|
CAPP-Seq
CAncer Personalized Profiling by deep Sequencing (CAPP-Seq) is a next-generation sequencing based method used to quantify circulating DNA in cancer (ctDNA). The method was introduced in 2014 by Ash Alizadeh and Maximilian Diehn’s laboratories at Stanford, as a tool for measuring Cell-free tumor DNA which is released from dead tumor cells into the blood and thus may reflect the entire tumor genome. This method can be generalized for any cancer type that is known to have recurrent mutations. CAPP-Seq can detect one molecule of mutant DNA in 10,000 molecules of healthy DNA. The original method was further refined in 2016 for ultra sensitive detection through integration of multiple error suppression strategies, termed integrated Digital Error Suppression (iDES). The use of ctDNA in this technique should not be confused with circulating tumor cells (CTCs); these are two different entities. Originally described as a method to detect and monitor lung cancers, CAPP-Seq has been succes ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell-free Tumour DNA
Circulating tumor DNA (ctDNA) is tumor-derived fragmented DNA in the bloodstream that is not associated with cells. ctDNA should not be confused with cell-free DNA (cfDNA), a broader term which describes DNA that is freely circulating in the bloodstream, but is not necessarily of tumor origin. Because ctDNA may reflect the entire tumor genome, it has gained traction for its potential clinical utility; " liquid biopsies" in the form of blood draws may be taken at various time points to monitor tumor progression throughout the treatment regimen. ctDNA originates directly from the tumor or from circulating tumor cells (CTCs), which describes viable, intact tumor cells that shed from primary tumors and enter the bloodstream or lymphatic system. The precise mechanism of ctDNA release is unclear. The biological processes postulated to be involved in ctDNA release include apoptosis and necrosis from dying cells, or active release from viable tumor cells. Studies in both human (healthy ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Circulating Tumor DNA
Circulating tumor DNA (ctDNA) is tumor-derived fragmented DNA in the bloodstream that is not associated with cells. ctDNA should not be confused with cell-free DNA (cfDNA), a broader term which describes DNA that is freely circulating in the bloodstream, but is not necessarily of tumor origin. Because ctDNA may reflect the entire tumor genome, it has gained traction for its potential clinical utility; "liquid biopsies" in the form of blood draws may be taken at various time points to monitor tumor progression throughout the treatment regimen. ctDNA originates directly from the tumor or from circulating tumor cells (CTCs), which describes viable, intact tumor cells that shed from primary tumors and enter the bloodstream or lymphatic system. The precise mechanism of ctDNA release is unclear. The biological processes postulated to be involved in ctDNA release include apoptosis and necrosis from dying cells, or active release from viable tumor cells. Studies in both human (healthy an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Recurrence Index
Recurrence Index (RI) in genomics denotes the number of mutations per kilobase of a given genomic locus of a patient carrying particular mutations. RI represents a patient level recurrence frequency estimated for somatic mutations. References {{reflist Genomics ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Copy Number Variations
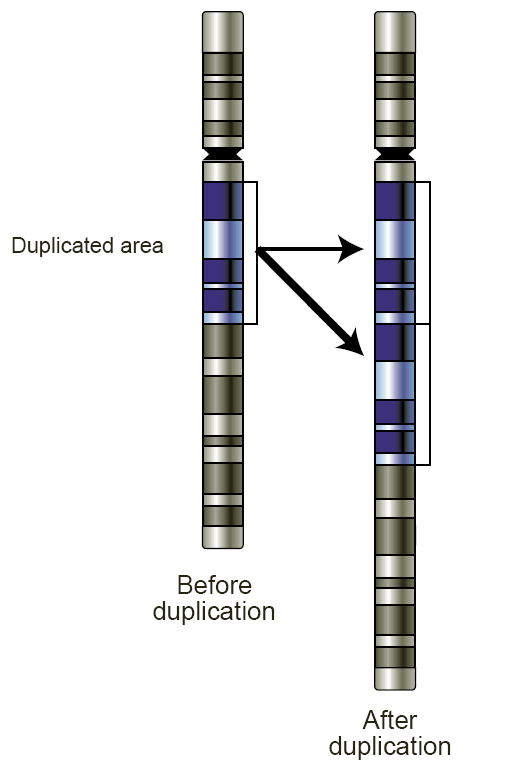
Copy number variation (CNV) is a phenomenon in which sections of the genome are repeated and the number of repeats in the genome varies between individuals. Copy number variation is a type of structural variation: specifically, it is a type of duplication or deletion event that affects a considerable number of base pairs. Approximately two-thirds of the entire human genome may be composed of repeats and 4.8–9.5% of the human genome can be classified as copy number variations. In mammals, copy number variations play an important role in generating necessary variation in the population as well as disease phenotype. Copy number variations can be generally categorized into two main groups: short repeats and long repeats. However, there are no clear boundaries between the two groups and the classification depends on the nature of the loci of interest. Short repeats include mainly dinucleotide repeats (two repeating nucleotides e.g. A-C-A-C-A-C...) and trinucleotide repeats. Long r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Structural Variations
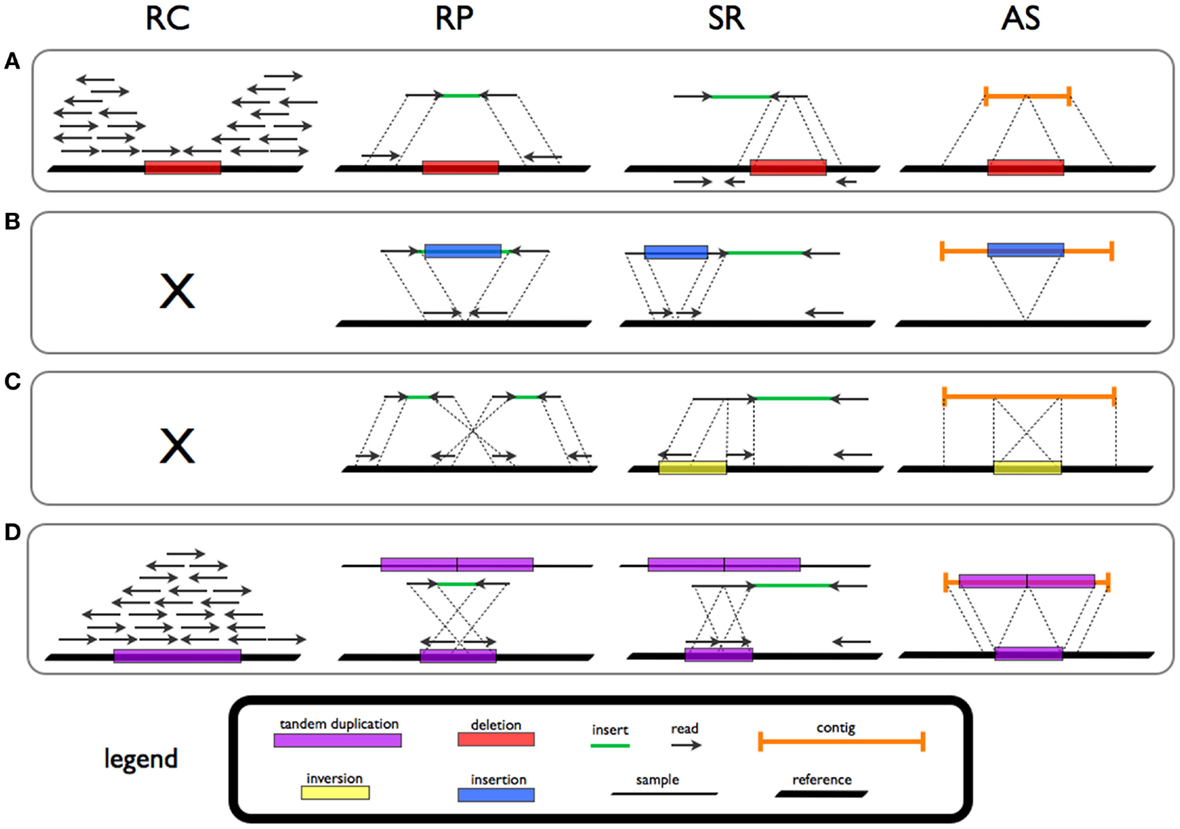
Genomic structural variation is the variation in structure of an organism's chromosome. It consists of many kinds of variation in the genome of one species, and usually includes microscopic and submicroscopic types, such as deletions, duplications, copy-number variants, insertions, inversions and translocations. Originally, a structure variation affects a sequence length about 1kb to 3Mb, which is larger than SNPs and smaller than chromosome abnormality (though the definitions have some overlap). However, the operational range of structural variants has widened to include events > 50bp. The definition of structural variation does not imply anything about frequency or phenotypical effects. Many structural variants are associated with genetic diseases, however many are not. Recent research about SVs indicates that SVs are more difficult to detect than SNPs. Approximately 13% of the human genome is defined as structurally variant in the normal population, and there are at least ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Indels
Indel is a molecular biology term for an insertion or deletion of bases in the genome of an organism. It is classified among small genetic variations, measuring from 1 to 10 000 base pairs in length, including insertion and deletion events that may be separated by many years, and may not be related to each other in any way. A microindel is defined as an indel that results in a net change of 1 to 50 nucleotides. In coding regions of the genome, unless the length of an indel is a multiple of 3, it will produce a frameshift mutation. For example, a common microindel which results in a frameshift causes Bloom syndrome in the Jewish or Japanese population. Indels can be contrasted with a point mutation. An indel inserts or deletes nucleotides from a sequence, while a point mutation is a form of substitution that ''replaces'' one of the nucleotides without changing the overall number in the DNA. Indels can also be contrasted with Tandem Base Mutations (TBM), which may result from ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Amplicon
