|
PA Clan
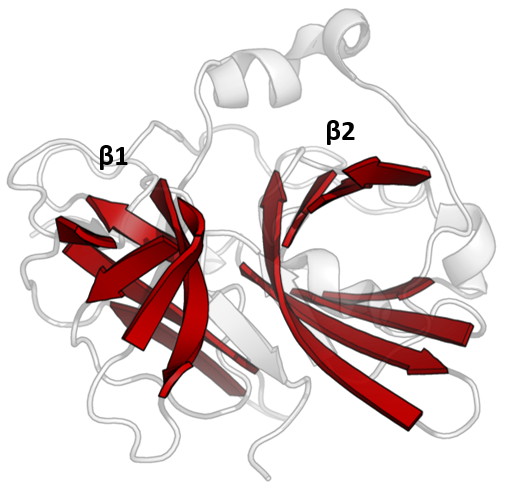
The PA clan ( Proteases of mixed nucleophile, superfamily A) is the largest group of proteases with common ancestry as identified by structural homology. Members have a chymotrypsin-like fold and similar proteolysis mechanisms but can have identity of <10%. The clan contains both and serine proteases (different nucleophiles). PA clan proteases can be found in , |
Protease
A protease (also called a peptidase, proteinase, or proteolytic enzyme) is an enzyme that catalysis, catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks Covalent bond, bonds. Proteases are involved in numerous biological pathways, including Digestion#Protein digestion, digestion of ingested proteins, protein catabolism (breakdown of old proteins), and cell signaling. In the absence of functional accelerants, proteolysis would be very slow, taking hundreds of years. Proteases can be found in all forms of life and viruses. They have independently convergent evolution, evolved multiple times, and different classes of protease can perform the same reaction by completely different catalytic mechanisms. Classification Based on catalytic residue Proteases can be classified into seven broad ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Convergent Evolution
Convergent evolution is the independent evolution of similar features in species of different periods or epochs in time. Convergent evolution creates analogous structures that have similar form or function but were not present in the last common ancestor of those groups. The cladistic term for the same phenomenon is Cladogram#Homoplasies, homoplasy. The recurrent evolution of flight is a classic example, as flying pterygota, insects, birds, pterosaurs, and bats have independently evolved the useful capacity of flight. Functionally similar features that have arisen through convergent evolution are ''analogous'', whereas ''homology (biology), homologous'' structures or traits have a common origin but can have dissimilar functions. Bird, bat, and pterosaur wings are analogous structures, but their forelimbs are homologous, sharing an ancestral state despite serving different functions. The opposite of convergence is divergent evolution, where related species evolve different trai ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tobacco Etch Virus Protease
TEV protease (, ''Tobacco Etch Virus nuclear-inclusion-a endopeptidase'') is a highly sequence-specific cysteine protease from Tobacco Etch Virus (TEV). It is a member of the PA clan of chymotrypsin-like proteases. Due to its high sequence specificity, TEV protease is frequently used for the controlled cleavage of fusion proteins ''in vitro'' and ''in vivo''. The consensus sequence recognized by TEV protease is Glu-Asn-Leu-Tyr-Phe-Gln-, -Ser, where ", " denotes cleaved peptide bond. Origin The tobacco etch virus encodes its entire genome as a single massive polyprotein (350 kDa). This is cleaved into functional units by the three proteases: P1 protease (1 cleavage site), helper-component protease (1 cleavage site) and TEV protease (7 cleavage sites).UniProt: TEV polyprotein: The native TEV protease also contains an internal self-cleavage site. This site is slowly cleaved to inactivate the enzyme (the physiological reason for this is unknown). Structure and function Th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Thrombin
Prothrombin (coagulation factor II) is encoded in the human by the F2-gene. It is proteolytically cleaved during the clotting process by the prothrombinase enzyme complex to form thrombin. Thrombin (Factor IIa) (, fibrose, thrombase, thrombofort, topical, thrombin-C, tropostasin, activated blood-coagulation factor II, E thrombin, beta-thrombin, gamma-thrombin) is a serine protease, that converts fibrinogen into strands of insoluble fibrin, as well as catalyzing many other coagulation-related reactions. History After the description of fibrinogen and fibrin, Alexander Schmidt hypothesised the existence of an enzyme that converts fibrinogen into fibrin in 1872. Prothrombin was discovered by Pekelharing in 1894. Physiology Synthesis Thrombin is produced by the enzymatic cleavage of two sites on prothrombin by activated Factor X (Xa). The activity of factor Xa is greatly enhanced by binding to activated Factor V (Va), termed the prothrombinase complex. Prothrombin is prod ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chymotrypsin
Chymotrypsin (, chymotrypsins A and B, alpha-chymar ophth, avazyme, chymar, chymotest, enzeon, quimar, quimotrase, alpha-chymar, alpha-chymotrypsin A, alpha-chymotrypsin) is a digestive enzyme component of pancreatic juice acting in the duodenum, where it performs proteolysis, the breakdown of proteins and polypeptides. Chymotrypsin preferentially cleaves peptide amide bonds where the side chain of the amino acid N-terminal to the scissile amide bond (the P1 position) is a large hydrophobic amino acid ( tyrosine, tryptophan, and phenylalanine). These amino acids contain an aromatic ring in their side chain that fits into a hydrophobic pocket (the S1 position) of the enzyme. It is activated in the presence of trypsin. The hydrophobic and shape complementarity between the peptide substrate P1 side chain and the enzyme S1 binding cavity accounts for the substrate specificity of this enzyme. Chymotrypsin also hydrolyzes other amide bonds in peptides at slower rates, particularly th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Structural Homology
A protein superfamily is the largest grouping (clade) of proteins for which common ancestry can be inferred (see homology). Usually this common ancestry is inferred from structural alignment and mechanistic similarity, even if no sequence similarity is evident. Sequence homology can then be deduced even if not apparent (due to low sequence similarity). Superfamilies typically contain several protein families which show sequence similarity within each family. The term ''protein clan'' is commonly used for protease and glycosyl hydrolases superfamilies based on the MEROPS and CAZy classification systems. Identification Superfamilies of proteins are identified using a number of methods. Closely related members can be identified by different methods to those needed to group the most evolutionarily divergent members. Sequence similarity Historically, the similarity of different amino acid sequences has been the most common method of inferring homology. Sequence similarity is ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MEROPS
MEROPS is an online database for peptidases (also known as proteases, proteinases and proteolytic enzymes) and their inhibitors. The classification scheme for peptidases was published by Rawlings & Barrett in 1993, and that for protein inhibitors by Rawlings ''et al.'' in 2004.Rawlings, N.D., Tolle, D.P. & Barrett, A.J. (2004) "Evolutionary families of peptidase inhibitors." ''Biochem J'' 378, 705-716. The most recent version, MEROPS 12.5, was released in September 2023. Overview The classification is based on similarities at the tertiary and primary structural levels. Comparisons are restricted to that part of the sequence directly involved in the reaction, which in the case of a peptidase must include the active site, and for a protein inhibitor the reactive site. The classification is hierarchical: sequences are assembled into families, and families are assembled into clans. Each peptidase, family, and clan has a unique identifier. Classification Family The families of pe ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Superfamily
A protein superfamily is the largest grouping (clade) of proteins for which common ancestry can be inferred (see homology (biology), homology). Usually this common ancestry is inferred from structural alignment and mechanistic similarity, even if no sequence similarity is evident. Sequence homology can then be deduced even if not apparent (due to low sequence similarity). Superfamilies typically contain several protein families which show sequence similarity within each family. The term ''protein clan'' is commonly used for protease and glycosyl hydrolases superfamilies based on the MEROPS and CAZy classification systems. Identification Superfamilies of proteins are identified using a number of methods. Closely related members can be identified by different methods to those needed to group the most evolutionarily divergent members. Sequence similarity Historically, the similarity of different amino acid sequences has been the most common method of inferring Sequence homology, h ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tobacco Etch Virus Protease
TEV protease (, ''Tobacco Etch Virus nuclear-inclusion-a endopeptidase'') is a highly sequence-specific cysteine protease from Tobacco Etch Virus (TEV). It is a member of the PA clan of chymotrypsin-like proteases. Due to its high sequence specificity, TEV protease is frequently used for the controlled cleavage of fusion proteins ''in vitro'' and ''in vivo''. The consensus sequence recognized by TEV protease is Glu-Asn-Leu-Tyr-Phe-Gln-, -Ser, where ", " denotes cleaved peptide bond. Origin The tobacco etch virus encodes its entire genome as a single massive polyprotein (350 kDa). This is cleaved into functional units by the three proteases: P1 protease (1 cleavage site), helper-component protease (1 cleavage site) and TEV protease (7 cleavage sites).UniProt: TEV polyprotein: The native TEV protease also contains an internal self-cleavage site. This site is slowly cleaved to inactivate the enzyme (the physiological reason for this is unknown). Structure and function Th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Crystallography
X-ray crystallography is the experimental science of determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to diffract in specific directions. By measuring the angles and intensities of the X-ray diffraction, a crystallographer can produce a three-dimensional picture of the density of electrons within the crystal and the positions of the atoms, as well as their chemical bonds, crystallographic disorder, and other information. X-ray crystallography has been fundamental in the development of many scientific fields. In its first decades of use, this method determined the size of atoms, the lengths and types of chemical bonds, and the atomic-scale differences between various materials, especially minerals and alloys. The method has also revealed the structure and function of many biological molecules, including vitamins, drugs, proteins and nucleic acids such as DNA. X-ray crystallography is still the prima ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
S1 Family
S1, S01, S.I, S-1, S.1, Š-1 or S 1 may refer to: Biology and chemistry * S1 nuclease, an enzyme that digests singled-stranded DNA and RNA * S1: Keep locked up, a safety phrase in chemistry * Primary somatosensory cortex, also known as S1 * Tegafur/gimeracil/oteracil, also known as S-1, a chemotherapy medication Entertainment * S1 (Indian TV channel), a Hindi-language channel * S1 (Swiss TV channel), a German-language channel * S1 (producer), a hip hop producer, member of the group Strange Fruit Project * S1 No. 1 Style, a Japanese adult video company * Gibson S-1, a guitar made by the Gibson Guitar Corporation * A member of the S1W (group) that later became part of the music group Public Enemy. Government * Bill S-1, a pro forma bill in Canadian Parliament * Form S-1, a U.S. Securities and Exchange Commission filing * S-1 Executive Committee, a United States government entity during World War II * S1 (military), an administrative position within military units Technology * ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Serine Protease
Serine proteases (or serine endopeptidases) are enzymes that cleave peptide bonds in proteins. Serine serves as the nucleophilic amino acid at the (enzyme's) active site. They are found ubiquitously in both eukaryotes and prokaryotes. Serine proteases fall into two broad categories based on their structure: chymotrypsin-like (trypsin-like) or subtilisin-like. Classification The MEROPS protease classification system counts 16 protein superfamily, superfamilies (as of 2013) each containing many protein family, families. Each superfamily uses the catalytic triad or dyad in a different protein fold and so represent convergent evolution of the catalytic mechanism. The majority belong to the S1 family of the PA clan (superfamily) of proteases. For protein superfamily, superfamilies, P: superfamily, containing a mixture of nucleophile class families, S: purely serine proteases. superfamily. Within each superfamily, protein family, families are designated by their catalytic nucl ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |