|
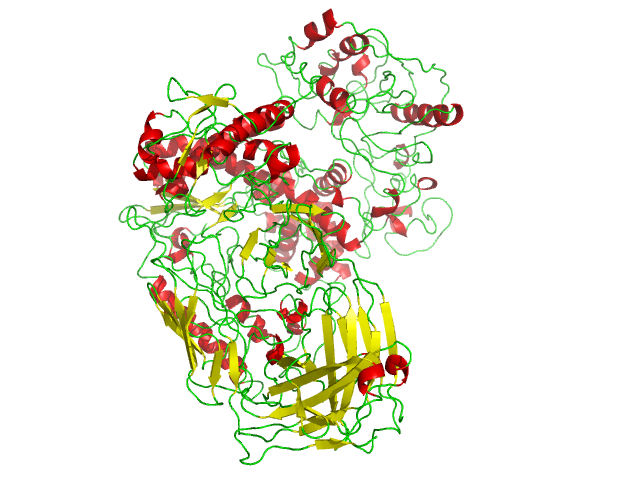
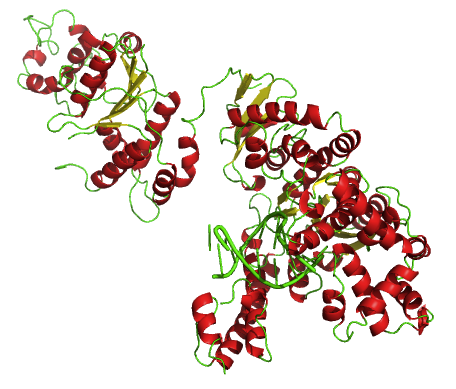
Taq Polymerase
''Taq'' polymerase is a thermostable DNA polymerase I named after the thermophilic eubacterial microorganism ''Thermus aquaticus,'' from which it was originally isolated by master's student Alice Chien et al. in 1976. Its name is often abbreviated to ''Taq'' or ''Taq'' pol. It is frequently used in the polymerase chain reaction (PCR), a method for greatly amplifying the quantity of short segments of DNA. ''T. aquaticus'' is a bacterium that lives in hot springs and hydrothermal vents, and ''Taq'' polymerase was identified as an enzyme able to withstand the protein-denaturing conditions (high temperature) required during PCR. Therefore, it replaced the DNA polymerase from '' E. coli'' originally used in PCR. Enzymatic properties ''Taqs optimum temperature for activity is 75–80 °C, with a half-life of greater than 2 hours at 92.5 °C, 40 minutes at 95 °C and 9 minutes at 97.5 °C, and can replicate a 1000 base pair strand of DNA in less than 10 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Thermostable DNA Polymerase
Thermostable DNA polymerases are DNA polymerases that originate from thermophiles, usually bacterial or archaeal species, and are therefore thermostable. They are used for the polymerase chain reaction and related methods for the amplification and modification of DNA. Properties Several DNA polymerases have been described with distinct properties that define their specific utilisation in a PCR, in real-time PCR or in an isothermal amplification. Being DNA polymerases, the thermostable DNA polymerases all have a 5'→3' polymerase activity, and either a 5'→3' or a 3'→5' exonuclease activity. Structure DNA polymerases are roughly shaped like a hand with a thumb, palm and fingers.T. A. Steitz''DNA polymerases: structural diversity and common mechanisms.''In: '' Journal of Biological Chemistry.'' Volume 274, issue 25, June 1999, p. 17395–17398, , .P. J. Rothwell, G. Waksman: ''Structure and mechanism of DNA polymerases.'' In: ''Advances in protein chemistry.'' Volu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Magnesium
Magnesium is a chemical element; it has Symbol (chemistry), symbol Mg and atomic number 12. It is a shiny gray metal having a low density, low melting point and high chemical reactivity. Like the other alkaline earth metals (group 2 of the periodic table), it occurs naturally only in combination with other elements and almost always has an oxidation state of +2. It reacts readily with air to form a thin Passivation (chemistry), passivation coating of magnesium oxide that inhibits further corrosion of the metal. The free metal burns with a brilliant-white light. The metal is obtained mainly by electrolysis of magnesium Salt (chemistry), salts obtained from brine. It is less dense than aluminium and is used primarily as a component in strong and lightweight magnesium alloy, alloys that contain aluminium. In the cosmos, magnesium is produced in large, aging stars by the sequential addition of three Helium nucleus, helium nuclei to a carbon nucleus. When such stars explo ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Kary Mullis
Kary Banks Mullis (December 28, 1944August 7, 2019) was an American biochemist. In recognition of his role in the invention of the polymerase chain reaction (PCR) technique, he shared the 1993 Nobel Prize in Chemistry with Michael Smith and was awarded the Japan Prize in the same year. PCR became a central technique in biochemistry and molecular biology, described by ''The New York Times'' as "highly original and significant, virtually dividing biology into the two epochs of before PCR and after PCR." Mullis downplayed humans' role in climate change, expressed doubt that HIV is the cause of AIDS, and professed a belief in astrology and the paranormal. He also practiced clandestine chemistry by producing LSD. Mullis's unscientific statements about topics outside his area of expertise have been named by '' Skeptical Inquirer'' as an instance of " Nobel disease". Early life and education Mullis was born in Lenoir, North Carolina, near the Blue Ridge Mountains, on December 2 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Ligase
DNA ligase is a type of enzyme that facilitates the joining of DNA strands together by catalyzing the formation of a phosphodiester bond. It plays a role in repairing single-strand breaks in duplex DNA in living organisms, but some forms (such as DNA ligase IV) may specifically repair double-strand breaks (i.e. a break in both complementary strands of DNA). Single-strand breaks are repaired by DNA ligase using the complementary strand of the double helix as a template, with DNA ligase creating the final phosphodiester bond to fully repair the DNA. DNA ligase is used in both DNA repair and DNA replication (see '' Mammalian ligases''). In addition, DNA ligase has extensive use in molecular biology laboratories for recombinant DNA experiments (see '' Research applications''). Purified DNA ligase is used in gene cloning to join DNA molecules together to form recombinant DNA. Enzymatic mechanism The mechanism of DNA ligase is to form two covalent phosphodiester bonds between ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Thymine
Thymine () (symbol T or Thy) is one of the four nucleotide bases in the nucleic acid of DNA that are represented by the letters G–C–A–T. The others are adenine, guanine, and cytosine. Thymine is also known as 5-methyluracil, a pyrimidine nucleobase. In RNA, thymine is replaced by the nucleobase uracil. Thymine was first isolated in 1893 by Albrecht Kossel and Albert Neumann from calf thymus glands, hence its name. Derivation As its alternate name (5-methyluracil) suggests, thymine may be derived by methylation of uracil at the 5th carbon. In RNA, thymine is replaced with uracil in most cases. In DNA, thymine (T) binds to adenine (A) via two hydrogen bonds, thereby stabilizing the nucleic acid structures. Thymine combined with deoxyribose creates the nucleoside deoxythymidine, which is synonymous with the term thymidine. Thymidine can be phosphorylated with up to three phosphoric acid groups, producing dTMP (deoxythymidine monophosphate), dTDP, or dTTP (for the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria and archaea; however plasmids are sometimes present in and eukaryotic organisms as well. Plasmids often carry useful genes, such as those involved in antibiotic resistance, virulence, secondary metabolism and bioremediation. While chromosomes are large and contain all the essential genetic information for living under normal conditions, plasmids are usually very small and contain additional genes for special circumstances. Artificial plasmids are widely used as vectors in molecular cloning, serving to drive the replication of recombinant DNA sequences within host organisms. In the laboratory, plasmids may be introduced into a cell via transformation. Synthetic plasmids are available for procurement over the internet by various vendors ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cloning Vector
A cloning vector is a small piece of DNA that can be stably maintained in an organism, and into which a foreign DNA fragment can be inserted for cloning purposes. The cloning vector may be DNA taken from a virus, the cell of a higher organism, or it may be the plasmid of a bacterium. The vector contains features that allow for the convenient insertion of a DNA fragment into the vector or its removal from the vector, for example through the presence of restriction sites. The vector and the foreign DNA may be treated with a restriction enzyme that cuts the DNA, and DNA fragments thus generated contain either blunt ends or overhangs known as sticky ends, and vector DNA and foreign DNA with compatible ends can then be joined by molecular ligation. After a DNA fragment has been cloned into a cloning vector, it may be further subcloned into another vector designed for more specific use. There are many types of cloning vectors, but the most commonly used ones are genetically engineer ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
TA Cloning
TA cloning (also known as rapid cloning or T cloning) is a subcloning technique that avoids the use of restriction enzymes and is easier and quicker than traditional subcloning. The technique relies on the ability of adenine (A) and thymine (T) (complementary basepairs) on different DNA fragments to hybridize and, in the presence of ligase, become ligated together. PCR products are usually amplified using Taq DNA polymerase which preferentially adds an adenine to the 3' end of the product. Such PCR amplified inserts are cloned into linearized vectors that have complementary 3' thymine overhangs. Procedure Creating the insert The insert is created by PCR using Taq polymerase. This polymerase lacks 3' to 5' proofreading activity and, with a high probability, adds a single, 3'-adenine overhang to each end of the PCR product. It is best if the PCR primers have guanines at the 5' end as this maximizes probability of Taq DNA polymerase adding the terminal adenosine overhang. Thermost ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Adenine
Adenine (, ) (nucleoside#List of nucleosides and corresponding nucleobases, symbol A or Ade) is a purine nucleotide base that is found in DNA, RNA, and Adenosine triphosphate, ATP. Usually a white crystalline subtance. The shape of adenine is complementary and pairs to either thymine in DNA or uracil in RNA. In cells adenine, as an independent molecule, is rare. It is almost always covalent bond, covalently bound to become a part of a larger biomolecule. Adenine has a central role in cellular respiration. It is part of adenosine triphosphate which provides the energy that drives and supports most activities in living cell (biology), cells, such as Protein biosynthesis, protein synthesis, chemical synthesis, muscle contraction, and nerve impulse propagation. In respiration it also participates as part of the cofactor (biochemistry), cofactors nicotinamide adenine dinucleotide, flavin adenine dinucleotide, and Coenzyme A. It is also part of adenosine, adenosine monophosphate, cy ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Pfu DNA Polymerase
''Pfu'' DNA polymerase is an enzyme found in the hyperthermophilic archaeon '' Pyrococcus furiosus'', where it functions to copy the organism's DNA during cell division (thermostable DNA polymerase). In the laboratory setting, ''Pfu'' is used to amplify DNA in the polymerase chain reaction (PCR), where the enzyme serves the central function of copying a new strand of DNA during each extension step. It is a family B DNA polymerase. It has an RNase H-like 3'-5' exonuclease domain, typical of B-family polymerase such as DNA polymerase II. Proofreading ability of ''Pfu'' polymerase ''Pfu'' DNA polymerase has superior thermostability and proofreading properties compared with ''Taq'' DNA polymerase. Unlike ''Taq'' DNA polymerase, ''Pfu'' DNA polymerase possesses 3' to 5' exonuclease proofreading activity, meaning that as the DNA is assembled from the 5' end to 3' end, the exonuclease activity immediately removes nucleotides misincorporated at the 3' end of the growing DNA strand. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exonuclease
Exonucleases are enzymes that work by cleaving nucleotides one at a time from the end (exo) of a polynucleotide chain. A hydrolyzing reaction that breaks phosphodiester bonds at either the 3′ or the 5′ end occurs. Its close relative is the endonuclease, which cleaves phosphodiester bonds in the middle (endo) of a polynucleotide chain. Eukaryotes and prokaryotes have three types of exonucleases involved in the normal turnover of mRNA: 5′ to 3′ exonuclease (Xrn1), which is a dependent decapping protein; 3′ to 5′ exonuclease, an independent protein; and poly(A)-specific 3′ to 5′ exonuclease. In both archaea and eukaryotes, one of the main routes of RNA degradation is performed by the multi-protein exosome complex, which consists largely of 3′ to 5′ exoribonucleases. Significance to polymerase RNA polymerase II is known to be in effect during transcriptional termination; it works with a 5' exonuclease (human gene Xrn2) to degrade the newly formed transcr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |