|
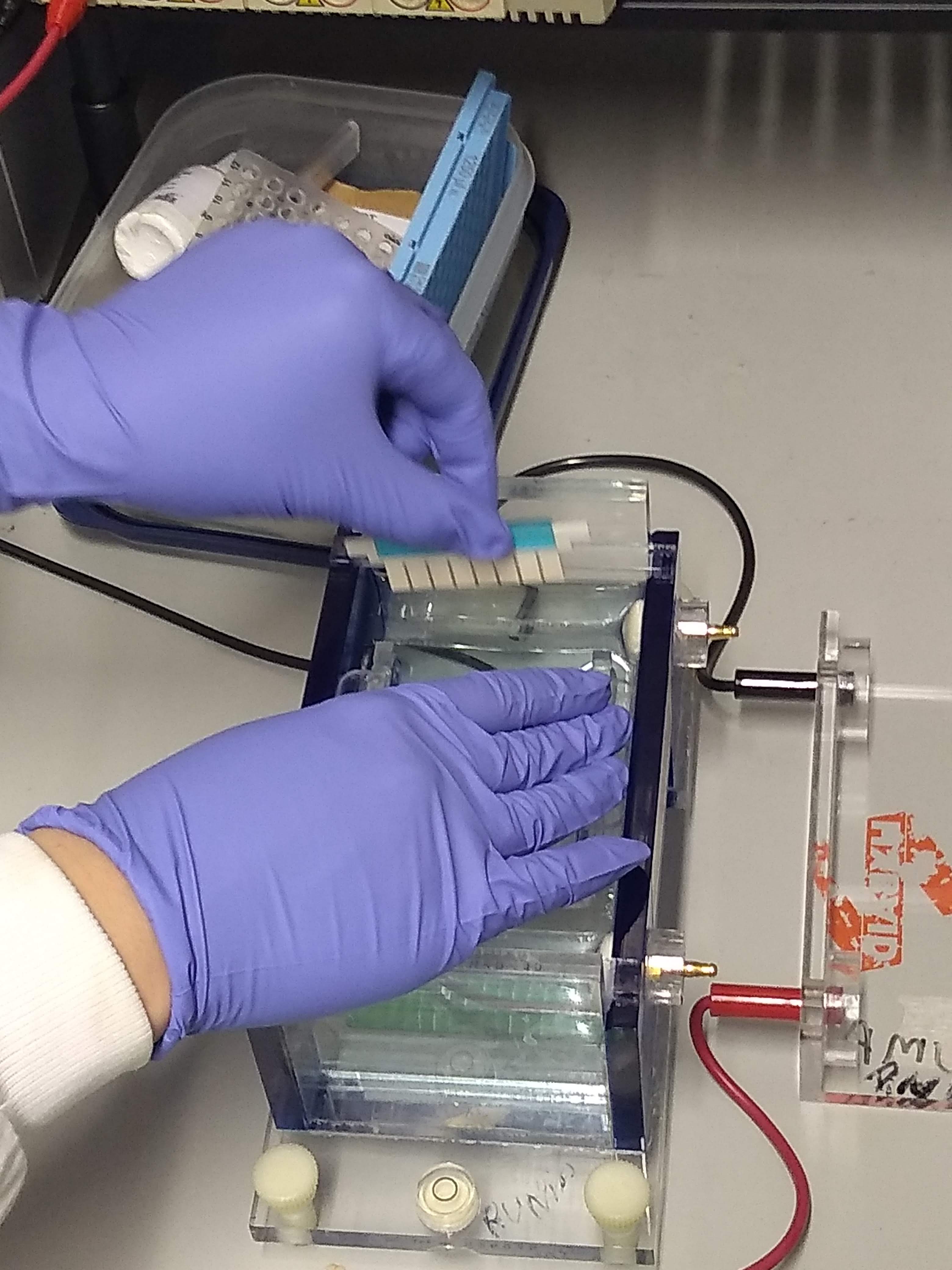
Genotyping
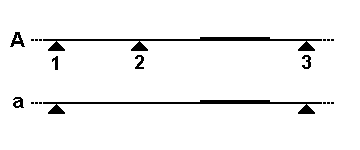
Genotyping is the process of determining differences in the genetic make-up (genotype) of an individual by examining the individual's DNA sequence using bioassay, biological assays and comparing it to another individual's sequence or a reference sequence. It reveals the alleles an individual has inherited from their parents. Traditionally genotyping is the use of DNA sequences to define biological Population, populations by use of molecular tools. It does not usually involve defining the genes of an individual. Techniques Restriction Fragment Length Polymorphisms A restriction fragment length polymorphism (RFLP) is a variation between different people at sites of the genome recognized by Restriction enzyme, restriction enzymes. DNA containing different restriction sites will be cut by bacterial restriction enzymes differently and this can be seen using gel electrophoresis. When running the sample through, a successfully cleaved sample will contain two bands, while the sample wit ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Single-nucleotide Polymorphism
In genetics and bioinformatics, a single-nucleotide polymorphism (SNP ; plural SNPs ) is a germline substitution of a single nucleotide at a specific position in the genome. Although certain definitions require the substitution to be present in a sufficiently large fraction of the population (e.g. 1% or more), many publications do not apply such a frequency threshold. For example, a Guanine, G nucleotide present at a specific location in a reference genome may be replaced by an Adenine, A in a minority of individuals. The two possible nucleotide variations of this SNP – G or A – are called alleles. SNPs can help explain differences in susceptibility to a wide range of diseases across a population. For example, a common SNP in the Factor H, CFH gene is associated with increased risk of age-related macular degeneration. Differences in the severity of an illness or response to treatments may also be manifestations of genetic variations caused by SNPs. For example, two ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Microarray
A DNA microarray (also commonly known as a DNA chip or biochip) is a collection of microscopic DNA spots attached to a solid surface. Scientists use DNA microarrays to measure the expression levels of large numbers of genes simultaneously or to genotype multiple regions of a genome. Each DNA spot contains picomoles (10−12 moles) of a specific DNA sequence, known as '' probes'' (or ''reporters'' or '' oligos''). These can be a short section of a gene or other DNA element that are used to hybridize a cDNA or cRNA (also called anti-sense RNA) sample (called ''target'') under high-stringency conditions. Probe-target hybridization is usually detected and quantified by detection of fluorophore-, silver-, or chemiluminescence-labeled targets to determine relative abundance of nucleic acid sequences in the target. The original nucleic acid arrays were macro arrays approximately 9 cm × 12 cm and the first computerized image based analysis was published in 1981. It was ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Fragment Length Polymorphism
In molecular biology, restriction fragment length polymorphism (RFLP) is a technique that exploits variations in homologous DNA sequences, known as polymorphisms, populations, or species or to pinpoint the locations of genes within a sequence. The term may refer to a polymorphism itself, as detected through the differing locations of restriction enzyme sites, or to a related laboratory technique by which such differences can be illustrated. In RFLP analysis, a DNA sample is digested into fragments by one or more restriction enzymes, and the resulting ''restriction fragments'' are then separated by gel electrophoresis according to their size. RFLP analysis is now largely obsolete due to the emergence of inexpensive DNA sequencing technologies, but it was the first DNA profiling technique inexpensive enough to see widespread application. RFLP analysis was an important early tool in genome mapping, localization of genes for genetic disorders, determination of risk for disease, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to make millions to billions of copies of a specific DNA sample rapidly, allowing scientists to amplify a very small sample of DNA (or a part of it) sufficiently to enable detailed study. PCR was invented in 1983 by American biochemist Kary Mullis at Cetus Corporation. Mullis and biochemist Michael Smith (chemist), Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of Ancient DNA, ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research and forensic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Amplified Fragment Length Polymorphism
Amplified fragment length polymorphism (AFLP-PCR or AFLP) is a PCR-based tool used in genetics research, DNA fingerprinting, and in the practice of genetic engineering. Developed in the early 1990s by Pieter Vos, AFLP uses restriction enzymes to digest genomic DNA, followed by ligation of adaptors to the sticky ends of the restriction fragments. A subset of the restriction fragments is then selected to be amplified. This selection is achieved by using primers complementary to the adaptor sequence, the restriction site sequence and a few nucleotides inside the restriction site fragments (as described in detail below). The amplified fragments are separated and visualized on denaturing on agarose gel electrophoresis, either through autoradiography or fluorescence methodologies, or via automated capillary sequencing instruments. Although AFLP should not be used as an acronym, it is commonly referred to as "Amplified fragment length polymorphism". However, the resulting data are ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genotype
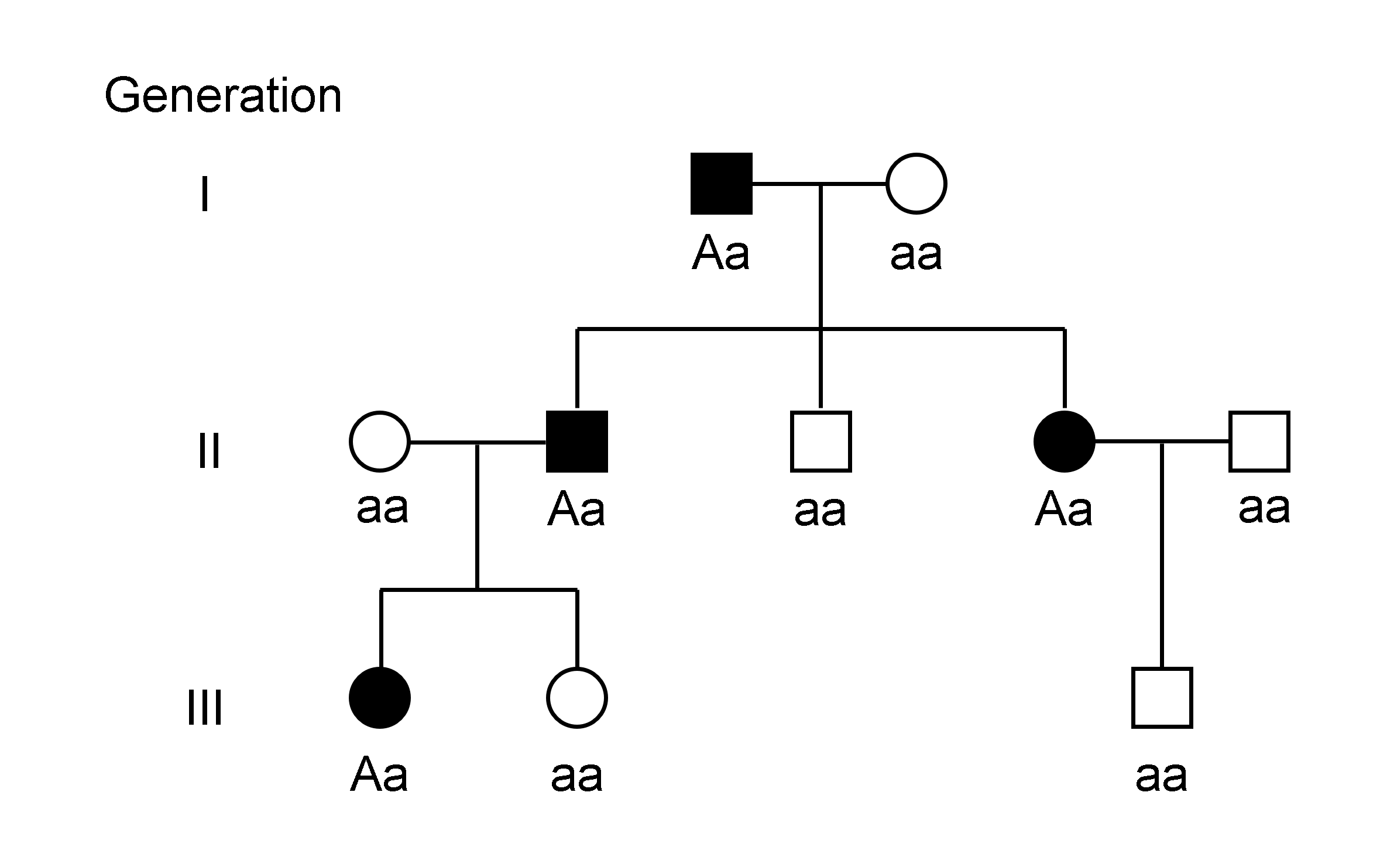
The genotype of an organism is its complete set of genetic material. Genotype can also be used to refer to the alleles or variants an individual carries in a particular gene or genetic location. The number of alleles an individual can have in a specific gene depends on the number of copies of each chromosome found in that species, also referred to as ploidy. In diploid species like humans, two full sets of chromosomes are present, meaning each individual has two alleles for any given gene. If both alleles are the same, the genotype is referred to as Zygosity, homozygous. If the alleles are different, the genotype is referred to as heterozygous. Genotype contributes to phenotype, the observable traits and characteristics in an individual or organism. The degree to which genotype affects phenotype depends on the trait. For example, the petal color in a pea plant is exclusively determined by genotype. The petals can be purple or white depending on the alleles present in the pea plan ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gel Electrophoresis
Gel electrophoresis is an electrophoresis method for separation and analysis of biomacromolecules (DNA, RNA, proteins, etc.) and their fragments, based on their size and charge through a gel. It is used in clinical chemistry to separate proteins by charge or size (IEF agarose, essentially size independent) and in biochemistry and molecular biology to separate a mixed population of DNA and RNA fragments by length, to estimate the size of DNA and RNA fragments, or to separate proteins by charge. Nucleic acid molecules are separated by applying an electric field to move the negatively charged molecules through a gel matrix of agarose, polyacrylamide, or other substances. Shorter molecules move faster and migrate farther than longer ones because shorter molecules migrate more easily through the pores of the gel. This phenomenon is called sieving. Proteins are separated by the charge in agarose because the pores of the gel are too large to sieve proteins. Gel electrophoresi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bacteria
Bacteria (; : bacterium) are ubiquitous, mostly free-living organisms often consisting of one Cell (biology), biological cell. They constitute a large domain (biology), domain of Prokaryote, prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit the air, soil, water, Hot spring, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria play a vital role in many stages of the nutrient cycle by recycling nutrients and the nitrogen fixation, fixation of nitrogen from the Earth's atmosphere, atmosphere. The nutrient cycle includes the decomposition of cadaver, dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, suc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Uracil
Uracil () (nucleoside#List of nucleosides and corresponding nucleobases, symbol U or Ura) is one of the four nucleotide bases in the nucleic acid RNA. The others are adenine (A), cytosine (C), and guanine (G). In RNA, uracil binds to adenine via two hydrogen bonds. In DNA, the uracil nucleobase is replaced by thymine (T). Uracil is a demethylated form of thymine. Uracil is a common and naturally occurring pyrimidine derivative. The name "uracil" was coined in 1885 by the German chemist Robert Behrend, who was attempting to synthesize derivatives of uric acid. Originally discovered in 1900 by Alberto Ascoli, it was isolated by hydrolysis of yeast nuclein; it was also found in bovine thymus and spleen, herring sperm, and wheat Cereal germ, germ. It is a planar, unsaturated compound that has the ability to absorb light. Uracil that was formed extraterrestrially has been detected in the Murchison meteorite, in near-Earth asteroid 162173 Ryugu, Ryugu, and possibly on the surface of th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cytosine
Cytosine () (symbol C or Cyt) is one of the four nucleotide bases found in DNA and RNA, along with adenine, guanine, and thymine ( uracil in RNA). It is a pyrimidine derivative, with a heterocyclic aromatic ring and two substituents attached (an amine group at position 4 and a keto group at position 2). The nucleoside of cytosine is cytidine. In Watson–Crick base pairing, it forms three hydrogen bonds with guanine. History Cytosine was discovered and named by Albrecht Kossel and Albert Neumann in 1894 when it was hydrolyzed from calf thymus tissues. A structure was proposed in 1903, and was synthesized (and thus confirmed) in the laboratory in the same year. In 1998, cytosine was used in an early demonstration of quantum information processing when Oxford University researchers implemented the Deutsch–Jozsa algorithm on a two qubit nuclear magnetic resonance quantum computer (NMRQC). In March 2015, NASA scientists reported the formation of cytosine, alon ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Deamination
Deamination is the removal of an amino group from a molecule. Enzymes that catalysis, catalyse this reaction are called deaminases. In the human body, deamination takes place primarily in the liver; however, it can also occur in the kidney. In situations of excess protein intake, deamination is used to break down amino acids for energy. The amino group is removed from the amino acid and converted to ammonia. The rest of the amino acid is made up of mostly carbon and hydrogen, and is recycled or oxidized for energy. Ammonia is toxic to the human system, and enzymes convert it to urea or uric acid by addition of carbon dioxide molecules (which is not considered a deamination process) in the urea cycle, which also takes place in the liver. Urea and uric acid can safely diffuse into the blood and then be excreted in urine. Deamination reactions in DNA Cytosine Spontaneous deamination is the hydrolysis reaction of cytosine into uracil, releasing ammonia in the process. This can occu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Evolution
Evolution is the change in the heritable Phenotypic trait, characteristics of biological populations over successive generations. It occurs when evolutionary processes such as natural selection and genetic drift act on genetic variation, resulting in certain characteristics becoming more or less common within a population over successive generations. The process of evolution has given rise to biodiversity at every level of biological organisation. The scientific theory of evolution by natural selection was conceived independently by two British naturalists, Charles Darwin and Alfred Russel Wallace, in the mid-19th century as an explanation for why organisms are adapted to their physical and biological environments. The theory was first set out in detail in Darwin's book ''On the Origin of Species''. Evolution by natural selection is established by observable facts about living organisms: (1) more offspring are often produced than can possibly survive; (2) phenotypic variatio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |