|
Nicotinamide Ribonucleoside Uptake Transporters
The Nicotinamide Ribonucleoside (NR) Uptake Permease (PnuC) FamilyTC# 4.B.1 is a family of transmembrane transporters that is part of the TOG superfamily. Close PnuC homologues are found in a wide range of Gram-negative and Gram-positive bacteria, archaea and eukaryotes. Function PnuC of ''Salmonella typhimurium'' and ''Haemophilus influenzae'' are believed to function cooperatively with NadR homologues, multifunctional proteins that together with PnuC, participate in NR phosphorylation, transport and transcriptional regulation. NadR, a cytoplasmic protein that is partly membrane associated, contains one well conserved and one poorly conserved mononucleotide-binding consensus sequence (G-X4 GKS). It drives transport and may render transport responsive to internal pyridine nucleotide levels. While its N-terminal half functions as a repressor, its C-terminal half functions as an NR kinase in a putative group translocation process. PnuC of ''Haemophilus influenzae'' The ''H. influ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
TOG Superfamily
The transporter-opsin-G protein-coupled receptor (TOG) superfamily is a protein superfamily of integral membrane proteins, usually of 7 or 8 transmembrane alpha-helical segments (TMSs). It includes (1) ion-translocating microbial rhodopsins and (2) G protein-coupled receptors (GPCRs), (3) Sweet sugar transporters, (4) nicotinamide ribonucleoside uptake permeases (PnuCTC# 4.B.1, (5) 4-toluene sulfonate uptake permeases (TSUP)TC# 2.A.102, (6) Ni2+–Co2+ transporters (NiCoT)TC# 2.A.52, (7) organic solute transporters (OST)TC# 2.A.82, (8) phosphate:Na+ symporters (PNaS)TC# 2.A.58 and (9) lysosomal cystine transporters (LCT)TC# 2.A.43. Families Currently recognized families Family (from la, familia) is a group of people related either by consanguinity (by recognized birth) or affinity (by marriage or other relationship). The purpose of the family is to maintain the well-being of its members and of society. Idea ... within the TOG Superfamily (with TC numbers in blue) inc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Proton Motive Force
Chemiosmosis is the movement of ions across a semipermeable membrane bound structure, down their electrochemical gradient. An important example is the formation of adenosine triphosphate (ATP) by the movement of hydrogen ions (H+) across a membrane during cellular respiration or photosynthesis. Hydrogen ions, or protons, will diffuse from a region of high proton concentration to a region of lower proton concentration, and an electrochemical concentration gradient of protons across a membrane can be harnessed to make ATP. This process is related to osmosis, the movement of water across a selective membrane, which is why it is called "chemiosmosis". ATP synthase is the enzyme that makes ATP by chemiosmosis. It allows protons to pass through the membrane and uses the free energy difference to phosphorylate adenosine diphosphate (ADP), making ATP. The generation of ATP by chemiosmosis occurs in mitochondria and chloroplasts, as well as in most bacteria and archaea. For in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transmembrane Transporters
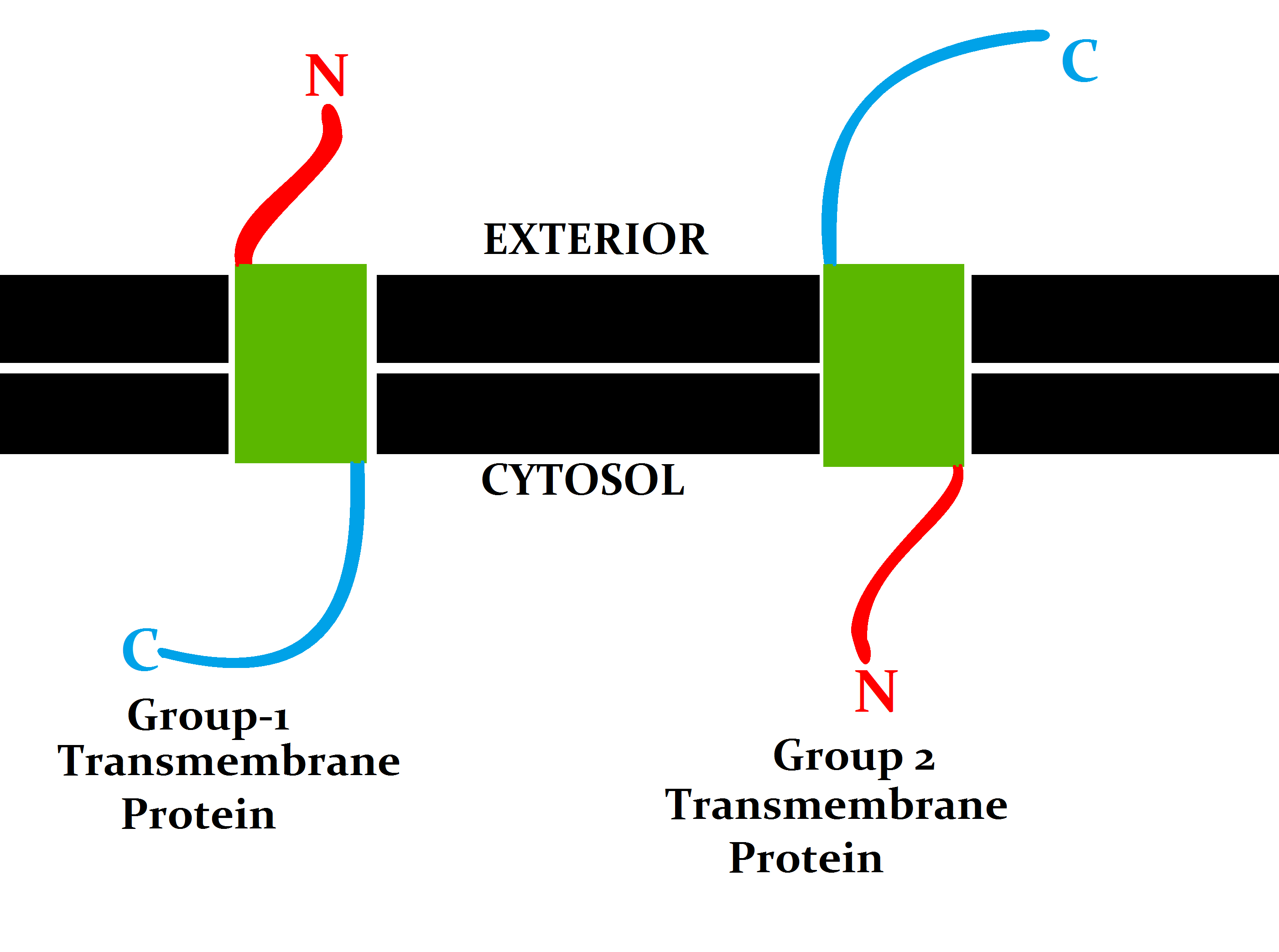
A transmembrane protein (TP) is a type of integral membrane protein that spans the entirety of the cell membrane. Many transmembrane proteins function as gateways to permit the transport of specific substances across the membrane. They frequently undergo significant conformational changes to move a substance through the membrane. They are usually highly hydrophobic and aggregate and precipitate in water. They require detergents or nonpolar solvents for extraction, although some of them (beta-barrels) can be also extracted using denaturing agents. The peptide sequence that spans the membrane, or the transmembrane segment, is largely hydrophobic and can be visualized using the hydropathy plot. Depending on the number of transmembrane segments, transmembrane proteins can be classified as single-span (or bitopic) or multi-span (polytopic). Some other integral membrane proteins are called monotopic, meaning that they are also permanently attached to the membrane, but do not pas ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Membrane Proteins
Membrane proteins are common proteins that are part of, or interact with, biological membranes. Membrane proteins fall into several broad categories depending on their location. Integral membrane proteins are a permanent part of a cell membrane and can either penetrate the membrane ( transmembrane) or associate with one or the other side of a membrane ( integral monotopic). Peripheral membrane proteins are transiently associated with the cell membrane. Membrane proteins are common, and medically important—about a third of all human proteins are membrane proteins, and these are targets for more than half of all drugs. Nonetheless, compared to other classes of proteins, determining membrane protein structures remains a challenge in large part due to the difficulty in establishing experimental conditions that can preserve the correct conformation of the protein in isolation from its native environment. Function Membrane proteins perform a variety of functions vital to the su ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Families
A protein family is a group of evolutionarily related proteins. In many cases, a protein family has a corresponding gene family, in which each gene encodes a corresponding protein with a 1:1 relationship. The term "protein family" should not be confused with family as it is used in taxonomy. Proteins in a family descend from a common ancestor and typically have similar three-dimensional structures, functions, and significant sequence similarity. The most important of these is sequence similarity (usually amino-acid sequence), since it is the strictest indicator of homology and therefore the clearest indicator of common ancestry. A fairly well developed framework exists for evaluating the significance of similarity between a group of sequences using sequence alignment methods. Proteins that do not share a common ancestor are very unlikely to show statistically significant sequence similarity, making sequence alignment a powerful tool for identifying the members of protein famil ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Rossmann Fold
The Rossmann fold is a tertiary fold found in proteins that bind nucleotides, such as enzyme cofactors FAD, NAD+, and NADP+. This fold is composed of alternating beta strands and alpha helical segments where the beta strands are hydrogen bonded to each other forming an extended beta sheet and the alpha helices surround both faces of the sheet to produce a three-layered sandwich. The classical Rossmann fold contains six beta strands whereas Rossmann-like folds, sometimes referred to as Rossmannoid folds, contain only five strands. The initial beta-alpha-beta (bab) fold is the most conserved segment of the Rossmann fold. The motif is named after Michael Rossmann who first noticed this structural motif in the enzyme lactate dehydrogenase in 1970 and who later observed that this was a frequently occurring motif in nucleotide binding proteins. Rossmann and Rossmannoid fold proteins are extremely common. They make up 20% of proteins with known structures in the Protein Data Bank, a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Helix-turn-helix
Helix-turn-helix is a DNA-binding protein (DBP). The helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two α helices, joined by a short strand of amino acids, that bind to the major groove of DNA. The HTH motif occurs in many proteins that regulate gene expression. It should not be confused with the helix–loop–helix motif. Discovery The discovery of the helix-turn-helix motif was based on similarities between several genes encoding transcription regulatory proteins from bacteriophage lambda and ''Escherichia coli'': Cro, CAP, and λ repressor, which were found to share a common 20–25 amino acid sequence that facilitates DNA recognition. Function The helix-turn-helix motif is a DNA-binding motif. The recognition and binding to DNA by helix-turn-helix proteins is done by the two α helices, one occupying the N-terminal end of the motif, the other at the C-terminus. In most cases, such as in the Cro repressor, the s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
C-terminus
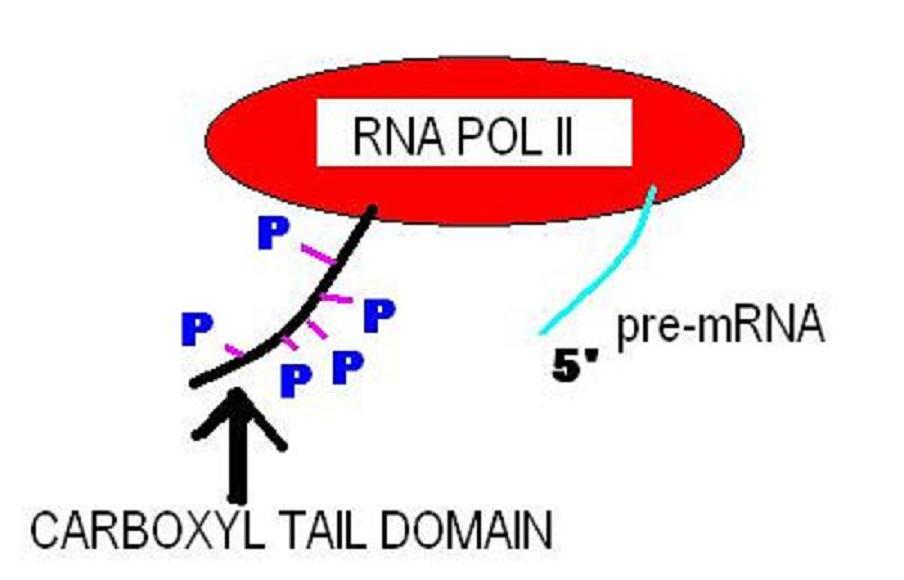
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When the protein is translated from messenger RNA, it is created from N-terminus to C-terminus. The convention for writing peptide sequences is to put the C-terminal end on the right and write the sequence from N- to C-terminus. Chemistry Each amino acid has a carboxyl group and an amine group. Amino acids link to one another to form a chain by a dehydration reaction which joins the amine group of one amino acid to the carboxyl group of the next. Thus polypeptide chains have an end with an unbound carboxyl group, the C-terminus, and an end with an unbound amine group, the N-terminus. Proteins are naturally synthesized starting from the N-terminus and ending at the C-terminus. Function C-terminal retention signals While the N-terminus of a protein often co ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alpha Helix
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located four residues earlier along the protein sequence. The alpha helix is also called a classic Pauling–Corey–Branson α-helix. The name 3.613-helix is also used for this type of helix, denoting the average number of residues per helical turn, with 13 atoms being involved in the ring formed by the hydrogen bond. Among types of local structure in proteins, the α-helix is the most extreme and the most predictable from sequence, as well as the most prevalent. Discovery In the early 1930s, William Astbury showed that there were drastic changes in the X-ray fiber diffraction of moist wool or hair fibers upon significant stretching. The data suggested that the unstretched fibers had a coiled molecular structure with a characteristic repeat of ≈. Astbu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transmembrane Domain
A transmembrane domain (TMD) is a membrane-spanning protein domain. TMDs generally adopt an alpha helix topological conformation, although some TMDs such as those in porins can adopt a different conformation. Because the interior of the lipid bilayer is hydrophobic, the amino acid residues in TMDs are often hydrophobic, although proteins such as membrane pumps and ion channels can contain polar residues. TMDs vary greatly in length, sequence, and hydrophobicity, adopting organelle-specific properties. Functions of transmembrane domains Transmembrane domains are known to perform a variety of functions. These include: * Anchoring transmembrane proteins to the membrane. *Facilitating molecular transport of molecules such as ions and proteins across biological membranes; usually hydrophilic residues and binding sites in the TMDs help in this process. *Signal transduction across the membrane; many transmembrane proteins, such as G protein-coupled receptors, receive extracel ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Integral Membrane Protein
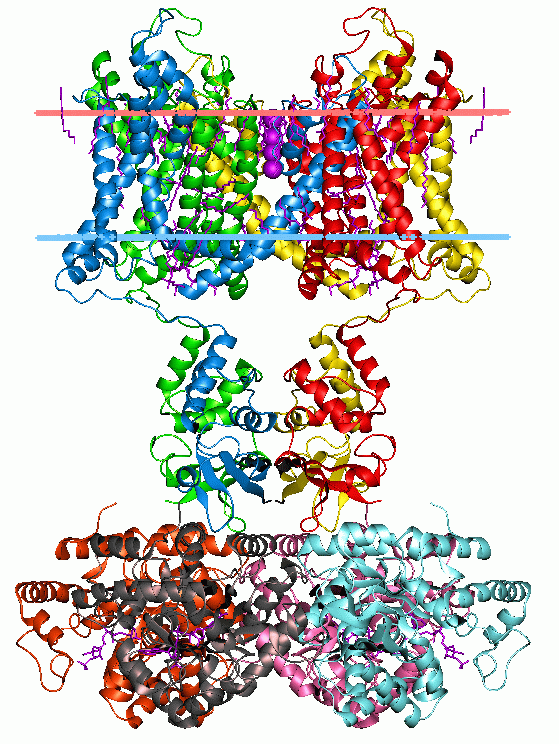
An integral, or intrinsic, membrane protein (IMP) is a type of membrane protein that is permanently attached to the biological membrane. All ''transmembrane proteins'' are IMPs, but not all IMPs are transmembrane proteins. IMPs comprise a significant fraction of the proteins encoded in an organism's genome. Proteins that cross the membrane are surrounded by annular lipids, which are defined as lipids that are in direct contact with a membrane protein. Such proteins can only be separated from the membranes by using detergents, nonpolar solvents, or sometimes denaturing agents. Structure Three-dimensional structures of ~160 different integral membrane proteins have been determined at atomic resolution by X-ray crystallography or nuclear magnetic resonance spectroscopy. They are challenging subjects for study owing to the difficulties associated with extraction and crystallization. In addition, structures of many water-soluble protein domains of IMPs are available in the Prote ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |

.png)