|
Auxiliary Metabolic Genes
Auxiliary metabolic genes (AMGs) are found in many bacteriophages but originated in bacterial cells. AMGs modulate host cell metabolism during infection so that the phage can replicate more efficiently. For instance, bacteriophages that infect the abundant marine cyanobacteria ''Synechococcus'' and ''Prochlorococcus'' (cyanophages) carry AMGs that have been acquired from their immediate host as well as more distantly-related bacteria. Cyanophage AMGs support a variety of functions including photosynthesis, carbon metabolism, nucleic acid synthesis and metabolism. AMGs also have broader ecological impacts beyond their host including their influence on biogeochemical cycling. Classes AMGs employ diverse functions including pathways not involved in metabolism despite what the name suggests. They are categorized in two classes based on their presence in the Kyoto Encyclopedia of Genes and Genomes (KEGG). AMGs do not encompass metabolic genes involved in typical viral functions, such ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bacteriophage
A bacteriophage (), also known informally as a phage (), is a virus that infects and replicates within bacteria. The term is derived . Bacteriophages are composed of proteins that Capsid, encapsulate a DNA or RNA genome, and may have structures that are either simple or elaborate. Their genomes may encode as few as four genes (e.g. Bacteriophage MS2, MS2) and as many as hundreds of genes. Phages replicate within the bacterium following the injection of their genome into its cytoplasm. Bacteriophages are among the most common and diverse entities in the biosphere. Bacteriophages are ubiquitous viruses, found wherever bacteria exist. It is estimated there are more than 1031 bacteriophages on the planet, more than every other organism on Earth, including bacteria, combined. Viruses are the most abundant biological entity in the water column of the world's oceans, and the second largest component of biomass after prokaryotes, where up to 9x108 virus, virions per millilitre have b ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Horizontal Gene Transfer
Horizontal gene transfer (HGT) or lateral gene transfer (LGT) is the movement of genetic material between organisms other than by the ("vertical") transmission of DNA from parent to offspring (reproduction). HGT is an important factor in the evolution of many organisms. HGT is influencing scientific understanding of higher-order evolution while more significantly shifting perspectives on bacterial evolution. Horizontal gene transfer is the primary mechanism for the spread of antibiotic resistance in bacteria, and plays an important role in the evolution of bacteria that can degrade novel compounds such as human-created Bactericide, pesticides and in the evolution, maintenance, and transmission of virulence. It often involves Temperateness (virology), temperate bacteriophages and plasmids. Genes responsible for antibiotic resistance in one species of bacteria can be transferred to another species of bacteria through various mechanisms of HGT such as Transformation (genetics), tr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protein-coding genes and non-coding genes. During gene expression (the synthesis of Gene product, RNA or protein from a gene), DNA is first transcription (biology), copied into RNA. RNA can be non-coding RNA, directly functional or be the intermediate protein biosynthesis, template for the synthesis of a protein. The transmission of genes to an organism's offspring, is the basis of the inheritance of phenotypic traits from one generation to the next. These genes make up different DNA sequences, together called a genotype, that is specific to every given individual, within the gene pool of the population (biology), population of a given species. The genotype, along with environmental and developmental factors, ultimately determines the phenotype ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome
A genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences (see non-coding DNA), and often a substantial fraction of junk DNA with no evident function. Almost all eukaryotes have mitochondrial DNA, mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplast DNA, chloroplasts with a chloroplast genome. The study of the genome is called genomics. The genomes of many organisms have been Whole-genome sequencing, sequenced and various regions have been annotated. The first genome to be sequenced was that of the virus φX174 in 1977; the first genome sequence of a prokaryote (''Haemophilus influenzae'') was published in 1995; the yeast (''Saccharomyces cerevisiae'') genome was the first eukaryotic genome to be sequenced in 1996. The Human Genome Project ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
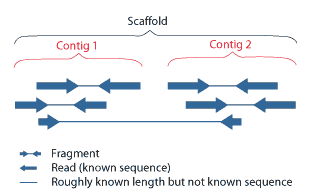
Contig
A contig (from ''contiguous'') is a set of overlapping DNA segments that together represent a consensus region of DNA.Gregory, S. ''Contig Assembly''. Encyclopedia of Life Sciences, 2005. In bottom-up sequencing projects, a contig refers to overlapping sequence data ( reads); in top-down sequencing projects, contig refers to the overlapping clones that form a physical map of the genome that is used to guide sequencing and assembly.Dear, P. H. ''Genome Mapping''. Encyclopedia of Life Sciences, 2005. . Contigs can thus refer both to overlapping DNA sequences and to overlapping physical segments (fragments) contained in clones depending on the context. Original definition of contig In 1980, Staden wrote: ''In order to make it easier to talk about our data gained by the shotgun method of sequencing we have invented the word "contig". A contig is a set of gel readings that are related to one another by overlap of their sequences. All gel readings belong to one and only one cont ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cyanophage
Cyanophages are viruses that infect cyanobacteria, also known as Cyanophyta or blue-green algae. Cyanobacteria are a phylum of bacteria that obtain their energy through the process of photosynthesis. Although cyanobacteria metabolize photoautotrophically like eukaryotic plants, they have Prokaryote, prokaryotic cell structure. Cyanophages can be found in both freshwater and marine environments. Marine and freshwater cyanophages have Regular icosahedron, icosahedral heads, which contain double-stranded DNA, attached to a tail by connector proteins. The size of the head and tail vary among species of cyanophages. Cyanophages infect a wide range of cyanobacteria and are key regulators of the cyanobacterial populations in aquatic environments, and may aid in the prevention of cyanobacterial blooms in freshwater and marine ecosystems. These blooms can pose a danger to humans and other animals, particularly in eutrophic freshwater lakes. Infection by these viruses is highly prevalent in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Membrane Vesicle Trafficking
Membrane vesicle trafficking in eukaryotic animal cells involves movement of biochemical signal molecules from synthesis-and-packaging locations in the Golgi body to specific release locations on the inside of the plasma membrane of the secretory cell. It takes place in the form of Golgi membrane-bound micro-sized vesicles, termed membrane vesicles (MVs). In this process, the packed cellular products are released or secreted outside the cell, across its membrane. On the other hand, the vesicular membrane is retained and recycled by the secretory cells. This phenomenon has a major role in synaptic neurotransmission, endocrine secretion, mucous secretion, granular-product secretion by neutrophils, and other phenomena. The scientists behind this discovery were awarded Nobel Prize for the year 2013. In prokaryotic, gram-negative bacterial cells, membrane vesicle trafficking is mediated through bacterial outer membrane bounded nano-sized vesicles, called outer membrane vesicles (OMVs). ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Transfer Agent
Gene transfer agents (GTAs) are DNA-containing virus-like particles that are produced by some bacteria and archaea and mediate horizontal gene transfer. Different GTA types have originated independently from viruses in several bacterial and archaeal lineages. These cells produce GTA particles containing short segments of the DNA present in the cell. After the particles are released from the producer cell, they can attach to related cells and inject their DNA into the cytoplasm. The DNA can then become part of the recipient cells' genome. GTAs are classified as viriforms in the ICTV taxonomy. Among the GTAs mentioned by the article, RcGTA and DsGTA are now in the family '' Rhodogtaviriformidae'', BaGTA in '' Bartogtaviriformidae'', and VSH-1 in '' Brachygtaviriformidae''. Dd1 and VTA do not yet have a classification. Discovery of gene transfer agents The first GTA system was discovered in 1974, when mixed cultures of ''Rhodobacter capsulatus'' strains produced a high frequenc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
National Center For Biotechnology Information
The National Center for Biotechnology Information (NCBI) is part of the National Library of Medicine (NLM), a branch of the National Institutes of Health (NIH). It is approved and funded by the government of the United States. The NCBI is located in Bethesda, Maryland, and was founded in 1988 through legislation sponsored by US Congressman Claude Pepper. The NCBI houses a series of databases relevant to biotechnology and biomedicine and is an important resource for bioinformatics tools and services. Major databases include GenBank for DNA sequences and PubMed, a bibliographic database for biomedical literature. Other databases include the NCBI Epigenomics database. All these databases are available online through the Entrez search engine. NCBI was directed by David Lipman, one of the original authors of the BLAST sequence alignment program and a widely respected figure in bioinformatics. GenBank NCBI had responsibility for making available the GenBank DNA seque ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MEROPS
MEROPS is an online database for peptidases (also known as proteases, proteinases and proteolytic enzymes) and their inhibitors. The classification scheme for peptidases was published by Rawlings & Barrett in 1993, and that for protein inhibitors by Rawlings ''et al.'' in 2004.Rawlings, N.D., Tolle, D.P. & Barrett, A.J. (2004) "Evolutionary families of peptidase inhibitors." ''Biochem J'' 378, 705-716. The most recent version, MEROPS 12.5, was released in September 2023. Overview The classification is based on similarities at the tertiary and primary structural levels. Comparisons are restricted to that part of the sequence directly involved in the reaction, which in the case of a peptidase must include the active site, and for a protein inhibitor the reactive site. The classification is hierarchical: sequences are assembled into families, and families are assembled into clans. Each peptidase, family, and clan has a unique identifier. Classification Family The families of pe ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
CAZy
CAZy is a database of Carbohydrate-Active enZYmes (CAZymes). The database contains a classification and associated information about enzymes involved in the synthesis, metabolism, and recognition of complex carbohydrates, i.e. disaccharides, oligosaccharides, polysaccharides, and glycoconjugates. Included in the database are families of glycoside hydrolases, glycosyltransferases, polysaccharide lyases, carbohydrate esterases, and non-catalytic carbohydrate-binding modules. The CAZy database also includes a classification of Auxiliary Activity redox enzymes involved in the breakdown of lignocellulose. CAZy was established in 1999 in order to provide online and constantly updated access to the protein sequence-based family classification of CAZymes, which was originally developed in early 1990s to classify the glycoside hydrolases. New entries are added shortly after they appear in the daily releases of GenBank. The rapid evolution of high-throughput DNA sequencing has resulted ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |



.png)
