|
H3K4me1
H3K4me1 is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the mono-methylation at the 4th lysine residue of the histone H3 protein and often associated with gene enhancers. Nomenclature H3K4me1 indicates monomethylation of lysine 4 on histone H3 protein subunit: Lysine methylation : This diagram shows the progressive methylation of a lysine residue. The mono-methylation (second from left) denotes the methylation present in H3K4me1. Understanding histone modifications The genomic DNA of eukaryotic cells is wrapped around special protein molecules known as histones. The complexes formed by the looping of the DNA are known as chromatin. The basic structural unit of chromatin is the nucleosome: this consists of the core octamer of histones (H2A, H2B, H3 and H4) as well as a linker histone and about 180 base pairs of DNA. These core histones are rich in lysine and arginine residues. The carboxyl (C) terminal end of these histon ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
KMT2D
Histone-lysine N-methyltransferase 2D (KMT2D), also known as MLL4 and sometimes MLL2 in humans and Mll4 in mice, is a major mammalian histone H3 lysine 4 (H3K4) mono-methyltransferase. It is part of a family of six Set1-like H3K4 methyltransferases that also contains KMT2A (or MLL1), KMT2B (or MLL2), KMT2C (or MLL3), KMT2F (or SET1A), and KMT2G (or SET1B). KMT2D is a large protein over 5,500 amino acids in size and is widely expressed in adult tissues. The protein co-localizes with lineage determining transcription factors on transcriptional enhancers and is essential for cell differentiation and embryonic development. It also plays critical roles in regulating cell fate transition, metabolism, and tumor suppression. Mutations in KMT2D cause human genetic conditions including Kabuki syndrome, another distinct congenital malformations disorder and various forms of cancer. Structure Gene In mice, KMT2D is coded by the ''Kmt2d'' gene located on chromosome 15F1. Its transcript ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
H3K27ac
H3K27ac is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates acetylation of the lysine residue at N-terminal position 27 of the histone H3 protein. H3K27ac is associated with the higher activation of transcription and therefore defined as an ''active enhancer'' mark. H3K27ac is found at both proximal and distal regions of transcription start site (TSS). Lysine acetylation and deacetylation Proteins are typically acetylated on lysine residues, and the acetylation reaction relies on acetyl-coenzyme A as the acetyl group donor. In histone acetylation and deacetylation, histone proteins are acetylated and deacetylated on lysine residues in the N-terminal tail as part of gene regulation. Typically, these reactions are catalyzed by enzymes with ''histone acetyltransferase'' (HAT) or ''histone deacetylase'' (HDAC) activity, although HATs and HDACs can modify the acetylation status of non-histone proteins as well. The regulation of t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
H3K4me3
H3K4me3 is an epigenetic modification to the DNA packaging protein Histone H3 that indicates tri-methylation at the 4th lysine residue of the histone H3 protein and is often involved in the regulation of gene expression. The name denotes the addition of three methyl groups ( trimethylation) to the lysine 4 on the histone H3 protein. H3 is used to package DNA in eukaryotic cells (including human cells), and modifications to the histone alter the accessibility of genes for transcription. H3K4me3 is commonly associated with the activation of transcription of nearby genes. H3K4 trimethylation regulates gene expression through chromatin remodeling by the NURF complex. This makes the DNA in the chromatin more accessible for transcription factors, allowing the genes to be transcribed and expressed in the cell. More specifically, H3K4me3 is found to positively regulate transcription by bringing histone acetylases and nucleosome remodelling enzymes (NURF). H3K4me3 also plays an importan ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histones
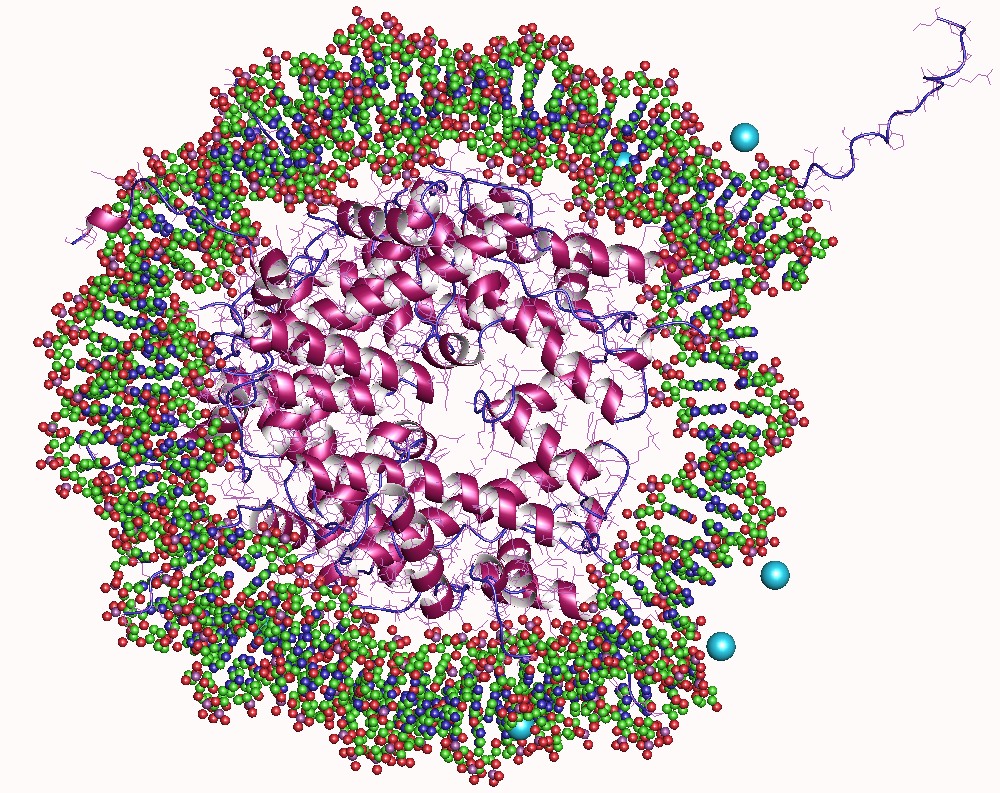
In biology, histones are highly Base (chemistry), basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei and in most Archaea, Archaeal Phylum, phyla. They act as spools around which DNA winds to create structural units called nucleosomes. Nucleosomes in turn are wrapped into 30-nanometer fibers that form tightly packed chromatin. Histones prevent DNA from becoming tangled and protect it from DNA damage. In addition, histones play important roles in gene regulation and DNA replication. Without histones, unwound DNA in chromosomes would be very long. For example, each human cell has about 1.8 meters of DNA if completely stretched out; however, when wound about histones, this length is reduced to about 9 micrometers (0.09 mm) of 30 nm diameter chromatin fibers. There are five families of histones, which are designated H1/H5 (linker histones), H2, H3, and H4 (core histones). The nucleosome core is formed of two H2A-H2B protein dimer ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ENCODE
The Encyclopedia of DNA Elements (ENCODE) is a public research project which aims "to build a comprehensive parts list of functional elements in the human genome." ENCODE also supports further biomedical research by "generating community resources of genomics data, software, tools and methods for genomics data analysis, and products resulting from data analyses and interpretations." The current phase of ENCODE (2016-2019) is adding depth to its resources by growing the number of cell types, data types, assays and now includes support for examination of the mouse genome. History ENCODE was launched by the US National Human Genome Research Institute (NHGRI) in September 2003. Intended as a follow-up to the Human Genome Project, the ENCODE project aims to identify all functional elements in the human genome. The project involves a worldwide consortium of research groups, and data generated from this project can be accessed through public databases. The initial release of ENCODE ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone H3
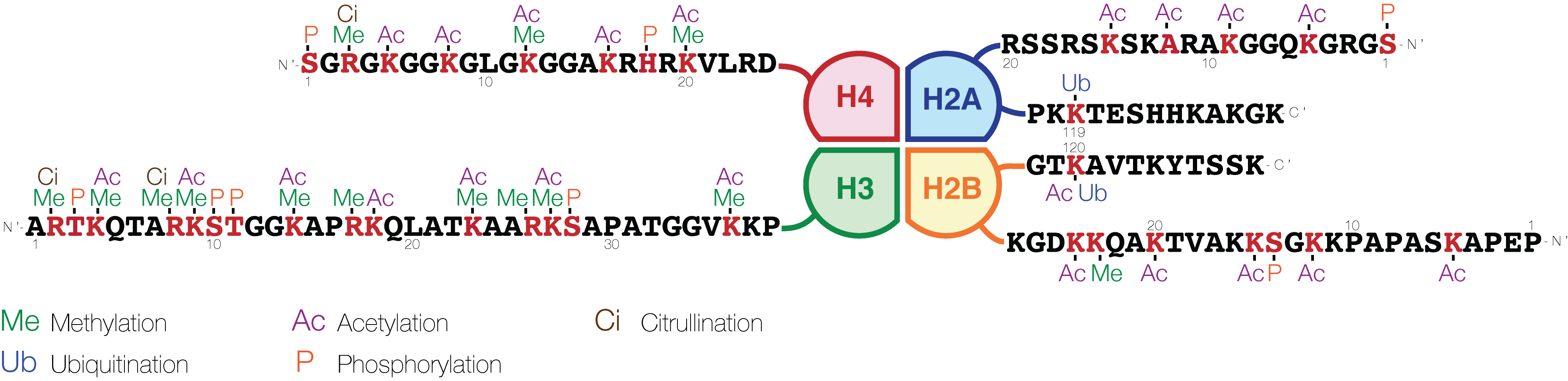
Histone H3 is one of the five main histones involved in the structure of chromatin in eukaryotic cells. Featuring a main globular domain and a long N-terminal end, N-terminal tail, H3 is involved with the structure of the nucleosomes of the 'beads on a string' structure. Histone proteins are highly post-translationally modified however Histone H3 is the most extensively modified of the five histones. The term "Histone H3" alone is purposely ambiguous in that it does not distinguish between sequence variants or modification state. Histone H3 is an important protein in the emerging field of epigenetics, where its sequence variants and variable modification states are thought to play a role in the dynamic and long term regulation of genes. Epigenetics and post-translational modifications The N-terminus of H3 protrudes from the globular nucleosome core and is susceptible to post-translational modification that influence cellular processes. These modifications include the covalen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
P300-CBP Coactivator Family
The p300-CBP coactivator family in humans is composed of two closely related transcriptional co-activating proteins (or coactivators): #p300 (also called EP300 or E1A binding protein p300) # CBP (also known as CREB-binding protein or CREBBP) Both p300 and CBP interact with numerous transcription factors and act to increase the expression of their target genes. Protein structure p300 and CBP have similar structures. Both contain five protein interaction domains: the nuclear receptor interaction domain (RID), the KIX domain (CREB and MYB interaction domain), the cysteine/histidine regions (TAZ1/CH1 and TAZ2/CH3) and the interferon response binding domain (IBiD). The last four domains, KIX, TAZ1, TAZ2 and IBiD of p300, each bind tightly to a sequence spanning both transactivation domains 9aaTADs of transcription factor p53. In addition p300 and CBP each contain a protein or histone acetyltransferase (PAT/HAT) domain and a bromodomain that binds acetylated lysines and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
H3K4
{{set index article ...
H3K4 is the fourth lysine residue on DNA packaging protein Histone H3. It can be marked by epigenetic modification by different amounts of methylation. Modifications include: *H3K4me1 * H3K4me2 *H3K4me3 H3K4me3 is an epigenetic modification to the DNA packaging protein Histone H3 that indicates tri-methylation at the 4th lysine residue of the histone H3 protein and is often involved in the regulation of gene expression. The name denotes the addit ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone Code
The histone code is a hypothesis that the transcription of genetic information encoded in DNA is in part regulated by chemical modifications (known as ''histone marks'') to histone proteins, primarily on their unstructured ends. Together with similar modifications such as DNA methylation it is part of the epigenetic code. Histones associate with DNA to form nucleosomes, which themselves bundle to form chromatin fibers, which in turn make up the more familiar chromosome. Histones are globular proteins with a flexible N-terminus (taken to be the tail) that protrudes from the nucleosome. Many of the histone tail modifications correlate very well to chromatin structure and both histone modification state and chromatin structure correlate well to gene expression levels. The critical concept of the histone code hypothesis is that the histone modifications serve to recruit other proteins by specific recognition of the modified histone via protein domains specialized for such purposes, ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
LSD1
Lysine-specific histone demethylase 1A (LSD1) also known as lysine (K)-specific demethylase 1A (KDM1A) is a protein that in humans is encoded by the ''KDM1A'' gene. LSD1 is a flavin-dependent monoamine oxidase, which can demethylate mono- and di-methylated lysines, specifically histone 3, lysine 4 (H3K4). Other reported methylated lysine substrates such as histone H3K9 and TP53 have not been biochemically validated. This enzyme plays a critical role in oocyte growth, embryogenesis, hematopoiesis and tissue-specific differentiation. LSD1 was the first histone demethylase to be discovered more than 30 years ago. Structure This gene encodes a nuclear protein containing a SWIRM domain, a FAD-binding motif, and an amine oxidase domain. This protein is a component of several complexes that include histone deacetylase and DNA methytransferase 1, all of which are associated with the repression of gene transcription. It is now known the LSD1 complex mediates a coordinated histone ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
H3K27me3
H3K27me3 is an epigenetic modification to the DNA packaging protein histone H3. It is a mark that indicates the tri-methylation of lysine 27 on histone H3 protein. This tri-methylation is associated with the Downregulation and upregulation, downregulation of nearby genes via the formation of Heterochromatin, heterochromatic regions. Nomenclature H3K27me3 indicates Methylation, trimethylation of lysine 27 on histone H3 protein subunit: Lysine methylation : This diagram shows the progressive methylation of a lysine residue. The tri-methylation (right) denotes the methylation present in H3K27me3. Understanding histone modifications The genomic DNA of eukaryotic cells is wrapped around special protein molecules known as histones. The complexes formed by the looping of the DNA are known as chromatin. The basic structural unit of chromatin is the nucleosome: this consists of the core octamer of histones (H2A, H2B, H3 and H4) as well as a linker histone and about 180 base pairs of D ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
H3K36me3
H3K36me3 is an epigenetic modification to the DNA packaging protein Histone H3. It is a mark that indicates the tri-methylation at the 36th lysine residue of the histone H3 protein and often associated with gene, gene bodies. There are diverse modifications at H3K36 and have many important biological processes. H3K36 has different acetylation and methylation states with no similarity to each other. Nomenclature H3K36me3 indicates Methylation, trimethylation of lysine 36 on histone H3 protein subunit: Lysine methylation : This diagram shows the progressive methylation of a lysine residue. The tri-methylation (right) denotes the methylation present in H3K36me3. Understanding histone modifications The genomic DNA of eukaryotic cells is wrapped around special protein molecules known as Histones. The complexes formed by the looping of the DNA are known as chromatin. The basic structural unit of chromatin is the nucleosome: this consists of the core octamer of histones (H2A, H2B ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |