|
Metagenomic
Metagenomics is the study of all genetic material from all organisms in a particular environment, providing insights into their composition, diversity, and functional potential. Metagenomics has allowed researchers to profile the microbial composition of environmental and clinical samples without the need for time-consuming culture of individual species. Metagenomics has transformed microbial ecology and evolutionary biology by uncovering previously hidden biodiversity and metabolic capabilities. As the cost of DNA sequencing continues to decline, metagenomic studies now routinely profile hundreds to thousands of samples, enabling large-scale exploration of microbial communities and their roles in health and global ecosystems. Metagenomic studies most commonly employ shotgun sequencing though long-read sequencing is being increasingly utilised as technologies advance. The field is also referred to as environmental genomics, ecogenomics, community genomics, or microbiomics a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Third-generation Sequencing
Third-generation sequencing (also known as long-read sequencing) is a class of DNA sequencing methods that have the capability to produce substantially longer reads (ranging from 10 kb to >1 Mb in length) than second generation sequencing, also known as next-generation sequencing methods. These methods emerged in 2008, characterized by technologies such as nanopore sequencing or single-molecule real-time sequencing, and continue to be developed. The ability to sequence longer reads has critical implications for both genome science and the study of biology in general. In structural variant calling, third generation sequencing has been found to outperform existing methods, even at a low depth of sequencing coverage. However, third generation sequencing data have much higher error rates than previous technologies, which can complicate downstream genome assembly and analysis of the resulting data. These technologies are undergoing active development and it is expected that there will ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
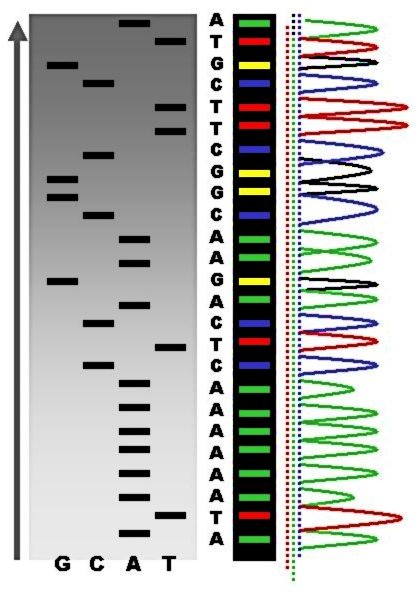
DNA Sequencing
DNA sequencing is the process of determining the nucleic acid sequence – the order of nucleotides in DNA. It includes any method or technology that is used to determine the order of the four bases: adenine, thymine, cytosine, and guanine. The advent of rapid DNA sequencing methods has greatly accelerated biological and medical research and discovery. Knowledge of DNA sequences has become indispensable for basic biological research, Genographic Project, DNA Genographic Projects and in numerous applied fields such as medical diagnosis, biotechnology, forensic biology, virology and biological systematics. Comparing healthy and mutated DNA sequences can diagnose different diseases including various cancers, characterize antibody repertoire, and can be used to guide patient treatment. Having a quick way to sequence DNA allows for faster and more individualized medical care to be administered, and for more organisms to be identified and cataloged. The rapid advancements in DNA seque ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Shotgun Sequencing
In genetics, shotgun sequencing is a method used for sequencing random DNA strands. It is named by analogy with the rapidly expanding, quasi-random shot grouping of a shotgun. The Sanger sequencing#Method, chain-termination method of DNA sequencing ("Sanger sequencing") can only be used for short DNA strands of 100 to 1000 base pairs. Due to this size limit, longer sequences are subdivided into smaller fragments that can be sequenced separately, and these sequences are sequence assembly, assembled to give the overall sequence. In shotgun sequencing, DNA is broken up randomly into numerous small segments, which are sequenced using the chain termination method to obtain ''reads''. Multiple overlapping reads for the target DNA are obtained by performing several rounds of this fragmentation and sequencing. Computer programs then use the overlapping ends of different reads to assemble them into a continuous sequence. Shotgun sequencing was one of the precursor technologies that was resp ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
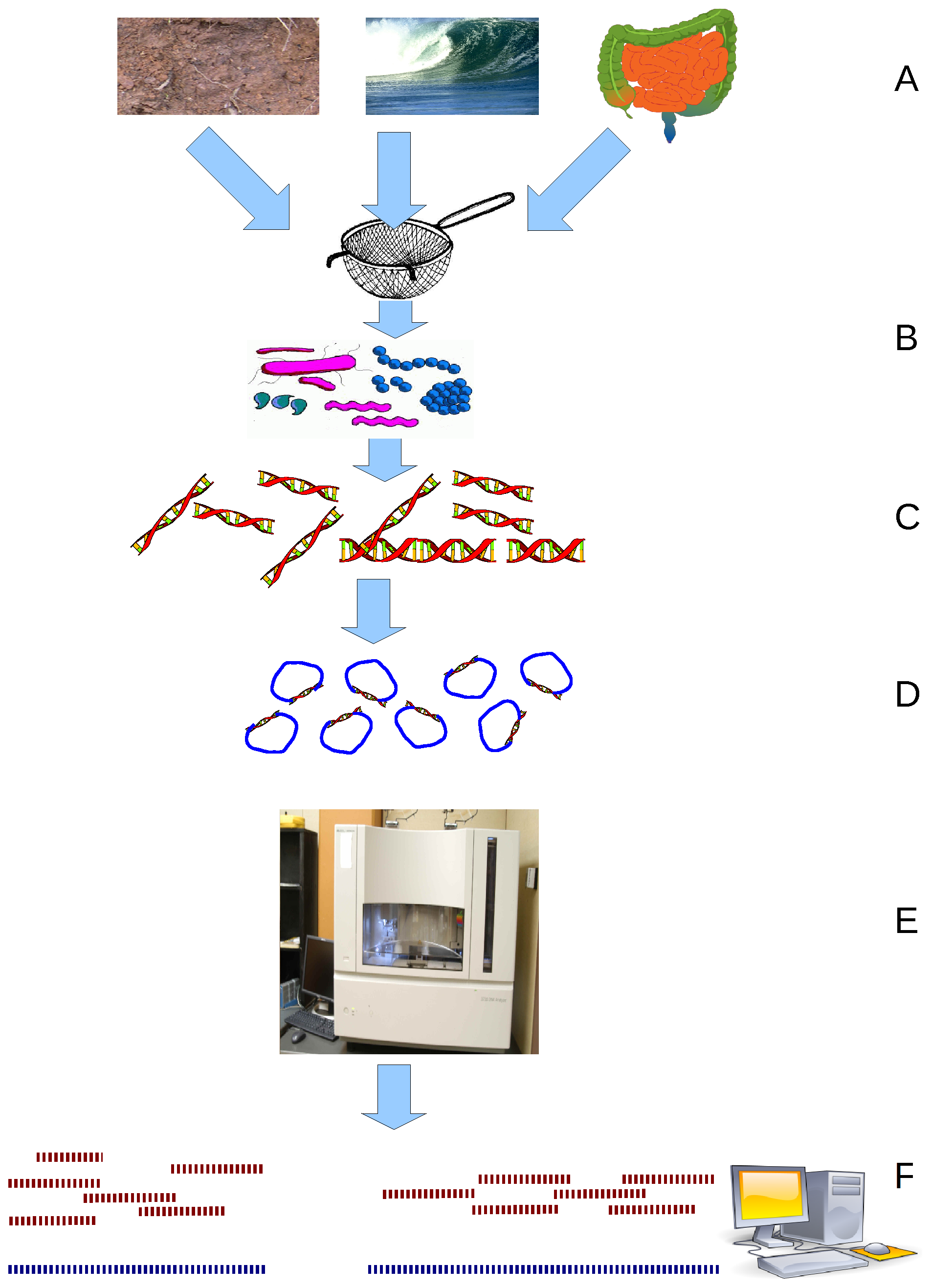
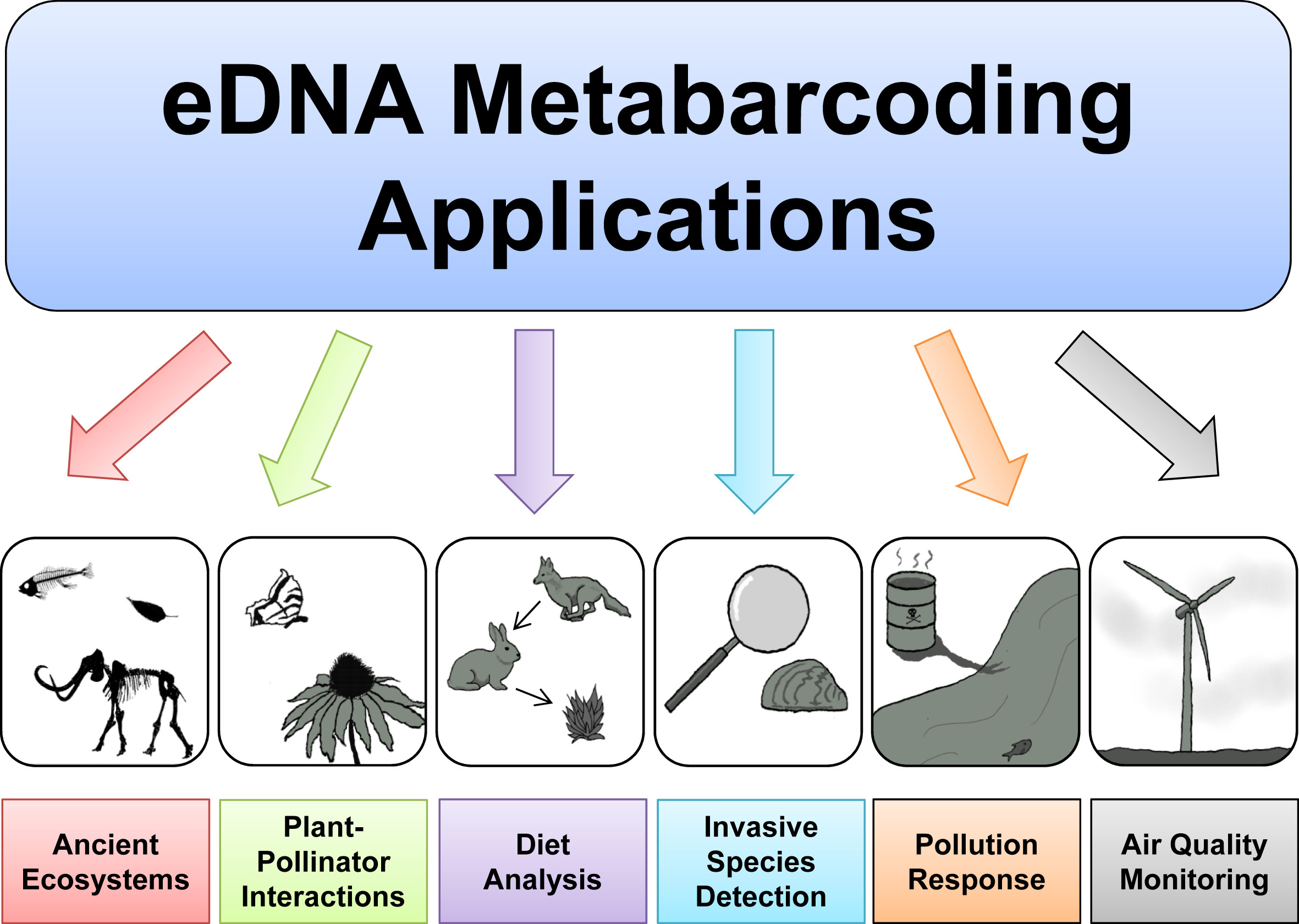
Metabarcoding
Metabarcoding is the DNA barcoding, barcoding of DNA/RNA (or Environmental DNA, eDNA/Environmental DNA, eRNA) in a manner that allows for the simultaneous identification of many taxa within the same sample. The main difference between barcoding and metabarcoding is that metabarcoding does not focus on one specific organism, but instead aims to determine species composition within a sample. A barcode consists of a short variable gene region (for example, see Algae DNA barcoding#Target regions, different markers/barcodes) which is useful for taxonomic assignment flanked by highly conserved gene regions which can be used for Primer (molecular biology), primer design. This idea of general barcoding originated in 2003 from researchers at the University of Guelph. The metabarcoding procedure, like general barcoding, proceeds in order through stages of DNA extraction, Polymerase chain reaction, PCR amplification, sequencing and data analysis. Different genes are used depending if the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Jo Handelsman
Jo Emily Handelsman (born March 19, 1959) is the Director of the Wisconsin Institute for Discovery at University of Wisconsin–Madison. She is also a Vilas Research Professor and a Howard Hughes Medical Institute Professor. Handelsman was appointed by President Barack Obama as the Associate Director for Science at the White House Office of Science and Technology Policy, where she served for three years until January 2017. She has been editor-in-chief of the academic journal '' DNA and Cell Biology'' and author of books on scientific education, most notably ''Scientific Teaching''. Education Handelsman was born on March 19, 1959, in New York City. She earned her Bachelor of Science degree in agronomy from Cornell University in 1979 and her Ph.D. in molecular biology from the University of Wisconsin–Madison in 1984. Career Handelsman secured a faculty position in plant pathology at the University of Wisconsin–Madison in 1985. She remained at Wisconsin until 2009, and then took ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
16S Ribosomal RNA
16S ribosomal RNA (or 16 S rRNA) is the RNA component of the 30S subunit of a prokaryotic ribosome ( SSU rRNA). It binds to the Shine-Dalgarno sequence and provides most of the SSU structure. The genes coding for it are referred to as 16S rRNA genes and are used in reconstructing phylogenies, due to the slow rates of evolution of this region of the gene. Carl Woese and George E. Fox were two of the people who pioneered the use of 16S rRNA in phylogenetics in 1977. Multiple sequences of the 16S rRNA gene can exist within a single bacterium. Terminology The descriptor ''16S'' refers to the size of these ribosomal subunits as reflected indirectly by the speed at which they sediment when samples are centrifuged. Thus ''16S'' means 16 Svedburg units. Functions * Like the large (23S) ribosomal RNA, it has a structural role, acting as a scaffold defining the positions of the ribosomal proteins. * The 3-end contains the anti- Shine-Dalgarno sequence, which binds upstream ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Jon Clardy
Jon Clardy (born May 16, 1943, Washington, D.C., United States) is currently the Hsien Wu and Daisy Yen Wu professor of biological chemistry and molecular pharmacology at Harvard Medical School. His research focuses on the isolation and structural characterization of natural products, and currently investigates the role of biologically active small molecules in mediating symbiotic interactions and disease. Biography Clardy grew up in Arlington, Virginia, United States, the oldest of four children. He attended Yale University where he received a B.S. in 1964 and was elected to Phi Beta Kappa. While he was always captivated by biology, during college he became more interested in chemistry. He performed undergraduate research in organic synthesis, directed by R. Stephen Berry, with an emphasis on benzyne. After graduating from Yale, he moved to Harvard University, where he received a Ph.D. in chemistry in 1969. He then accepted a faculty position in the Chemistry Department at Iowa ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Conserved Sequence
In evolutionary biology, conserved sequences are identical or similar sequences in nucleic acids ( DNA and RNA) or proteins across species ( orthologous sequences), or within a genome ( paralogous sequences), or between donor and receptor taxa ( xenologous sequences). Conservation indicates that a sequence has been maintained by natural selection. A highly conserved sequence is one that has remained relatively unchanged far back up the phylogenetic tree, and hence far back in geological time. Examples of highly conserved sequences include the RNA components of ribosomes present in all domains of life, the homeobox sequences widespread amongst eukaryotes, and the tmRNA in bacteria. The study of sequence conservation overlaps with the fields of genomics, proteomics, evolutionary biology, phylogenetics, bioinformatics and mathematics. History The discovery of the role of DNA in heredity, and observations by Frederick Sanger of variation between animal insulins in 194 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RRNA
Ribosomal ribonucleic acid (rRNA) is a type of non-coding RNA which is the primary component of ribosomes, essential to all cells. rRNA is a ribozyme which carries out protein synthesis in ribosomes. Ribosomal RNA is transcribed from ribosomal DNA (rDNA) and then bound to ribosomal proteins to form SSU rRNA, small and LSU rRNA, large ribosome subunits. rRNA is the physical and mechanical factor of the ribosome that forces transfer RNA (tRNA) and messenger RNA (mRNA) to process and Translation (biology), translate the latter into proteins. Ribosomal RNA is the predominant form of RNA found in most cells; it makes up about 80% of cellular RNA despite never being translated into proteins itself. Ribosomes are composed of approximately 60% rRNA and 40% ribosomal proteins, though this ratio differs between Prokaryote, prokaryotes and Eukaryote, eukaryotes. Structure Although the primary structure of rRNA sequences can vary across organisms, Base pair, base-pairing within these sequ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Species
A species () is often defined as the largest group of organisms in which any two individuals of the appropriate sexes or mating types can produce fertile offspring, typically by sexual reproduction. It is the basic unit of Taxonomy (biology), classification and a taxonomic rank of an organism, as well as a unit of biodiversity. Other ways of defining species include their karyotype, DNA sequence, morphology (biology), morphology, behaviour, or ecological niche. In addition, palaeontologists use the concept of the chronospecies since fossil reproduction cannot be examined. The most recent rigorous estimate for the total number of species of eukaryotes is between 8 and 8.7 million. About 14% of these had been described by 2011. All species (except viruses) are given a binomial nomenclature, two-part name, a "binomen". The first part of a binomen is the name of a genus to which the species belongs. The second part is called the specific name (zoology), specific name or the specific ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is a branch of biology that seeks to understand the molecule, molecular basis of biological activity in and between Cell (biology), cells, including biomolecule, biomolecular synthesis, modification, mechanisms, and interactions. Though cells and other microscopic structures had been observed in living organisms as early as the 18th century, a detailed understanding of the mechanisms and interactions governing their behavior did not emerge until the 20th century, when technologies used in physics and chemistry had advanced sufficiently to permit their application in the biological sciences. The term 'molecular biology' was first used in 1945 by the English physicist William Astbury, who described it as an approach focused on discerning the underpinnings of biological phenomena—i.e. uncovering the physical and chemical structures and properties of biological molecules, as well as their interactions with other molecules and how these interactions explain observ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Archaea
Archaea ( ) is a Domain (biology), domain of organisms. Traditionally, Archaea only included its Prokaryote, prokaryotic members, but this has since been found to be paraphyletic, as eukaryotes are known to have evolved from archaea. Even though the domain Archaea Cladistics, cladistically includes eukaryotes, the term "archaea" (: archaeon , from the Greek "ἀρχαῖον", which means ancient) in English still generally refers specifically to prokaryotic members of Archaea. Archaea were initially Taxonomy (biology), classified as bacteria, receiving the name archaebacteria (, in the Archaebacteria Kingdom (biology), kingdom), but this term has fallen out of use. Archaeal cells have unique properties separating them from Bacteria and Eukaryote, Eukaryota. Archaea are further divided into multiple recognized phylum, phyla. Classification is difficult because most have not been Isolation (microbiology), isolated in a laboratory and have been detected only by their Gene, gene s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |