|
Centromeric
The centromere links a pair of sister chromatids together during cell division. This constricted region of chromosome connects the sister chromatids, creating a short arm (p) and a long arm (q) on the chromatids. During mitosis, spindle fibers attach to the centromere via the kinetochore. The physical role of the centromere is to act as the site of assembly of the kinetochores – a highly complex multiprotein structure that is responsible for the actual events of chromosome segregation – i.e. binding microtubules and signaling to the cell cycle machinery when all chromosomes have adopted correct attachments to the spindle, so that it is safe for cell division to proceed to completion and for cells to enter anaphase. There are, broadly speaking, two types of centromeres. "Point centromeres" bind to specific proteins that recognize particular DNA sequences with high efficiency. Any piece of DNA with the point centromere DNA sequence on it will typically form a centromer ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
|
 |
Kinetochore
A kinetochore (, ) is a disc-shaped protein structure associated with duplicated chromatids in eukaryotic cells where the spindle fibers attach during cell division to pull sister chromatids apart. The kinetochore assembles on the centromere and links the chromosome to microtubule polymers from the mitotic spindle during mitosis and meiosis. The term kinetochore was first used in a footnote in a 1934 Cytology book by Lester W. Sharp and commonly accepted in 1936. Sharp's footnote reads: "The convenient term ''kinetochore'' (= movement place) has been suggested to the author by J. A. Moore", likely referring to John Alexander Moore who had joined Columbia University as a freshman in 1932. Monocentric organisms, including vertebrates, fungi, and most plants, have a single centromeric region on each chromosome which assembles a single, localized kinetochore. Holocentric organisms, such as nematodes and some plants, assemble a kinetochore along the entire length of a chrom ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
|
Kinetochores
A kinetochore (, ) is a disc-shaped protein structure associated with duplicated chromatids in eukaryotic cells where the spindle fibers attach during cell division to pull sister chromatids apart. The kinetochore assembles on the centromere and links the chromosome to microtubule polymers from the mitotic spindle during mitosis and meiosis. The term kinetochore was first used in a footnote in a 1934 Cytology book by Lester W. Sharp and commonly accepted in 1936. Sharp's footnote reads: "The convenient term ''kinetochore'' (= movement place) has been suggested to the author by J. A. Moore", likely referring to John Alexander Moore who had joined Columbia University as a freshman in 1932. Monocentric organisms, including vertebrates, fungi, and most plants, have a single centromeric region on each chromosome which assembles a single, localized kinetochore. Holocentric organisms, such as nematodes and some plants, assemble a kinetochore along the entire length of a chromosome. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
|
 |
Telocentric
The centromere links a pair of sister chromatids together during cell division. This constricted region of chromosome connects the sister chromatids, creating a short arm (p) and a long arm (q) on the chromatids. During mitosis, spindle fibers attach to the centromere via the kinetochore. The physical role of the centromere is to act as the site of assembly of the kinetochores – a highly complex multiprotein structure that is responsible for the actual events of chromosome segregation – i.e. binding microtubules and signaling to the cell cycle machinery when all chromosomes have adopted correct attachments to the spindle, so that it is safe for cell division to proceed to completion and for cells to enter anaphase. There are, broadly speaking, two types of centromeres. "Point centromeres" bind to specific proteins that recognize particular DNA sequences with high efficiency. Any piece of DNA with the point centromere DNA sequence on it will typically form a centromere if ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
 |
Karyotype
A karyotype is the general appearance of the complete set of metaphase chromosomes in the cells of a species or in an individual organism, mainly including their sizes, numbers, and shapes. Karyotyping is the process by which a karyotype is discerned by determining the chromosome complement of an individual, including the number of chromosomes and any abnormalities. A karyogram or idiogram is a graphical depiction of a karyotype, wherein chromosomes are organized in pairs, ordered by size and position of centromere for chromosomes of the same size. Karyotyping generally combines light microscopy and photography, and results in a photomicrographic (or simply micrographic) karyogram. In contrast, a schematic karyogram is a designed graphic representation of a karyotype. In schematic karyograms, just one of the sister chromatids of each chromosome is generally shown for brevity, and in reality they are generally so close together that they look as one on photomicrographs as well u ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
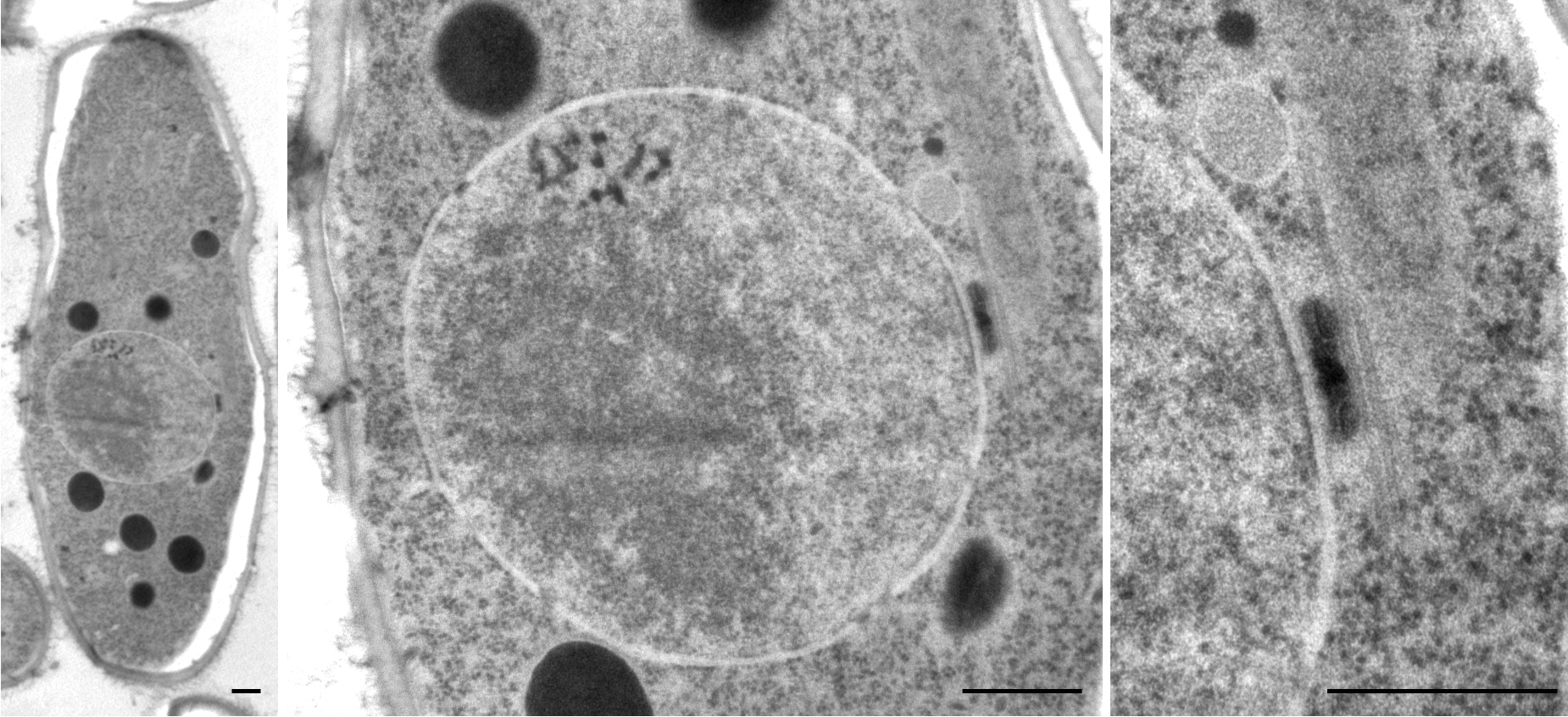 |
Schizosaccharomyces Pombe
''Schizosaccharomyces pombe'', also called "fission yeast", is a species of yeast used in traditional brewing and as a model organism in molecular and cell biology. It is a unicellular eukaryote, whose cells are rod-shaped. Cells typically measure 3 to 4 micrometres in diameter and 7 to 14 micrometres in length. Its genome, which is approximately 14.1 million base pairs, is estimated to contain 4,970 protein-coding genes and at least 450 non-coding RNAs. These cells maintain their shape by growing exclusively through the cell tips and divide by medial fission to produce two daughter cells of equal size, which makes them a powerful tool in cell cycle research. Fission yeast was isolated in 1893 by Paul Lindner from East African millet beer. The species name ''pombe'' is the Swahili word for beer. It was first developed as an experimental model in the 1950s: by Urs Leupold for studying genetics, and by Murdoch Mitchison for studying the cell cycle. Paul Nurse, a fission yeast r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
|
Cohesin
Cohesin is a protein complex that mediates sister chromatid cohesion, homologous recombination, and DNA looping. Cohesin is formed of SMC3, SMC1, SCC1 and SCC3 ( SA1 or SA2 in humans). Cohesin holds sister chromatids together after DNA replication until anaphase when removal of cohesin leads to separation of sister chromatids. The complex forms a ring-like structure and it is believed that sister chromatids are held together by entrapment inside the cohesin ring. Cohesin is a member of the SMC family of protein complexes which includes Condensin, MukBEF and SMC-ScpAB. Cohesin was separately discovered in budding yeast by Douglas Koshland and Kim Nasmyth. Structure Cohesin is a multi-subunit protein complex, made up of SMC1, SMC3, RAD21 and SCC3 (SA1 or SA2). SMC1 and SMC3 are members of the Structural Maintenance of Chromosomes (SMC) family. SMC proteins have two main structural characteristics: an ATP-binding cassette-like 'head' domain with ATPase activity ( ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
|
|
Human Genome
The human genome is a complete set of nucleic acid sequences for humans, encoded as DNA within the 23 chromosome pairs in cell nuclei and in a small DNA molecule found within individual mitochondria. These are usually treated separately as the nuclear genome and the mitochondrial genome. Human genomes include both protein-coding DNA sequences and various types of DNA that does not encode proteins. The latter is a diverse category that includes DNA coding for non-translated RNA, such as that for ribosomal RNA, transfer RNA, ribozymes, small nuclear RNAs, and several types of regulatory RNAs. It also includes promoters and their associated gene-regulatory elements, DNA playing structural and replicatory roles, such as scaffolding regions, telomeres, centromeres, and origins of replication, plus large numbers of transposable elements, inserted viral DNA, non-functional pseudogenes and simple, highly-repetitive sequences. Introns make up a large percentage of non-coding DN ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
|
|
Chromosome 13 (human)
Chromosome 13 is one of the 23 pairs of chromosomes in humans. People normally have two copies of this chromosome. Chromosome 13 spans about 114 million base pairs (the building material of DNA) and represents between 3.5 and 4% of the total DNA in cells. Genes Number of genes The following are some of the gene count estimates of human chromosome 13. Because researchers use different approaches to genome annotation their predictions of the number of genes on each chromosome varies (for technical details, see gene prediction). Among various projects, the collaborative consensus coding sequence project ( CCDS) takes an extremely conservative strategy. So CCDS's gene number prediction represents a lower bound on the total number of human protein-coding genes. Gene list The following is a partial list of genes on human chromosome 13. For complete list, see the link in the infobox on the right. Diseases and disorders The following diseases and disorders are some of those ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |