in replication, recombination and repair processes such as nucleotide excision repair and homologous recombination. It also plays roles in responding to damaged DNA. Structure RPA is a heterotrimer, composed of the subunits RPA1 (RPA70) (70kDa subunit), RPA2 (RPA32) (32kDa subunit) and RPA3 (RPA14) (14kDa subunit). The three RPA subunits contain six OB-folds (oligonucleotide/oligosaccharide binding), with DNA-binding domains (DBD) designated DBDs A-F, that bind RPA to single-stranded DNA. DBDs A, B, C and F are located on RPA1, DBD D is located on RPA2, and DBD E is located on RPA3. DBDs C, D, and E make up the trimerization core of the protein with flexible linker regions connecting them all together. Due to these flexible linker regions RPA is consider ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RAD52
RAD52 homolog (S. cerevisiae), also known as RAD52, is a protein which in humans is encoded by the ''RAD52'' gene. Function The protein encoded by this gene shares similarity with ''Saccharomyces cerevisiae'' Rad52, a protein important for DNA double-strand break repair and homologous recombination. This gene product was shown to bind single-stranded DNA ends, and mediate the DNA-DNA interaction necessary for the annealing of complementary DNA strands. It was also found to interact with DNA recombination protein RAD51, which suggested its role in RAD51-related DNA recombination and repair. ''aka'' RAD52 homolog, DNA repair protein ''Homo sapiens'' (human)">nowiki>''Homo sapiens'' (human)/ref> Role in DNA recombination repair RAD52 mediates RAD51 function in homologous recombinational repair (HRR) in both yeast ''Saccharomyces cerevisiae'' and in mammalian cells of mice and humans. However, the RAD52 protein has distinctly different functions in HRR of yeast and humans. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RecA
RecA is a 38 kilodalton protein essential for the repair and maintenance of DNA in bacteria. Structural and functional homologs to RecA have been found in all kingdoms of life. RecA serves as an archetype for this class of homologous DNA repair proteins. The homologous protein is called RAD51 in eukaryotes and RadA in archaea. RecA has multiple activities, all related to DNA repair. In the bacterial SOS response, it functions as a co-protease in the autocatalytic cleavage of the LexA repressor and the λ repressor. Function Homologous recombination The RecA protein binds strongly and in long clusters to ssDNA to form a nucleoprotein filament. The protein has multiple DNA binding sites, and thus can hold a single strand and double strand together. This feature makes it possible to catalyze a DNA synapsis reaction between a DNA double helix and a complementary region of single-stranded DNA. The RecA-ssDNA filament searches for sequence similarity along the dsDNA. A disordere ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RAD51
DNA repair protein RAD51 homolog 1 is a protein encoded by the gene ''RAD51''. The enzyme encoded by this gene is a member of the RAD51 protein family which assists in repair of DNA double strand breaks. RAD51 family members are homologous to the bacterial RecA, Archaeal RadA, and yeast Rad51. The protein is highly conserved in most eukaryotes, from yeast to humans. The name RAD51 derives from ''radiation sensitive protein 51''. Variants Two alternatively spliced transcript variants of this gene have been reported, which encode distinct proteins. Transcript variants utilizing alternative polyA signals also exist. Family In mammals, seven recA-like genes have been identified: Rad51, Rad51L1/B, Rad51L2/C, Rad51L3/D, XRCC2, XRCC3, and DMC1/Lim15. All of these proteins, with the exception of meiosis-specific DMC1, are essential for development in mammals. Rad51 is a member of thRecA-like NTPases Function In humans, RAD51 is a 339-amino acid protein that plays ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Double-strand Break Repair Models That Act Via Homologous Recombination
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA and RNA. Dictated by specific hydrogen bonding patterns, "Watson–Crick" (or "Watson–Crick–Franklin") base pairs (guanine–cytosine and adenine–thymine) allow the DNA helix to maintain a regular helical structure that is subtly dependent on its nucleotide sequence. The complementary nature of this based-paired structure provides a redundant copy of the genetic information encoded within each strand of DNA. The regular structure and data redundancy provided by the DNA double helix make DNA well suited to the storage of genetic information, while base-pairing between DNA and incoming nucleotides provides the mechanism through which DNA polymerase replicates DNA and RNA polymerase transcribes DNA into RNA. Many DNA-binding proteins ca ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BRCA2
''BRCA2'' and BRCA2 () are human genes and their protein products, respectively. The official symbol (BRCA2, italic for the gene, nonitalic for the protein) and the official name (originally breast cancer 2; currently BRCA2, DNA repair associated) are gene nomenclature, maintained by the HUGO Gene Nomenclature Committee. One alternative symbol, FANCD1, recognizes its association with the FANC proteins, FANC protein complex. Orthologs, styled ''Brca2'' and Brca2, are common in other vertebrate species. May 2021 ''BRCA2'' is a human tumor suppressor gene (specifically, a caretaker gene), found in all humans; its protein, also called by the synonym breast cancer type 2 susceptibility protein, is responsible for repairing DNA. ''BRCA2'' and ''BRCA1'' are normally expressed in the cells of breast and other tissue, where they help repair damaged DNA or destroy cells if DNA cannot be repaired. They are involved in the repair of chromosome, chromosomal damage with an important role in th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |