|
Ribonuclease
Ribonuclease (commonly abbreviated RNase) is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within the EC 2.7 (for the phosphorolytic enzymes) and 3.1 (for the hydrolytic enzymes) classes of enzymes. Function All organisms studied contain many RNases of two different classes, showing that RNA degradation is a very ancient and important process. As well as clearing of cellular RNA that is no longer required, RNases play key roles in the maturation of all RNA molecules, both messenger RNAs that carry genetic material for making proteins and non-coding RNAs that function in varied cellular processes. In addition, active RNA degradation systems are the first defense against RNA viruses and provide the underlying machinery for more advanced cellular immune strategies such as RNAi. Some cells also secrete copious quantities of non-specific ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNase H
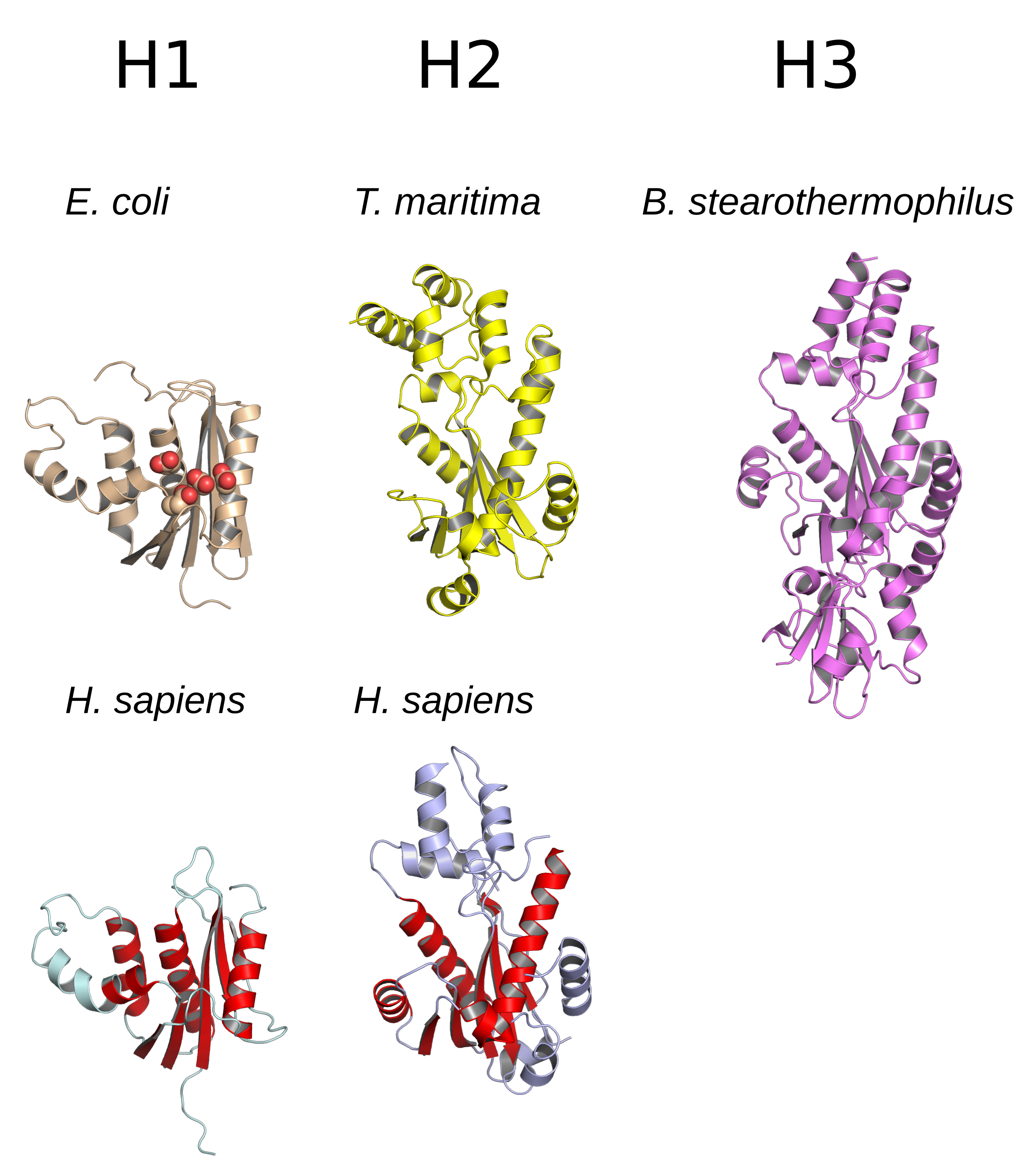
Ribonuclease H (abbreviated RNase H or RNH) is a family of non-sequence-specific endonuclease enzymes that catalyze the cleavage of RNA in an RNA/ DNA substrate via a hydrolytic mechanism. Members of the RNase H family can be found in nearly all organisms, from bacteria to archaea to eukaryotes. The family is divided into evolutionarily related groups with slightly different substrate preferences, broadly designated ribonuclease H1 and H2. The human genome encodes both H1 and H2. Human ribonuclease H2 is a heterotrimeric complex composed of three subunits, mutations in any of which are among the genetic causes of a rare disease known as Aicardi–Goutières syndrome. A third type, closely related to H2, is found only in a few prokaryotes, whereas H1 and H2 occur in all domains of life. Additionally, RNase H1-like retroviral ribonuclease H domains occur in multidomain reverse transcriptase proteins, which are encoded by retroviruses such as HIV and are required for viral rep ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNase III
Ribonuclease III (RNase III or RNase C)(BREND3.1.26.3 is a type of ribonuclease that recognizes dsRNA and cleaves it at specific targeted locations to transform them into mature RNAs. These enzymes are a group of endoribonucleases that are characterized by their ribonuclease domain, which is labelled the RNase III domain. They are ubiquitous compounds in the cell and play a major role in pathways such as RNA precursor synthesis, RNA Silencing, and the ''pnp'' autoregulatory mechanism. Types of RNase III The RNase III superfamily is divided into four known classes: 1, 2, 3, and 4. Each class is defined by its domain structure.Liang Y-H, Lavoie M, Comeau M-A, Elela SA, Ji X. Structure of a Eukaryotic RNase III Post-Cleavage Complex Reveals a Double- Ruler Mechanism for Substrate Selection. Molecular cell. 2014;54(3):431-444. doi:10.1016/j.molcel.2014.03.006. Class 1 RNase III *Class 1 RNase III enzymes have a homodimeric structure whose function is to cleave dsRNA into multiple ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNase T1
Ribonuclease (commonly abbreviated RNase) is a type of nuclease that catalyzes the degradation of RNA into smaller components. Ribonucleases can be divided into endoribonucleases and exoribonucleases, and comprise several sub-classes within the EC 2.7 (for the phosphorolytic enzymes) and 3.1 (for the hydrolytic enzymes) classes of enzymes. Function All organisms studied contain many RNases of two different classes, showing that RNA degradation is a very ancient and important process. As well as clearing of cellular RNA that is no longer required, RNases play key roles in the maturation of all RNA molecules, both messenger RNAs that carry genetic material for making proteins and non-coding RNAs that function in varied cellular processes. In addition, active RNA degradation systems are the first defense against RNA viruses and provide the underlying machinery for more advanced cellular immune strategies such as RNAi. Some cells also secrete copious quantities of non-specific RNa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNase P
Ribonuclease P (, ''RNase P'') is a type of ribonuclease which cleaves RNA. RNase P is unique from other RNases in that it is a ribozyme – a ribonucleic acid that acts as a catalyst in the same way that a protein-based enzyme would. Its function is to cleave off an extra, or precursor, sequence of RNA on tRNA molecules. Further, RNase P is one of two known multiple turnover ribozymes in nature (the other being the ribosome), the discovery of which earned Sidney Altman and Thomas Cech the Nobel Prize in Chemistry in 1989: in the 1970s, Altman discovered the existence of precursor tRNA with flanking sequences and was the first to characterize RNase P and its activity in processing of the 5' leader sequence of precursor tRNA. Recent findings also reveal that RNase P has a new function. It has been shown that human nuclear RNase P is required for the normal and efficient transcription of various small noncoding RNAs, such as tRNA, 5S rRNA, SRP RNA and U6 snRNA genes, which ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribonuclease A
Pancreatic ribonuclease family (, ''RNase'', ''RNase I'', ''RNase A'', ''pancreatic RNase'', ''ribonuclease I'', ''endoribonuclease I'', ''ribonucleic phosphatase'', ''alkaline ribonuclease'', ''ribonuclease'', ''gene S glycoproteins'', ''Ceratitis capitata alkaline ribonuclease'', ''SLSG glycoproteins'', ''gene S locus-specific glycoproteins'', ''S-genotype-assocd. glycoproteins'', ''ribonucleate 3'-pyrimidino-oligonucleotidohydrolase'') is a superfamily of pyrimidine-specific endonucleases found in high quantity in the pancreas of certain mammals and of some reptiles. Specifically, the enzymes are involved in endonucleolytic cleavage of 3'-phosphomononucleotides and 3'-phosphooligonucleotides ending in C-P or U-P with 2',3'-cyclic phosphate intermediates. Ribonuclease can unwind the RNA helix by complexing with single-stranded RNA; the complex arises by an extended multi-site cation-anion interaction between lysine and arginine residues of the enzyme and phosphate groups of the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribonuclease Inhibitor
Ribonuclease inhibitor (RI) is a large (~450 residues, ~49 kDa), acidic (pI ~4.7), leucine-rich repeat protein that forms extremely tight complexes with certain ribonucleases. It is a major cellular protein, comprising ~0.1% of all cellular protein by weight, and appears to play an important role in regulating the lifetime of RNA. RI has a surprisingly high cysteine content (~6.5%, cf. 1.7% in typical proteins) and is sensitive to oxidation. RI is also rich in leucine (21.5%, compared to 9% in typical proteins) and commensurately lower in other hydrophobic residues, esp. valine, isoleucine, methionine, tyrosine, and phenylalanine. Structure RI is the classic leucine-rich repeat protein, consisting of alternating α-helices and β-strands along its backbone. These secondary structure elements wrap around in a curved, right-handed solenoid that resembles a horseshoe. The parallel β-strands and α-helices form the inner and outer wall of the horseshoe, respectively. The s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNase L
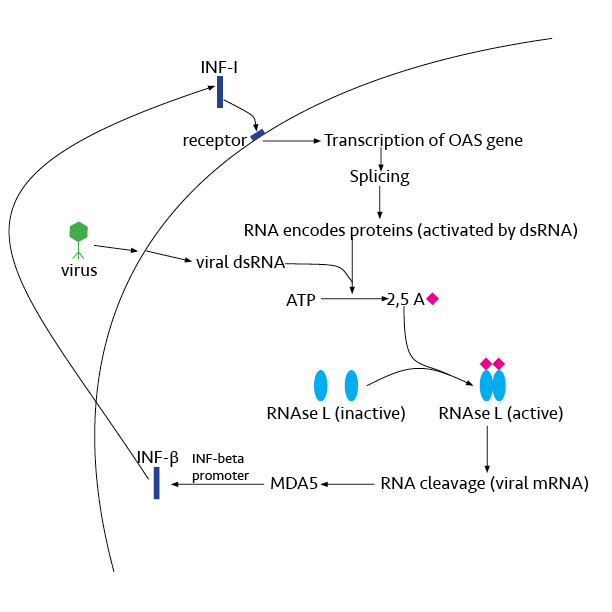
Ribonuclease L or RNase L (for ''latent''), known sometimes as ribonuclease 4 or 2'-5' oligoadenylate synthetase-dependent ribonuclease — is an interferon (IFN)-induced ribonuclease which, upon activation, destroys all RNA within the cell (both cellular and viral). RNase L is an enzyme that in humans is encoded by the ''RNASEL'' gene. This gene encodes a component of the interferon-regulated 2'-5'oligoadenylate (2'-5'A) system that functions in the antiviral and antiproliferative roles of interferons. RNase L is activated by dimerization, which occurs upon 2'-5'A binding, and results in cleavage of all RNA in the cell. This can lead to activation of MDA5, an RNA helicase involved in the production of interferons. Synthesis and activation RNase L is present in very minute quantities during the normal cell cycle. When interferon binds to cell receptors, it activates transcription of around 300 genes to bring about the antiviral state. Among the enzymes produced is RNase L ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
TRNA
Transfer RNA (abbreviated tRNA and formerly referred to as sRNA, for soluble RNA) is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes), that serves as the physical link between the mRNA and the amino acid sequence of proteins. tRNAs genes from Bacteria are typically shorter (mean = 77.6 bp) than tRNAs from Archaea (mean = 83.1 bp) and eukaryotes (mean = 84.7 bp). The mature tRNA follows an opposite pattern with tRNAs from Bacteria being usually longer (median = 77.6 nt) than tRNAs from Archaea (median = 76.8 nt), with eukaryotes exhibiting the shortest mature tRNAs (median = 74.5 nt). Transfer RNA (tRNA) does this by carrying an amino acid to the protein synthesizing machinery of a cell called the ribosome. Complementation of a 3-nucleotide codon in a messenger RNA (mRNA) by a 3-nucleotide anticodon of the tRNA results in protein synthesis based on the mRNA code. As such, tRNAs are a necessary component of translation, the biological ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exoribonuclease
An exoribonuclease is an exonuclease ribonuclease, which are enzymes that degrade RNA by removing terminal nucleotides from either the 5' end or the 3' end of the RNA molecule. Enzymes that remove nucleotides from the 5' end are called ''5'-3' exoribonucleases'', and enzymes that remove nucleotides from the 3' end are called ''3'-5' exoribonucleases''. Exoribonucleases can use either water to cleave the nucleotide-nucleotide bond (which is called hydrolytic activity) or inorganic phosphate (which is called phosphorolytic activity). Hydrolytic exoribonucleases are classified under EC number 3.1 and phosphorolytic exoribonucleases under EC number 2.7.7. As the phosphorolytic enzymes use inorganic phosphate to cleave bonds they release nucleotide diphosphates, whereas the hydrolytic enzymes (which use water) release nucleotide monosphosphates. Exoribonucleases exist in all kingdoms of life, the bacteria, archaea, and eukaryotes. Exoribonucleases are involved in the degradation ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Dissociation Constant
In chemistry, biochemistry, and pharmacology, a dissociation constant (K_D) is a specific type of equilibrium constant that measures the propensity of a larger object to separate (dissociate) reversibly into smaller components, as when a complex falls apart into its component molecules, or when a salt splits up into its component ions. The dissociation constant is the inverse of the association constant. In the special case of salts, the dissociation constant can also be called an ionization constant. For a general reaction: : A_\mathit B_\mathit \mathit A + \mathit B in which a complex \ce_x \ce_y breaks down into ''x'' A subunits and ''y'' B subunits, the dissociation constant is defined as : K_D = \frac where and ''x'' B''y''are the equilibrium concentrations of A, B, and the complex A''x'' B''y'', respectively. One reason for the popularity of the dissociation constant in biochemistry and pharmacology is that in the frequently en ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribozyme
Ribozymes (ribonucleic acid enzymes) are RNA molecules that have the ability to catalyze specific biochemical reactions, including RNA splicing in gene expression, similar to the action of protein enzymes. The 1982 discovery of ribozymes demonstrated that RNA can be both genetic material (like DNA) and a biological catalyst (like protein enzymes), and contributed to the RNA world hypothesis, which suggests that RNA may have been important in the evolution of prebiotic self-replicating systems. The most common activities of natural or in vitro-evolved ribozymes are the cleavage or ligation of RNA and DNA and peptide bond formation. For example, the smallest ribozyme known (GUGGC-3') can aminoacylate a GCCU-3' sequence in the presence of PheAMP. Within the ribosome, ribozymes function as part of the large subunit ribosomal RNA to link amino acids during protein synthesis. They also participate in a variety of RNA processing reactions, including RNA splicing, viral replication, and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |