|
Amplified Ribosomal DNA Restriction Analysis
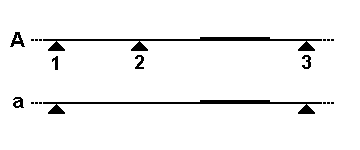
Amplified rDNA (Ribosomal DNA) Restriction Analysis is the extension of the technique of RFLP (restriction fragment length polymorphism) to the gene encoding the small (16s) ribosomal subunit of bacteria. The technique involves an enzymatic amplification using primers directed at the conserved regions at the ends of the 16s gene, followed by digestion using tetracutter Restriction enzymes. The pattern obtained is said to be representative of the species analysed. Patterns obtained from several restriction enzymes can be used to phylogenetically characterize cultured isolates and 16s genes obtained through cloning from community DNA ARDRA rationale and procedure Vaneechoutte ''et al.'', were among the first to use this method and applied it to characterize Mycobacterium species and Acinetobacter species Numerous other studies have applied ARDRA for different bacterial taxa. Based on the simple formula for the frequency of random occurrence of a Restriction site, a 4-bp sequence can o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Fragment Length Polymorphism
In molecular biology, restriction fragment length polymorphism (RFLP) is a technique that exploits variations in homologous DNA sequences, known as polymorphisms, populations, or species or to pinpoint the locations of genes within a sequence. The term may refer to a polymorphism itself, as detected through the differing locations of restriction enzyme sites, or to a related laboratory technique by which such differences can be illustrated. In RFLP analysis, a DNA sample is digested into fragments by one or more restriction enzymes, and the resulting ''restriction fragments'' are then separated by gel electrophoresis according to their size. RFLP analysis is now largely obsolete due to the emergence of inexpensive DNA sequencing technologies, but it was the first DNA profiling technique inexpensive enough to see widespread application. RFLP analysis was an important early tool in genome mapping, localization of genes for genetic disorders, determination of risk for disease, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
16S Ribosomal RNA
16S ribosomal RNA (or 16 S rRNA) is the RNA component of the 30S subunit of a prokaryotic ribosome ( SSU rRNA). It binds to the Shine-Dalgarno sequence and provides most of the SSU structure. The genes coding for it are referred to as 16S rRNA genes and are used in reconstructing phylogenies, due to the slow rates of evolution of this region of the gene. Carl Woese and George E. Fox were two of the people who pioneered the use of 16S rRNA in phylogenetics in 1977. Multiple sequences of the 16S rRNA gene can exist within a single bacterium. Terminology The descriptor ''16S'' refers to the size of these ribosomal subunits as reflected indirectly by the speed at which they sediment when samples are centrifuged. Thus ''16S'' means 16 Svedburg units. Functions * Like the large (23S) ribosomal RNA, it has a structural role, acting as a scaffold defining the positions of the ribosomal proteins. * The 3-end contains the anti- Shine-Dalgarno sequence, which binds upstream ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
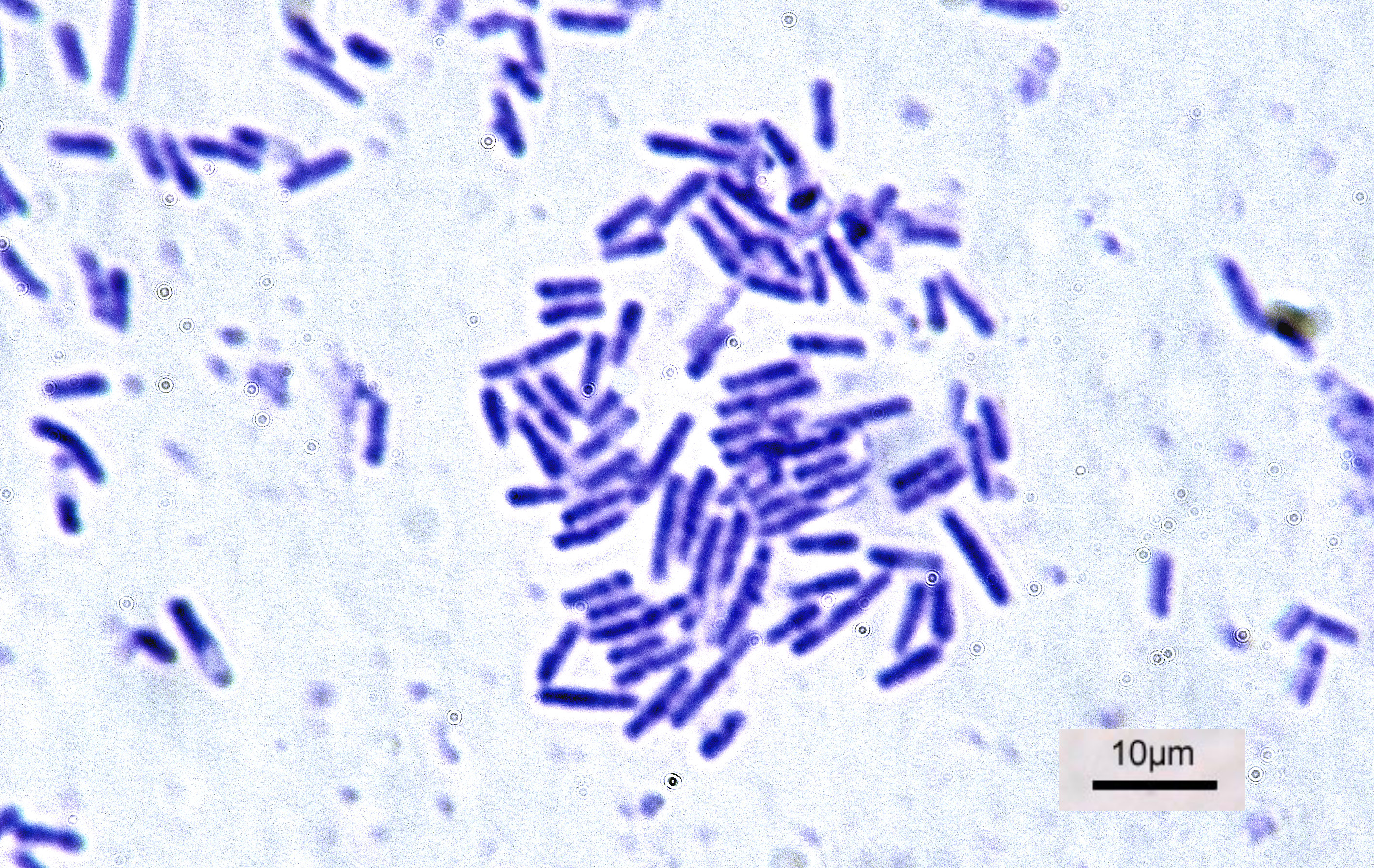
Bacteria
Bacteria (; : bacterium) are ubiquitous, mostly free-living organisms often consisting of one Cell (biology), biological cell. They constitute a large domain (biology), domain of Prokaryote, prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit the air, soil, water, Hot spring, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria play a vital role in many stages of the nutrient cycle by recycling nutrients and the nitrogen fixation, fixation of nitrogen from the Earth's atmosphere, atmosphere. The nutrient cycle includes the decomposition of cadaver, dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, suc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Enzymes
A restriction enzyme, restriction endonuclease, REase, ENase or'' restrictase '' is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class of the broader endonuclease group of enzymes. Restriction enzymes are commonly classified into five types, which differ in their structure and whether they cut their DNA substrate at their recognition site, or if the recognition and cleavage sites are separate from one another. To cut DNA, all restriction enzymes make two incisions, once through each sugar-phosphate backbone (i.e. each strand) of the DNA double helix. These enzymes are found in bacteria and archaea and provide a defense mechanism against invading viruses. Inside a prokaryote, the restriction enzymes selectively cut up ''foreign'' DNA in a process called ''restriction digestion''; meanwhile, host DNA is protected by a modification enzyme (a methyltransferase) that modifi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Random Amplification Of Polymorphic DNA
Random amplified polymorphic DNA (RAPD), pronounced "rapid", is a type of polymerase chain reaction (PCR), but the segments of DNA that are amplified are random. The scientist performing RAPD creates several arbitrary, short primers (10–12 nucleotides), then proceeds with the PCR using a large template of genomic DNA, hoping that fragments will amplify. By resolving the resulting patterns, a semi-unique profile can be gleaned from an RAPD reaction. No knowledge of the DNA sequence of the targeted genome is required, as the primers will bind somewhere in the sequence, but it is not certain exactly where. This makes the method popular for comparing the DNA of biological systems that have not had the attention of the scientific community, or in a system in which relatively few DNA sequences are compared (it is not suitable for forming a cDNA databank). Because it relies on a large, intact DNA template sequence, it has some limitations in the use of degraded DNA samples. Its re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
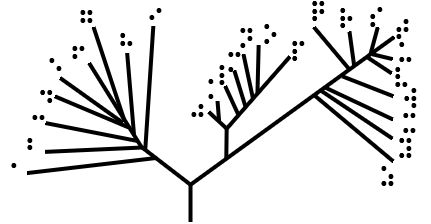
Cladogram
A cladogram (from Greek language, Greek ''clados'' "branch" and ''gramma'' "character") is a diagram used in cladistics to show relations among organisms. A cladogram is not, however, an Phylogenetic tree, evolutionary tree because it does not show how ancestors are related to descendants, nor does it show how much they have changed, so many differing evolutionary trees can be consistent with the same cladogram. A cladogram uses lines that branch off in different directions ending at a clade, a group of organisms with a last common ancestor. There are many shapes of cladograms but they all have lines that branch off from other lines. The lines can be traced back to where they branch off. These branching off points represent a hypothetical ancestor (not an actual entity) which can be inferred to exhibit the traits shared among the terminal taxa above it. This hypothetical ancestor might then provide clues about the order of evolution of various features, adaptation, and other e ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phylogram
A phylogenetic tree or phylogeny is a graphical representation which shows the evolutionary history between a set of species or taxa during a specific time.Felsenstein J. (2004). ''Inferring Phylogenies'' Sinauer Associates: Sunderland, MA. In other words, it is a branching diagram or a tree showing the evolutionary relationships among various biological species or other entities based upon similarities and differences in their physical or genetic characteristics. In evolutionary biology, all life on Earth is theoretically part of a single phylogenetic tree, indicating common ancestry. Phylogenetics is the study of phylogenetic trees. The main challenge is to find a phylogenetic tree representing optimal evolutionary ancestry between a set of species or taxa. Computational phylogenetics (also phylogeny inference) focuses on the algorithms involved in finding optimal phylogenetic tree in the phylogenetic landscape. Phylogenetic trees may be rooted or unrooted. In a ''rooted'' phy ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
BioNumerics
BioNumerics is a bioinformatics desktop software application that manages microbiological data. It is developed by Applied Maths NV, a bioMérieux company. History BioNumerics was first released in 1998. PulseNet, a network run by the Centers for Disease Control and Prevention (CDC), uses BioNumerics to compare pulsed field gel electrophoresis (PFGE) patterns and whole genome sequences from different bacterial strains. CaliciNet, an outbreak surveillance network for noroviruses, is another example of a network which uses BioNumerics to submit norovirus sequences and basic epidemiologic information to a central database. Features The basis of BioNumerics is a database consisting of entries. The entries correspond to the individual organisms or samples under study and are characterized by a unique key and by a number of user-defined information fields. Each entry in a database may be characterized by one or more experiments that can be linked easil ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |