Nonsense Mediated Decay on:
[Wikipedia]
[Google]
[Amazon]
 Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all
Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all
 Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all
Nonsense-mediated mRNA decay (NMD) is a surveillance pathway that exists in all eukaryotes
The eukaryotes ( ) constitute the domain of Eukaryota or Eukarya, organisms whose cells have a membrane-bound nucleus. All animals, plants, fungi, seaweeds, and many unicellular organisms are eukaryotes. They constitute a major group of ...
. Its main function is to reduce errors in gene expression by eliminating mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
transcripts that contain premature stop codon
In molecular biology, a stop codon (or termination codon) is a codon (nucleotide triplet within messenger RNA) that signals the termination of the translation process of the current protein. Most codons in messenger RNA correspond to the additio ...
s. Translation of these aberrant mRNAs could, in some cases, lead to deleterious gain-of-function or dominant-negative activity of the resulting proteins.
NMD was first described in human cells and in yeast almost simultaneously in 1979. This suggested broad phylogenetic conservation and an important biological role of this intriguing mechanism. NMD was discovered when it was realized that cells often contain unexpectedly low concentrations of mRNAs that are transcribed from alleles carrying nonsense mutation
In genetics, a nonsense mutation is a point mutation in a sequence of DNA that results in a ''nonsense codon'', or a premature stop codon in the transcribed mRNA, and leads to a truncated, incomplete, and possibly nonfunctional protein product. No ...
s. Nonsense mutations code for a premature stop codon which causes the protein to be shortened. The truncated protein may or may not be functional, depending on the severity of what is not translated. In human genetics, NMD has the possibility to not only limit the translation of abnormal proteins, but it can occasionally cause detrimental effects in specific genetic mutations.
NMD functions to regulate numerous biological functions in a diverse range of cells, including the synaptic plasticity of neurons which may shape adult behavior.
Pathway
While many of the proteins involved in NMD are not conserved between species, in ''Saccharomyces cerevisiae'' (yeast), there are three main factors in NMD:UPF1
Regulator of nonsense transcripts 1 (or Up-frameshift suppressor 1 homolog) is a protein that in humans is encoded by the ''UPF1'' gene.
Function
This gene encodes a protein that is part of a post-splicing multiprotein complex, the exon junct ...
, UPF2
Regulator of nonsense transcripts 2 is a protein that in humans is encoded by the ''UPF2'' gene.
Function
This gene encodes a protein that is part of a post-splicing multiprotein complex, the exon junction complex, involved in both mRNA nucle ...
and UPF3 (UPF3A
Regulator of nonsense transcripts 3A is a protein that in humans is encoded by the ''UPF3A'' gene.
This gene encodes a protein that is part of a post-splicing multiprotein complex involved in both mRNA nuclear export and mRNA surveillance. The en ...
and UPF3B in humans), that make up the conserved core of the NMD pathway. All three of these factors are trans-acting In the field of molecular biology, ''trans''-acting (''trans''-regulatory, ''trans''-regulation), in general, means "acting from a different molecule" (''i.e.'', intermolecular). It may be considered the opposite of ''cis''-acting (''cis''-regula ...
elements called up-frameshift (UPF) proteins. In mammals, UPF2 and UPF3 are part of the exon-exon junction complex (EJC) bound to mRNA after splicing along with other proteins, eIF4AIII, MLN51, and the Y14/MAGOH heterodimer, which also function in NMD. UPF1 phosphorylation is controlled by the proteins SMG-1, SMG-5, SMG-6 and SMG-7.
The process of detecting aberrant transcripts occurs during translation
Translation is the communication of the semantics, meaning of a #Source and target languages, source-language text by means of an Dynamic and formal equivalence, equivalent #Source and target languages, target-language text. The English la ...
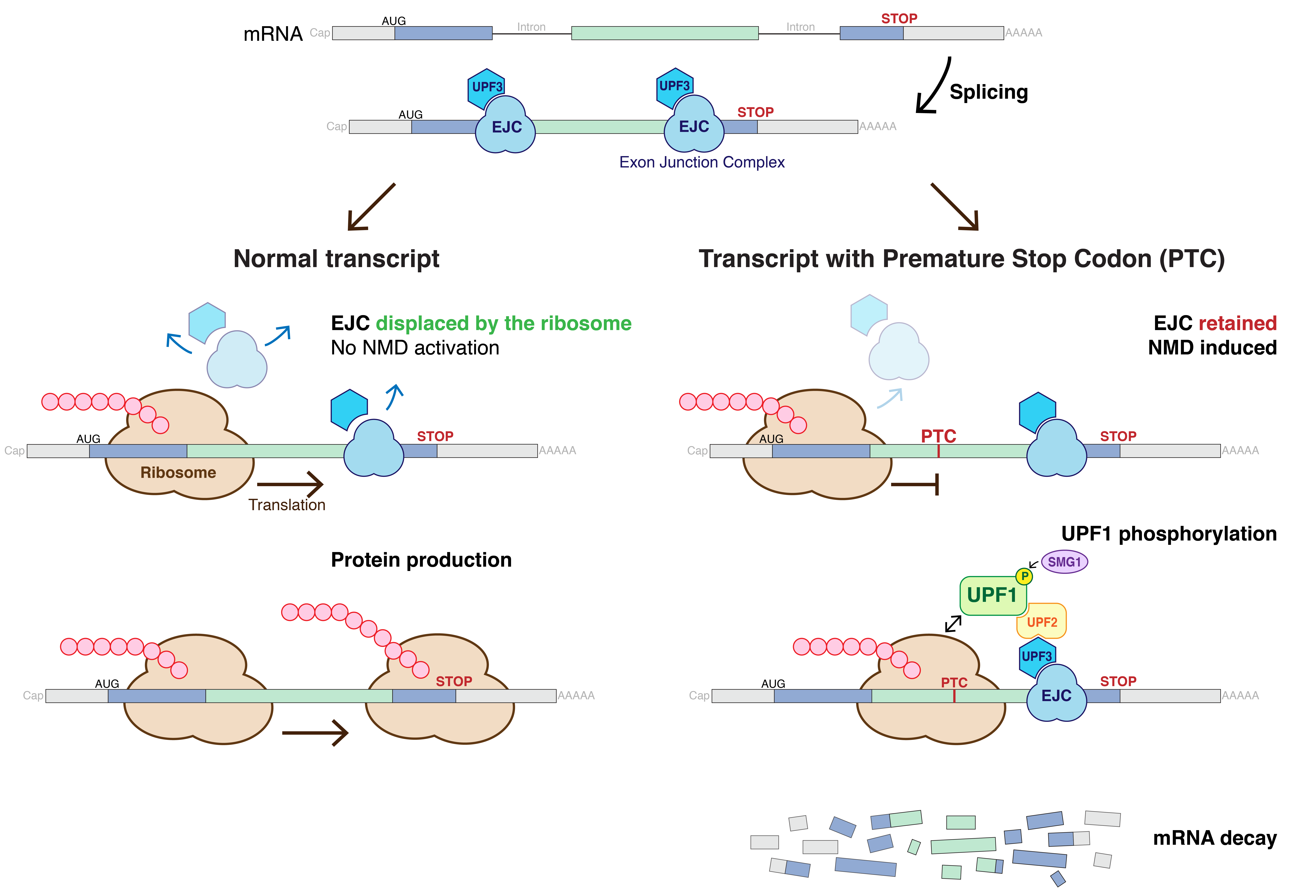
of the mRNA. A popular model for the detection of aberrant transcripts in mammals suggests that during the first round of translation, the ribosome
Ribosomes () are molecular machine, macromolecular machines, found within all cell (biology), cells, that perform Translation (biology), biological protein synthesis (messenger RNA translation). Ribosomes link amino acids together in the order s ...
removes the exon-exon junction complexes bound to the mRNA after splicing occurs. If after this first round of translation, any of these proteins remain bound to the mRNA, NMD is activated. Exon-exon junction complexes located downstream of a stop codon
In molecular biology, a stop codon (or termination codon) is a codon (nucleotide triplet within messenger RNA) that signals the termination of the translation process of the current protein. Most codons in messenger RNA correspond to the additio ...
are not removed from the transcript because the ribosome is released before reaching them. Termination of translation leads to the assembly of a complex composed of UPF1, SMG1 and the release factors, eRF1 and eRF3, on the mRNA. If an EJC is left on the mRNA because the transcript contains a premature stop codon, then UPF1 comes into contact with UPF2 and UPF3, triggering the phosphorylation
In biochemistry, phosphorylation is described as the "transfer of a phosphate group" from a donor to an acceptor. A common phosphorylating agent (phosphate donor) is ATP and a common family of acceptor are alcohols:
:
This equation can be writ ...
of UPF1. In vertebrates, the location of the last exon-junction complex relative to the termination codon usually determines whether the transcript will be subjected to NMD or not. If the termination codon is downstream of or within about 50 nucleotides of the final exon-junction complex then the transcript is translated normally. However, if the termination codon is further than about 50 nucleotides upstream of any exon-junction complexes, then the transcript is down regulated by NMD. The phosphorylated UPF1 then interacts with SMG-5, SMG-6 and SMG-7, which promote the dephosphorylation of UPF1. SMG-7 is thought to be the terminating effector in NMD, as it accumulates in P-bodies
In cellular biology, P-bodies, or processing bodies, are distinct foci formed by phase separation within the cytoplasm of a eukaryotic cell consisting of many enzymes involved in mRNA turnover. P-bodies are highly conserved structures and have ...
, which are cytoplasmic sites for mRNA decay. In both yeast and human cells, the major pathway for mRNA decay is initiated by the removal of the 5’ cap followed by degradation by XRN1, an exoribonuclease enzyme. The other pathway by which mRNA is degraded is by deadenylation
Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In euka ...
from 3’-5'.
In addition to the well recognized role of NMD in removing aberrant transcripts, there are transcripts that contain introns within their 3' untranslated regions (UTRs). These messages are predicted to be NMD-targets yet they (e.g., activity-regulated cytoskeleton-associated protein, known as Arc) can play crucial biologic functions suggesting that NMD may have physiologically relevant roles.
Mechanism and regulation
NMD is a cellular mechanism that degrades mRNAs containing premature termination codons (PTCs), which can arise from mutations. Comprehensive analyses large scale genetics and gene expression datasets have enabled the systemic identification of the mechanism of NMD and its efficiency. # EJC model: NMD is typically triggered when a PTC is located upstream of the last exon junction complex (EJC). If the PTC is downstream of the last EJC, NMD is often inefficient. # Start-proximal effect: PTCs located near the start codon can evade NMD. This evasion is associated with the presence of downstream in-frame stop codons, which can allow the ribosome to bypass the PTC and continue translation. # Exon length and distance to normal stop codon: Long exons and large distances between the PTC and the normal stop codon are associated with inefficient NMD. This suggests that the spatial configuration of the mRNA can influence the accessibility of NMD machinery. # mRNA turnover rate: Transcripts with rapid turnover rates tend to attenuate the effects of NMD. This means that mRNAs that are quickly degraded by other mechanisms may not be efficiently targeted by NMD. # RNA-binding protein motifs: Certain RNA-binding protein motifs near the PTC or within the 3′UTR can modulate NMD efficiency. These motifs can either enhance or inhibit the recognition of PTCs by NMD machinery, depending on their specific interactions with NMD factors. # Generalized frameshift. As a response to poor nutritional conditions, the bulk of theyeast
Yeasts are eukaryotic, single-celled microorganisms classified as members of the fungus kingdom (biology), kingdom. The first yeast originated hundreds of millions of years ago, and at least 1,500 species are currently recognized. They are est ...
transcriptome undergoes −1 ribosome frameshifts which leads to an accelerated co-translational mRNA decay. Under such conditions NMD-dependent degradation represents at least one-third of the total mRNA decay. Less optimal codons are a key factor for ribosomes to induce out-of-frame mRNA decay. This mechanism appears to be conserved from bacteria
Bacteria (; : bacterium) are ubiquitous, mostly free-living organisms often consisting of one Cell (biology), biological cell. They constitute a large domain (biology), domain of Prokaryote, prokaryotic microorganisms. Typically a few micr ...
to humans
Humans (''Homo sapiens'') or modern humans are the most common and widespread species of primate, and the last surviving species of the genus ''Homo''. They are Hominidae, great apes characterized by their Prehistory of nakedness and clothing ...
.
Mutations
Although nonsense-mediated mRNA decay reduces nonsense codons, mutations can occur that lead to various health problems and diseases in humans. A dominant-negative or deleterious gain-of-function mutation can occur if premature terminating (nonsense) codons are translated. NMD is becoming increasingly evident in the way it modifies phenotypic consequences because of the broad way it controls gene expression. For instance, the blood disorderBeta thalassemia
Beta-thalassemia (β-thalassemia) is an genetic disorder, inherited hemoglobinopathy, blood disorder, a form of thalassemia resulting in variable outcomes ranging from clinically asymptomatic to severe anemia individuals. It is caused by reduce ...
is inherited and caused by mutations within the upstream region of the β-globin gene. An individual carrying only one affected allele will have no or extremely low levels of the mutant β-globin mRNA. An even more severe form of the disease can occur called thalassemia intermedia or ‘inclusion body’ thalassemia. Instead of decreased mRNA levels, a mutant transcript produces truncated β chains, which in turn leads to a clinical phenotype in the heterozygote.
Nonsense-mediated decay mutations can also contribute to Marfan syndrome
Marfan syndrome (MFS) is a multi-systemic genetic disorder that affects the connective tissue. Those with the condition tend to be tall and thin, with dolichostenomelia, long arms, legs, Arachnodactyly, fingers, and toes. They also typically ha ...
. This disorder is caused by mutations in the fibrillin 1 (FBN1) gene and is resulted from a dominant negative interaction between mutant and wild-type fibrillin-1 gene.
Regulating immunogenic frameshift-derived antigens
NMD plays a role in the regulation of immunogenic frameshift-derived antigens. Frameshift mutations often result in the production of aberrant proteins that can be recognized as neoantigens by the immune system, particularly in cancer cells. However, frameshift mutations often lead to the translation of an out-of-frame PTC that can activate NMD to degrade these mutant mRNAs before they are translated into proteins, thereby reducing the presentation of these potentially immunogenic peptides on the cell surface via HLA class I molecules. This modulation of immunogenicity means that frameshift-derived neoantigens only contribute to the response to immune checkpoint inhibition if they arise from mutations in parts of the genome that are not recognized by NMD.Research applications
This pathway has a significant effect in the way genes are translated, restricting the amount of gene expression. It is still a new field in genetics, but its role in research has already led scientists to uncover numerous explanations for gene regulation. Studying nonsense-mediated decay has allowed scientists to determine the causes for certain heritable diseases and dosage compensation in mammals.Heritable diseases
Theproopiomelanocortin
Pro-opiomelanocortin (POMC) is a precursor polypeptide with 241 amino acid residues. POMC is Protein biosynthesis, synthesized in Corticotropic cell, corticotrophs of the anterior pituitary from the 267-amino-acid-long Precursor polypeptide, pol ...
gene (POMC) is expressed in the hypothalamus, in the pituitary gland. It yields a range of biologically active peptides and hormones and undergoes tissue-specific posttranslational processing to yield a range of biologically active peptides producing adrenocorticotropic hormone (ACTH), b-endorphin, and a-, b- and c-melanocyte-stimulating hormone (MSH). These peptides then interact with different melanocortin receptors (MCRs) and are involved in a wide range of processes including the regulation of body weight (MC3R and MC4R), adrenal steroidogenesis (MC2R) and hair pigmentation (MC1R). Published in the British Associations of Dermatologists in 2012, Lack of red hair phenotype in a North-African obese child homozygous for a novel POMC null mutation showed nonsense-mediated decay RNA evaluation in a hair pigment chemical analysis. They found that inactivating the POMC gene mutation results in obesity, adrenal insufficiency, and red hair. This has been seen in both in humans and mice. In this experiment they described a 3-year-old boy from Rome, Italy. He was a source of focus because he had Addison's disease
Addison's disease, also known as primary adrenal insufficiency, is a rare long-term endocrine disorder characterized by inadequate production of the steroid hormones cortisol and aldosterone by the two outer layers of the cells of the adr ...
and early onset obesity. They collected his DNA and amplified it using PCR. Sequencing analysis revealed a homozygous single substitution determining a stop codon. This caused an aberrant protein and the corresponding amino acid sequence indicated the exact position of the homozygous nucleotide. The substitution was localized in exon 3 and nonsense mutation at codon 68. The results from this experiment strongly suggest that the absence of red hair in non-European patients with early onset obesity and hormone deficiency does not exclude the occurrence of POMC mutations. By sequencing the patients DNA they found that this novel mutation has a collection of symptoms because of a malfunctioning nonsense-mediated mRNA decay surveillance pathway.
Dosage compensation
There has been evidence that the nonsense-mediated mRNA decay pathway participates in X chromosome dosage compensation in mammals. In higher eukaryotes with dimorphic sex chromosomes, such as humans and fruit flies, males have oneX chromosome
The X chromosome is one of the two sex chromosomes in many organisms, including mammals, and is found in both males and females. It is a part of the XY sex-determination system and XO sex-determination system. The X chromosome was named for its u ...
, whereas females have two. These organisms have evolved a mechanism that compensates not only for the different number of sex chromosomes between the two sexes, but also for the differing X/autosome ratios. In this genome-wide survey, the scientists found that autosomal genes are more likely to undergo nonsense-mediated decay than X-linked genes. This is because NMD fine tunes X chromosomes and it was demonstrated by inhibiting the pathway. The results were that balanced gene expression between X and autosomes gene expression was decreased by 10–15% no matter the method of inhibition. The NMD pathway is skewed towards depressing expression of larger population or autosomal genes than x-linked ones. In conclusion, the data supports the view that the coupling of alternative splicing and NMD is a pervasive means of gene expression regulation.
Designing CRISPR-Cas9 experiments
The implications of NMD are significant when designing CRISPR-Cas9 experiments, particularly those aimed at gene inactivation. CRISPR-Cas9 introduces double-strand breaks that can lead to insertions or deletions (indels), often resulting in frameshift mutations and PTCs. If these PTCs are located in regions that trigger NMD, the resulting mRNAs will be rapidly degraded, leading to effective gene knockdown. However, if the PTCs are in regions that evade NMD, the mutant mRNAs may be translated into truncated proteins, potentially retaining partial function and leading to incomplete gene inactivation. Therefore, understanding and incorporating NMD rules into the design of single guide RNAs (sgRNAs) is essential for achieving desired outcomes in CRISPR-Cas9 experiments. Tools such as NMDetective can predict the likelihood of NMD triggering based on the location of PTCs, thereby aiding in the design of more effective gene-editing strategies.See also
*Non-stop decay
Non-stop decay (NSD) is a cellular mechanism of mRNA surveillance to detect mRNA molecules lacking a stop codon and prevent these mRNAs from translation. The non-stop decay pathway releases ribosomes that have reached the far 3' end of an mRNA and ...
, another mRNA surveillance mechanism
* mRNA surveillance
mRNA surveillance mechanisms are pathways utilized by organisms to ensure fidelity and quality of messenger RNA (mRNA) molecules. There are a number of surveillance mechanisms present within cells. These mechanisms function at various steps of th ...
References
Further reading
* * * * * * * *External links
* {{cite web , url = https://archive.today/20121228092407/http://db.yeastgenome.org/cgi-bin/GO/go.pl?goid=184 , title = Yeast mRNA catabolism , work = Saccharomyces Genome Database , publisher = Stanford University Gene expression