In molecular biology, an amplicon is a piece of DNA or RNA that is the source and/or product of amplification or replication events. It can be formed artificially, using various methods including polymerase chain reactions (PCR) or ligase chain reactions (LCR), or naturally through gene duplication. In this context, ''amplification'' refers to the production of one or more copies of a genetic fragment or target sequence, specifically the amplicon. As it refers to the product of an amplification reaction, ''amplicon'' is used interchangeably with common laboratory terms, such as "PCR product." Artificial amplification is used in research, forensics, and medicine for purposes that include detection and quantification of infectious agents, identification of human remains, and extracting genotypes from human hair. Natural gene duplication plays a major role in evolution. It is also implicated in several forms of human cancer including primary mediastinal B cell lymphoma and Hodgkin ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Digital Polymerase Chain Reaction
Digital polymerase chain reaction (digital PCR, DigitalPCR, dPCR, or dePCR) is a biotechnological refinement of conventional polymerase chain reaction methods that can be used to directly quantify and clonally amplify nucleic acids strands including DNA, cDNA, or RNA. The key difference between dPCR and traditional PCR lies in the method of measuring nucleic acids amounts, with the former being a more precise method than PCR, though also more prone to error in the hands of inexperienced users. A "digital" measurement quantitatively and discretely measures a certain variable, whereas an “analog” measurement extrapolates certain measurements based on measured patterns. PCR carries out one reaction per single sample. dPCR also carries out a single reaction within a sample, however the sample is separated into a large number of partitions and the reaction is carried out in each partition individually. This separation allows a more reliable collection and sensitive measurement of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Metastatic Disease
Metastasis is a pathogenic agent's spread from an initial or primary site to a different or secondary site within the host's body; the term is typically used when referring to metastasis by a cancerous tumor. The newly pathological sites, then, are metastases (mets). It is generally distinguished from cancer invasion, which is the direct extension and penetration by cancer cells into neighboring tissues. Cancer occurs after cells are genetically altered to proliferate rapidly and indefinitely. This uncontrolled proliferation by mitosis produces a primary heterogeneic tumour. The cells which constitute the tumor eventually undergo metaplasia, followed by dysplasia then anaplasia, resulting in a malignant phenotype. This malignancy allows for invasion into the circulation, followed by invasion to a second site for tumorigenesis. Some cancer cells known as circulating tumor cells acquire the ability to penetrate the walls of lymphatic or blood vessels, after which they are able t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Peripheral Blood Lymphocytes
Peripheral blood lymphocytes (PBL) are mature lymphocytes that circulate in the blood, rather than localising to organs (such as the spleen or lymph nodes). They comprise T cells, NK cells and B cells B cells, also known as B lymphocytes, are a type of white blood cell of the lymphocyte subtype. They function in the humoral immunity component of the adaptive immune system. B cells produce antibody molecules which may be either secreted or .... References Lymphocytes {{lymphatic-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at Cetus Corporation; Mullis and biochemist Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |