|
RAD17
Cell cycle checkpoint protein RAD17 is a protein that in humans is encoded by the ''RAD17'' gene. Function The protein encoded by this gene is highly similar to the gene product of Schizosaccharomyces pombe rad17, a cell cycle checkpoint gene required for cell cycle arrest and DNA damage repair in response to DNA damage. This protein shares strong similarity with DNA replication factor C (RFC), and can form a complex with RFCs. This protein binds to chromatin prior to DNA damage and is phosphorylated by ATR after the damage. This protein recruits the RAD1-RAD9-HUS1 checkpoint protein complex onto chromatin after DNA damage, which may be required for its phosphorylation. The phosphorylation of this protein is required for the DNA-damage-induced cell cycle G2 arrest, and is thought to be a critical early event during checkpoint signaling in DNA-damaged cells. Eight alternatively spliced transcript variants of this gene, which encode four distinct proteins, have been reported. Me ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RAD1 Homolog
Cell cycle checkpoint protein RAD1 is a protein that in humans is encoded by the ''RAD1'' gene. Function This gene encodes a component of a heterotrimeric cell cycle checkpoint complex, known as the 9-1-1 complex, that is activated to stop cell cycle progression in response to DNA damage or incomplete DNA replication. The 9-1-1 complex is recruited by RAD17 to affected sites where it may attract specialized DNA polymerases and other DNA repair effectors. Alternatively spliced transcript variants encoding different isoforms of this gene have been described. Meiosis During meiosis, double-strand breaks occur in DNA that initiate homologous recombination, recombination. Recombination is a process that repairs the breaks and also promotes faithful chromosome segregation. In yeast the 9-1-1 complex (including RAD1) facilitates meiotic recombination. An alternative, but inaccurate, mechanism for repairing double-strand breaks is non-homologous end joining. In the rice plant, the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
HUS1
Checkpoint protein HUS1 is a protein that in humans is encoded by the ''HUS1'' gene. Function The protein encoded by this gene is a component of an evolutionarily conserved, genotoxin-activated checkpoint complex that is involved in the cell cycle arrest in response to DNA damage. This protein forms a heterotrimeric complex with checkpoint proteins RAD9 and RAD1. In response to DNA damage, the trimeric complex interacts with another protein complex consisting of checkpoint protein RAD17 and four small subunits of the replication factor C (RFC), which loads the combined complex onto the chromatin. The DNA damage induced chromatin binding has been shown to depend on the activation of the checkpoint kinase ATM, and is thought to be an early checkpoint signaling event. In somatic cells In somatic cells the RAD9-RAD1-HUS1 (9-1-1) complex responds to DNA damage by promoting DNA repair. In meiosis In flies, worms and yeast, the 9-1-1 complex is necessary for meiotic checkpoint fu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ataxia Telangiectasia And Rad3 Related
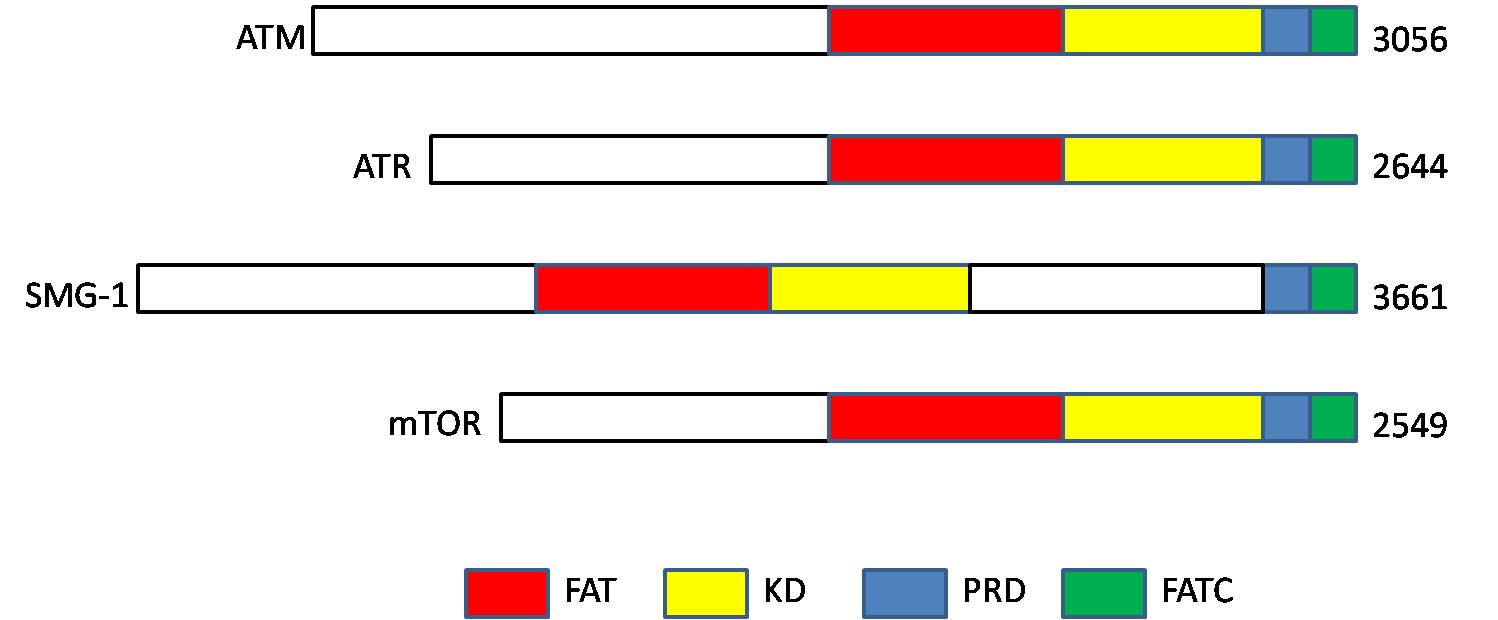
Serine/threonine-protein kinase ATR, also known as ataxia telangiectasia and Rad3-related protein (ATR) or FRAP-related protein 1 (FRP1), is an enzyme that, in humans, is encoded by the ''ATR'' gene. It is a large kinase of about 301.66 kDa. ATR belongs to the phosphatidylinositol 3-kinase-related kinase protein family. ATR is activated in response to single strand breaks, and works with ATM to ensure genome integrity. Function ATR is a serine/threonine-specific protein kinase that is involved in sensing DNA damage and activating the DNA damage checkpoint, leading to cell cycle arrest in eukaryotes. ATR is activated in response to persistent single-stranded DNA, which is a common intermediate formed during DNA damage detection and repair. Single-stranded DNA occurs at stalled replication forks and as an intermediate in DNA repair pathways such as nucleotide excision repair and homologous recombination repair. ATR is activated during more persistent issues with DNA damage ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
NHP2L1
NHP2-like protein 1 is a protein that in humans is encoded by the ''SNU13'' gene. Function Originally named because of its sequence similarity to the ''Saccharomyces cerevisiae'' NHP2 (non-histone protein 2), this protein appears to be a highly conserved nuclear protein that is a component of the 4/U6.U5 tri-snRNP. It binds to the 5' stem-loop of U4 snRNA. Two transcript variants encoding the same protein have been found for this gene. Interactions SNU13 has been shown to interact with RAD17 Cell cycle checkpoint protein RAD17 is a protein that in humans is encoded by the ''RAD17'' gene. Function The protein encoded by this gene is highly similar to the gene product of Schizosaccharomyces pombe rad17, a cell cycle checkpoint gene .... References Further reading * * * * * * * * * * * * {{protein-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ataxia Telangiectasia Mutated
ATM serine/threonine kinase or Ataxia-telangiectasia mutated, symbol ATM, is a serine/threonine protein kinase that is recruited and activated by DNA repair#Double-strand breaks, DNA double-strand breaks (Canonical pathway, canonical pathway), oxidative stress, topoisomerase cleavage complexes, splicing intermediates, R-loops and in some cases by DNA repair, single-strand DNA breaks. It phosphorylates several key proteins that initiate activation of the DNA damage cell cycle checkpoint, checkpoint, leading to cell cycle arrest, DNA repair or apoptosis. Several of these targets, including p53, CHK2, BRCA1, NBS1 and H2AX are tumor suppressors. In 1995, the gene was discovered by Yosef Shiloh who named its product ATM since he found that its mutations are responsible for the disorder ataxia–telangiectasia#Cause, ataxia–telangiectasia. In 1998, the Shiloh and Michael B. Kastan, Kastan laboratories independently showed that ATM is a protein kinase whose activity is enhanced by DNA ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Synaptonemal Complex
The synaptonemal complex (SC) is a protein structure that forms between homologous chromosomes (two pairs of sister chromatids) during meiosis and is thought to mediate synapsis and recombination during prophase I during meiosis in eukaryotes. It is currently thought that the SC functions primarily as a scaffold to allow interacting chromatids to complete their crossover activities. Composition The synaptonemal complex is a tripartite structure consisting of two parallel lateral regions and a central element. This "tripartite structure" is seen during the pachytene stage of the first meiotic prophase, both in males and in females during gametogenesis. Previous to the pachytene stage, during leptonema, the lateral elements begin to form and they initiate and complete their pairing during the zygotene stage. After pachynema ends, the SC usually becomes disassembled and can no longer be identified. In humans, three specific components of the synaptonemal complex have been ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
POLE (enzyme)
DNA polymerase epsilon is a member of the DNA polymerase family of enzymes found in eukaryotes. It is composed of the following four subunits: POLE (central catalytic unit), POLE2 (subunit 2), POLE3 (subunit 3), and POLE4 (subunit 4). Recent evidence suggests that it plays a major role in leading strand DNA synthesis and nucleotide and base excision repair. Research had conducted to study nucleotide excision repair DNA synthesis by DNA polymerase epsilon in the presence of PCNA (proliferating cell nuclear antigen), RFC (replication factor C) and RPA (replication protein A). Either DNA polymerase epsilon or DNA polymerase delta along with DNA ligase DNA ligase is a type of enzyme that facilitates the joining of DNA strands together by catalyzing the formation of a phosphodiester bond. It plays a role in repairing single-strand breaks in duplex DNA in living organisms, but some forms (such ... can be used to repair UV-damaged DNA. However, it is found that DNA polymeras ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme catalysis, catalysing metabolic reactions, DNA replication, Cell signaling, responding to stimuli, providing Cytoskeleton, structure to cells and Fibrous protein, organisms, and Intracellular transport, transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the Nucleic acid sequence, nucleotide sequence of their genes, and which usually results in protein folding into a specific Protein structure, 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called pep ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protein-coding genes and non-coding genes. During gene expression (the synthesis of Gene product, RNA or protein from a gene), DNA is first transcription (biology), copied into RNA. RNA can be non-coding RNA, directly functional or be the intermediate protein biosynthesis, template for the synthesis of a protein. The transmission of genes to an organism's offspring, is the basis of the inheritance of phenotypic traits from one generation to the next. These genes make up different DNA sequences, together called a genotype, that is specific to every given individual, within the gene pool of the population (biology), population of a given species. The genotype, along with environmental and developmental factors, ultimately determines the phenotype ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RAD51
DNA repair protein RAD51 homolog 1 is a protein encoded by the gene ''RAD51''. The enzyme encoded by this gene is a member of the RAD51 protein family which assists in repair of DNA double strand breaks. RAD51 family members are homologous to the bacterial RecA, Archaeal RadA, and yeast Rad51. The protein is highly conserved in most eukaryotes, from yeast to humans. The name RAD51 derives from ''radiation sensitive protein 51''. Variants Two alternatively spliced transcript variants of this gene have been reported, which encode distinct proteins. Transcript variants utilizing alternative polyA signals also exist. Family In mammals, seven recA-like genes have been identified: Rad51, Rad51L1/B, Rad51L2/C, Rad51L3/D, XRCC2, XRCC3, and DMC1/Lim15. All of these proteins, with the exception of meiosis-specific DMC1, are essential for development in mammals. Rad51 is a member of thRecA-like NTPases Function In humans, RAD51 is a 339-amino acid protein that plays ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Homologous Recombination
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids (usually DNA as in Cell (biology), cellular organisms but may be also RNA in viruses). Homologous recombination is widely used by cells to accurately DNA repair, repair harmful DNA breaks that occur on both strands of DNA, known as double-strand breaks (DSB), in a process called homologous recombinational repair (HRR). Homologous recombination also produces new combinations of DNA sequences during meiosis, the process by which eukaryotes make gamete cells, like sperm and ovum, egg cells in animals. These new combinations of DNA represent genetic variation in offspring, which in turn enables populations to Adaptation, adapt during the course of evolution. Homologous recombination is also used in horizontal gene transfer to exchange genetic material between different strains and species ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |