|
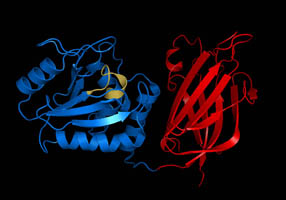
Phosphatase
In biochemistry, a phosphatase is an enzyme that uses water to cleave a phosphoric acid Ester, monoester into a phosphate ion and an Alcohol (chemistry), alcohol. Because a phosphatase enzyme catalysis, catalyzes the hydrolysis of its Substrate (chemistry), substrate, it is a subcategory of hydrolases. Phosphatase enzymes are essential to many biological functions, because phosphorylation (e.g. by protein kinases) and dephosphorylation (by phosphatases) serve diverse roles in cell growth, cellular regulation and cell signaling, signaling. Whereas phosphatases remove phosphate groups from molecules, kinases catalyze the transfer of phosphate groups to molecules from Adenosine triphosphate, ATP. Together, kinases and phosphatases direct a form of post-translational modification that is essential to the cell's regulatory network. Phosphatase enzymes are not to be confused with phosphorylase enzymes, which catalyze the transfer of a phosphate group from hydrogen phosphate to an acce ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphatome
The phosphatome of an organism is the set of phosphatase genes in its genome. Phosphatases are enzymes that catalyze the removal of phosphate from biomolecules. Over half of all cellular proteins are modified by phosphorylation which typically controls their functions. Protein phosphorylation is controlled by the opposing actions of protein phosphatases and protein kinases. Most phosphorylation sites are not linked to a specific phosphatase, so the phosphatome approach allows a global analysis of dephosphorylation, screening to find the phosphatase responsible for a given reaction, and comparative studies between different phosphatases, similar to how protein kinase research has been impacted by the kinome approach. Protein Phosphatome Protein phosphatases remove phosphates from proteins, usually on Serine, Threonine, and Tyrosine residues, reversing the action of protein kinases. The PTP family of protein phosphatases is tyrosine-specific, and several other families (PPPL, PPM, H ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Dephosphorylation
In biochemistry, dephosphorylation is the removal of a phosphate () group from an organic compound by hydrolysis. It is a reversible post-translational modification. Dephosphorylation and its counterpart, phosphorylation, activate and deactivate enzymes by detaching or attaching phosphoric esters and anhydrides. A notable occurrence of dephosphorylation is the conversion of Adenosine triphosphate, ATP to Adenosine diphosphate, ADP and inorganic phosphate. Dephosphorylation employs a type of hydrolytic enzyme, or hydrolase, which cleaves ester bonds. The prominent hydrolase subclass used in dephosphorylation is phosphatase, which removes phosphate groups by hydrolysing phosphoric acid monoesters into a phosphate ion and a molecule with a free hydroxyl (–OH) group. The reversible phosphorylation-dephosphorylation reaction occurs in every physiological process, making proper function of protein phosphatases necessary for organism viability. Because protein dephosphorylation is a k ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Calcineurin
Calcineurin (CaN) is a calcium and calmodulin dependent serine/threonine protein phosphatase (also known as protein phosphatase 3, and calcium-dependent serine-threonine phosphatase). It activates the T cells of the immune system and can be blocked by drugs. Calcineurin activates nuclear factor of activated T cell cytoplasmic (NFATC1, NFATc), a transcription factor, by dephosphorylation, dephosphorylating it. The activated NFATc is then Protein targeting#Post-translational translocation, translocated into the nucleus, where it upregulates the expression of interleukin 2 (IL-2), which, in turn, stimulates the growth and differentiation of the Cell immunity, T cell response. Calcineurin is the target of a class of drugs called Immunosuppressive drug#Drugs acting on immunophilins, calcineurin inhibitors, which include ciclosporin, voclosporin, pimecrolimus and tacrolimus. Structure Calcineurin is a heterodimer of a 61-kD calmodulin-binding catalytic subunit, calcineurin A and a 19- ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Kinase
In biochemistry, a kinase () is an enzyme that catalyzes the transfer of phosphate groups from high-energy, phosphate-donating molecules to specific substrates. This process is known as phosphorylation, where the high-energy ATP molecule donates a phosphate group to the substrate molecule. As a result, kinase produces a phosphorylated substrate and ADP. Conversely, it is referred to as dephosphorylation when the phosphorylated substrate donates a phosphate group and ADP gains a phosphate group (producing a dephosphorylated substrate and the high energy molecule of ATP). These two processes, phosphorylation and dephosphorylation, occur four times during glycolysis. Kinases are part of the larger family of phosphotransferases. Kinases should not be confused with phosphorylases, which catalyze the addition of inorganic phosphate groups to an acceptor, nor with phosphatases, which remove phosphate groups (dephosphorylation). The phosphorylation state of a molecule, whether it ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Phosphorylase
In biochemistry, phosphorylases are enzymes that catalyze the addition of a phosphate group from an inorganic phosphate (phosphate+hydrogen) to an acceptor. :A-B + P A + P-B They include allosteric enzymes that catalyze the production of glucose-1-phosphate from a glucan such as glycogen, starch or maltodextrin. Phosphorylase is also a common name used for glycogen phosphorylase in honor of Earl W. Sutherland Jr., who in the late 1930s discovered it as the first phosphorylase. Function Phosphorylases should not be confused with phosphatases, which remove phosphate groups. In more general terms, phosphorylases are enzymes that catalyze the addition of a phosphate group from an inorganic phosphate (phosphate + hydrogen) to an acceptor, not to be confused with a phosphatase (a hydrolase) or a kinase (a phosphotransferase). A phosphatase removes a phosphate group from a donor using water, whereas a kinase transfers a phosphate group from a donor (usually ATP) to an accep ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hydrolase
In biochemistry, hydrolases constitute a class of enzymes that commonly function as biochemical catalysts that use water to break a chemical bond: :\ce \quad \xrightarrowtext\quad \ce This typically results in dividing a larger molecule into smaller molecules. Some common examples of hydrolase enzymes are esterases including lipases, phosphatases, glycosidases, peptidases, and nucleosidases. Esterases cleave ester bonds in lipids and phosphatases cleave phosphate groups off molecules. An example of crucial esterase is acetylcholine esterase, which assists in transforming the neuron impulse into the acetate group after the hydrolase breaks the acetylcholine into choline and acetic acid. Acetic acid is an important metabolite in the body and a critical intermediate for other reactions such as glycolysis. Lipases hydrolyze glycerides. Glycosidases cleave sugar molecules off carbohydrates and peptidases hydrolyze peptide bonds. Nucleosidases hydrolyze the bonds of nucleo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzyme
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different molecules known as product (chemistry), products. Almost all metabolism, metabolic processes in the cell (biology), cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme, pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts include Ribozyme, catalytic RNA molecules, also called ribozymes. They are sometimes descr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Kinases
A protein kinase is a kinase which selectively modifies other proteins by covalently adding phosphates to them (phosphorylation) as opposed to kinases which modify lipids, carbohydrates, or other molecules. Phosphorylation usually results in a functional change of the target protein ( substrate) by changing enzyme activity, cellular location, or association with other proteins. The human genome contains about 500 protein kinase genes and they constitute about 2% of all human genes. There are two main types of protein kinase. The great majority are serine/threonine kinases, which phosphorylate the hydroxyl groups of serines and threonines in their targets. Most of the others are tyrosine kinases, although additional types exist. Protein kinases are also found in bacteria and plants. Up to 30% of all human proteins may be modified by kinase activity, and kinases are known to regulate the majority of cellular pathways, especially those involved in signal transduction. Chemical acti ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Adenosine Triphosphate
Adenosine triphosphate (ATP) is a nucleoside triphosphate that provides energy to drive and support many processes in living cell (biology), cells, such as muscle contraction, nerve impulse propagation, and chemical synthesis. Found in all known forms of life, it is often referred to as the "molecular unit of currency" for intracellular energy transfer. When consumed in a Metabolism, metabolic process, ATP converts either to adenosine diphosphate (ADP) or to adenosine monophosphate (AMP). Other processes regenerate ATP. It is also a Precursor (chemistry), precursor to DNA and RNA, and is used as a coenzyme. An average adult human processes around 50 kilograms (about 100 mole (unit), moles) daily. From the perspective of biochemistry, ATP is classified as a nucleoside triphosphate, which indicates that it consists of three components: a nitrogenous base (adenine), the sugar ribose, and the Polyphosphate, triphosphate. Structure ATP consists of three parts: a sugar, an amine base ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Active Site
In biology and biochemistry, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of amino acid residues that form temporary bonds with the substrate, the ''binding site'', and residues that catalyse a reaction of that substrate, the ''catalytic site''. Although the active site occupies only ~10–20% of the volume of an enzyme, it is the most important part as it directly catalyzes the chemical reaction. It usually consists of three to four amino acids, while other amino acids within the protein are required to maintain the tertiary structure of the enzymes. Each active site is evolved to be optimised to bind a particular substrate and catalyse a particular reaction, resulting in high specificity. This specificity is determined by the arrangement of amino acids within the active site and the structure of the substrates. Sometimes enzymes also need to bind with some cofactors to fulfil their functio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Structural Motif
In a chain-like biological molecule, such as a protein or nucleic acid, a structural motif is a common three-dimensional structure that appears in a variety of different, evolutionarily unrelated molecules. A structural motif does not have to be associated with a sequence motif; it can be represented by different and completely unrelated sequences in different proteins or RNA. In nucleic acids Depending upon the sequence and other conditions, nucleic acids can form a variety of structural motifs which is thought to have biological significance. ;Stem-loop: Stem-loop intramolecular base pairing is a pattern that can occur in single-stranded DNA or, more commonly, in RNA. The structure is also known as a hairpin or hairpin loop. It occurs when two regions of the same strand, usually complementary in nucleotide sequence when read in opposite directions, base-pair to form a double helix that ends in an unpaired loop. The resulting structure is a key building block of many ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |