|
Cleaved Amplified Polymorphic Sequence
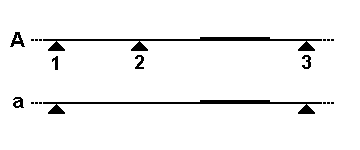
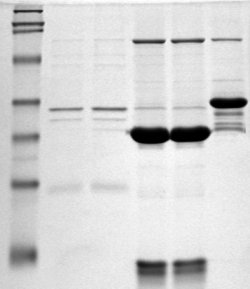
The cleaved amplified polymorphic sequence (CAPS) method is a technique in molecular biology for the analysis of genetic markers. It is an extension to the restriction fragment length polymorphism (RFLP) method, using polymerase chain reaction (PCR) to more quickly analyse the results. Like RFLP, CAPS works on the principle that genetic differences between individuals can create or abolish restriction endonuclease restriction sites, and that these differences can be detected in the resulting DNA fragment length after digestion. In the CAPS method, PCR amplification is directed across the altered restriction site, and the products digested with the restriction enzyme. When fractionated by agarose or polyacrylamide gel electrophoresis, the digested PCR products will give readily distinguishable patterns of bands. Alternatively, the amplified segment can be analyzed by allele-specific oligonucleotide (ASO) probes, a process that can often be done by a simple dot blot. See also * RF ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Biology
Molecular biology is a branch of biology that seeks to understand the molecule, molecular basis of biological activity in and between Cell (biology), cells, including biomolecule, biomolecular synthesis, modification, mechanisms, and interactions. Though cells and other microscopic structures had been observed in living organisms as early as the 18th century, a detailed understanding of the mechanisms and interactions governing their behavior did not emerge until the 20th century, when technologies used in physics and chemistry had advanced sufficiently to permit their application in the biological sciences. The term 'molecular biology' was first used in 1945 by the English physicist William Astbury, who described it as an approach focused on discerning the underpinnings of biological phenomena—i.e. uncovering the physical and chemical structures and properties of biological molecules, as well as their interactions with other molecules and how these interactions explain observ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Marker
A genetic marker is a gene or DNA sequence with a known location on a chromosome that can be used to identify individuals or species. It can be described as a variation (which may arise due to mutation or alteration in the genomic loci) that can be observed. A genetic marker may be a short DNA sequence, such as a sequence surrounding a single base-pair change ( single nucleotide polymorphism, SNP), or a long one, like minisatellites. Background For many years, gene mapping was limited to identifying organisms by traditional phenotypes markers. This included genes that encoded easily observable characteristics, such as blood types or seed shapes. The insufficient number of these types of characteristics in several organisms limited the possible mapping efforts. This prompted the development of gene markers, which could identify genetic characteristics that are not readily observable in organisms (such as protein variation). Types Some commonly used types of genetic markers ar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Fragment Length Polymorphism
In molecular biology, restriction fragment length polymorphism (RFLP) is a technique that exploits variations in homologous DNA sequences, known as polymorphisms, populations, or species or to pinpoint the locations of genes within a sequence. The term may refer to a polymorphism itself, as detected through the differing locations of restriction enzyme sites, or to a related laboratory technique by which such differences can be illustrated. In RFLP analysis, a DNA sample is digested into fragments by one or more restriction enzymes, and the resulting ''restriction fragments'' are then separated by gel electrophoresis according to their size. RFLP analysis is now largely obsolete due to the emergence of inexpensive DNA sequencing technologies, but it was the first DNA profiling technique inexpensive enough to see widespread application. RFLP analysis was an important early tool in genome mapping, localization of genes for genetic disorders, determination of risk for disease, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to make millions to billions of copies of a specific DNA sample rapidly, allowing scientists to amplify a very small sample of DNA (or a part of it) sufficiently to enable detailed study. PCR was invented in 1983 by American biochemist Kary Mullis at Cetus Corporation. Mullis and biochemist Michael Smith (chemist), Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of Ancient DNA, ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research and forensic ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Endonuclease
A restriction enzyme, restriction endonuclease, REase, ENase or'' restrictase '' is an enzyme that cleaves DNA into fragments at or near specific recognition sites within molecules known as restriction sites. Restriction enzymes are one class of the broader endonuclease group of enzymes. Restriction enzymes are commonly classified into five types, which differ in their structure and whether they cut their DNA enzyme substrate (biology), substrate at their recognition site, or if the recognition and cleavage sites are separate from one another. To cut DNA, all restriction enzymes make two incisions, once through each backbone chain, sugar-phosphate backbone (i.e. each strand) of the DNA double helix. These enzymes are found in bacteria and archaea and provide a defense mechanism against invading viruses. Inside a prokaryote, the restriction enzymes selectively cut up ''foreign'' DNA in a process called ''restriction digestion''; meanwhile, host DNA is protected by a modification ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Agarose Gel Electrophoresis
Agarose gel electrophoresis is a method of gel electrophoresis used in biochemistry, molecular biology, genetics, and clinical chemistry to separate a mixed population of macromolecules such as DNA or proteins in a matrix of agarose, one of the two main components of agar. The proteins may be separated by charge and/or size (isoelectric focusing agarose electrophoresis is essentially size independent), and the DNA and RNA fragments by length. Biomolecules are separated by applying an electric field to move the charged molecules through an agarose matrix, and the biomolecules are separated by size in the agarose gel matrix. Agarose gel is easy to cast, has relatively fewer charged groups, and is particularly suitable for separating DNA of size range most often encountered in laboratories, which accounts for the popularity of its use. The separated DNA may be viewed with stain, most commonly under UV light, and the DNA fragments can be extracted from the gel with relative ease. Most ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polyacrylamide Gel Electrophoresis
Polyacrylamide gel electrophoresis (PAGE) is a technique widely used in biochemistry, forensic chemistry, genetics, molecular biology and biotechnology to separate biological macromolecules, usually proteins or nucleic acids, according to their Electrophoresis, electrophoretic mobility. Electrophoretic mobility is a function of the length, conformation, and charge of the molecule. Polyacrylamide gel electrophoresis is a powerful tool used to analyze RNA samples. When polyacrylamide gel is denatured after electrophoresis, it provides information on the sample composition of the RNA species. Hydration reaction, Hydration of acrylonitrile results in formation of acrylamide molecules () by nitrile hydratase. Acrylamide monomer is in a powder state before addition of water. Acrylamide is toxic to the human nervous system, therefore all safety measures must be followed when working with it. Acrylamide is soluble in water and upon addition of free-radical initiators it polymerizes resu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Allele-specific Oligonucleotide
An anti-sense oligonucleotide (ASO) is a short piece of synthetic DNA complementary to the sequence of a variable target DNA. It acts as a probe for the presence of the target in a Southern blot assay or, more commonly, in the simpler dot blot assay. It is a common tool used in genetic testing, forensics, and molecular biology research. An ASO is typically an oligonucleotide of 15–21 nucleotide bases in length. It is designed (and used) in a way that makes it specific for only one version, or allele, of the DNA being tested. The length of the ASO, which strand it is chosen from, and the conditions by which it is bound to (and washed from) the target DNA all play a role in its specificity. These probes can usually be designed to detect a difference of as little as 1 base in the target's genetic sequence, a basic ability in the assay of single-nucleotide polymorphisms (SNPs), important in genotype analysis and the Human Genome Project. To be detected after it has bound to its ta ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Dot Blot
A dot blot (or slot blot) is a technique in molecular biology used to detect proteins. It represents a simplification of the western blot method, with the exception that the proteins to be detected are not first separated by electrophoresis. Instead, the sample is applied directly on a membrane in a single spot, and the blotting procedure is performed. The technique offers significant savings in time, as chromatography or gel electrophoresis, and the complex blotting procedures for the gel are not required. However, it offers no information on the size of the target protein. Uses Performing a dot blot is similar in idea to performing a western blot, with the advantage of faster speed and lower cost. Dot blots are also performed to screen the binding capabilities of an antibody. Methodology A general dot blot protocol involves spotting 1–2 microliters of a samples onto a nitrocellulose Nitrocellulose (also known as cellulose nitrate, flash paper, flash cotton, guncot ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Restriction Fragment Length Polymorphism
In molecular biology, restriction fragment length polymorphism (RFLP) is a technique that exploits variations in homologous DNA sequences, known as polymorphisms, populations, or species or to pinpoint the locations of genes within a sequence. The term may refer to a polymorphism itself, as detected through the differing locations of restriction enzyme sites, or to a related laboratory technique by which such differences can be illustrated. In RFLP analysis, a DNA sample is digested into fragments by one or more restriction enzymes, and the resulting ''restriction fragments'' are then separated by gel electrophoresis according to their size. RFLP analysis is now largely obsolete due to the emergence of inexpensive DNA sequencing technologies, but it was the first DNA profiling technique inexpensive enough to see widespread application. RFLP analysis was an important early tool in genome mapping, localization of genes for genetic disorders, determination of risk for disease, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Profiling Techniques
Deoxyribonucleic acid (; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a Nucleic acid double helix, double helix. The polymer carries genetics, genetic instructions for the development, functioning, growth and reproduction of all known organisms and many viruses. DNA and ribonucleic acid (RNA) are nucleic acids. Alongside proteins, lipids and complex carbohydrates (polysaccharides), nucleic acids are one of the four major types of macromolecules that are essential for all known forms of life. The two DNA strands are known as polynucleotides as they are composed of simpler monomeric units called nucleotides. Each nucleotide is composed of one of four nitrogenous base, nitrogen-containing nucleobases (cytosine [C], guanine [G], adenine [A] or thymine [T]), a monosaccharide, sugar called deoxyribose, and a Organophosphate, phosphate group. The nucleotides are joined to one another in a chain by covalent bonds (known as the Phosphod ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |