|
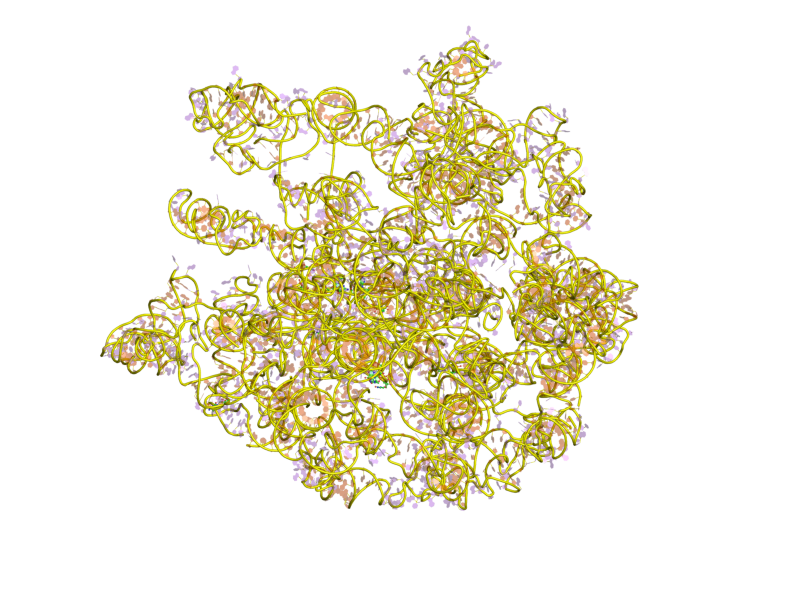
RNase E
Ribonuclease E is a bacterial ribonuclease that participates in the processing of ribosomal RNA (9S to 5S rRNA) and the chemical degradation of bulk cellular RNA. Cellular localization RNase E was suggested to be a part of the cell membrane protein complex, as it sediments with ribosomes and crude membranes. Microscopy has localized tagged RNase E to the inner cytoplasmic membrane or a helical cytoskeletal structure closely associated with the inner layer. Protein structure This enzyme contains 1,061 residues and separates into two distinct functional regions, which are a large domain located at the 5'N-terminus and a small domain located at the 3' C-terminus. While N-terminal half forms a catalytic domain, C-terminal half forms a degradosome scaffolding domain. A metal-binding pocket separates them in the middle of the RNase E protein structure. Although degradosome formation does not play a key role for E. coli growth, the deletion of the C-terminal half has been found to ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
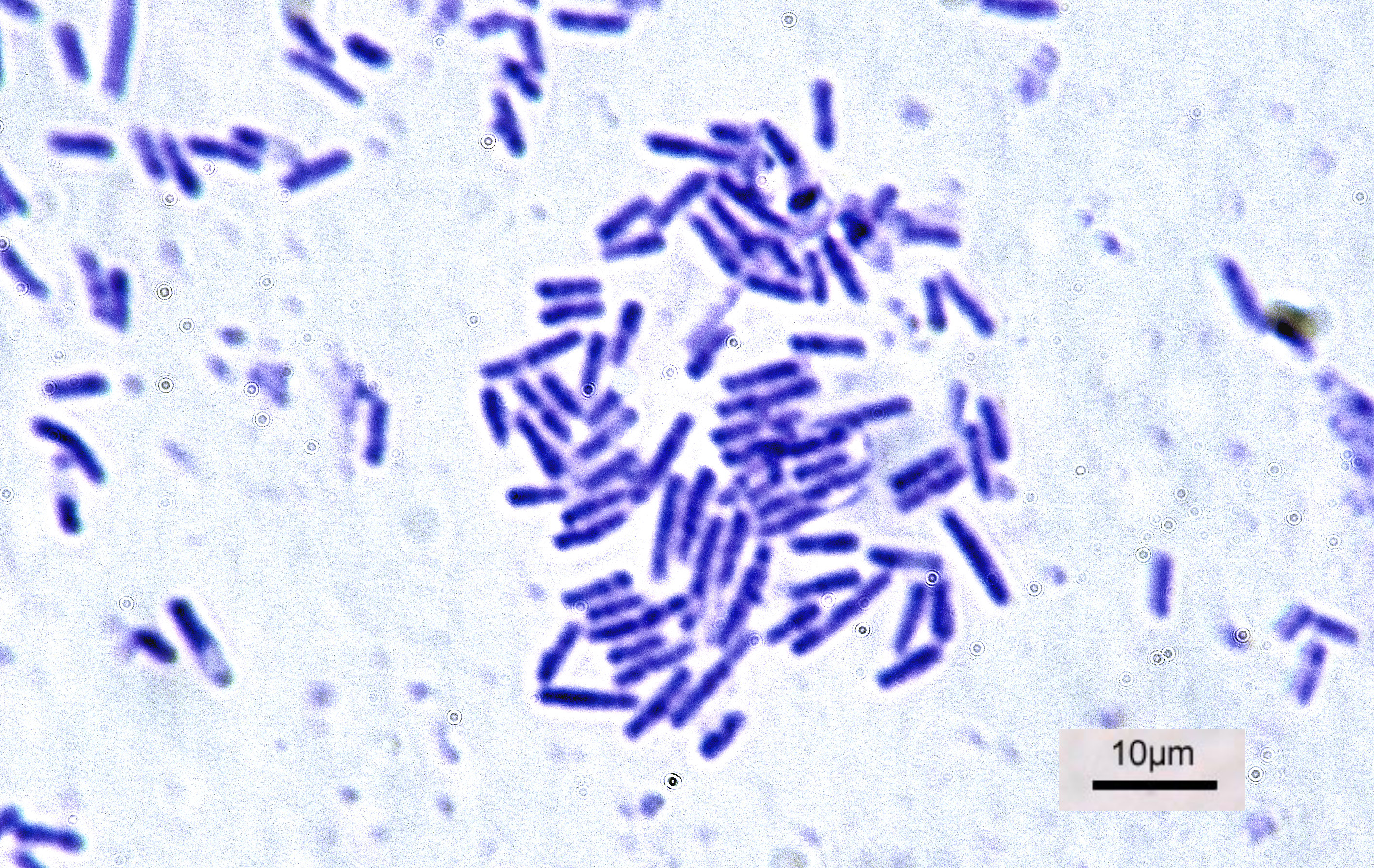
Bacterial
Bacteria (; : bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit the air, soil, water, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria play a vital role in many stages of the nutrient cycle by recycling nutrients and the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition of dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. Bacteria also live in mutualistic, commensal an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and the ribosome creates the protein utilizing amino acids carried by transfer RNA (tRNA). This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, genetic inf ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Homology
Sequence homology is the homology (biology), biological homology between DNA sequence, DNA, RNA sequence, RNA, or Protein primary structure, protein sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry because of three phenomena: either a speciation event (orthologs), or a Gene duplication, duplication event (paralogs), or else a Horizontal gene transfer, horizontal (or lateral) gene transfer event (xenologs). Homology among DNA, RNA, or proteins is typically inferred from their nucleotide or amino acid sequence similarity. Significant similarity is strong evidence that two sequences are related by evolutionary changes from a common ancestral sequence. Sequence alignment, Alignments of multiple sequences are used to indicate which regions of each sequence are homologous. Identity, similarity, and conservation The term "percent homology" is often used to mean "sequence similarity”, that is the percen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exoribonuclease
An exoribonuclease is an exonuclease ribonuclease, which are enzymes that degrade RNA by removing terminal nucleotides from either the 5' end or the 3' end of the RNA molecule. Enzymes that remove nucleotides from the 5' end are called ''5'-3' exoribonucleases'', and enzymes that remove nucleotides from the 3' end are called ''3'-5' exoribonucleases''. Exoribonucleases can use either water to cleave the nucleotide-nucleotide bond (which is called hydrolytic activity) or inorganic phosphate (which is called phosphorolytic activity). Hydrolytic exoribonucleases are classified under EC number 3.1 and phosphorolytic exoribonucleases under EC number 2.7.7. As the phosphorolytic enzymes use inorganic phosphate to cleave bonds they release nucleotide diphosphates, whereas the hydrolytic enzymes (which use water) release nucleotide monosphosphates. Exoribonucleases exist in all kingdoms of life, the bacteria, archaea, and eukaryotes. Exoribonucleases are involved in the degradati ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cedecea
''Cedecea'' is a genus of extremely rare bacteria of the family Enterobacteriaceae. The name of this genus was derived from CDC, the abbreviation for the Centers for Disease Control where the initial members of this genus were discovered. This genus resembles no other group of Enterobacteriaceae.Grimont, P. A. D., Grimont, F., Farmer III, J. J., and Asbury, M. A. (1981)''Cedecea davisae'' gen. nov., sp. nov. and ''Cedecea lapagei'' sp. nov., New ''Enterobacteriaceae'' from clinical specimens.''International Journal of Systematic Bacteriology'' 31, 317-326. ''Cedecea'' bacteria are Gram-negative, bacillus in shape, motile, nonencapsulated, and non-spore-forming.Mawardi, H., Pavlakis, M., Mandelbrot, D., Woo, S. B. (2010) Sirolimus oral ulcer with ''Cedecea davisae'' superinfection.''Transpl Infect Dis'' 12, 446-450.Abate,G., Qureshi, S., and Mazumder, S. A. (2011)''Cedecea davisae'' bacteremia in a neutropenic patient with acute myeloid leukemia.''Journal of Infection'' 63, 83-85. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
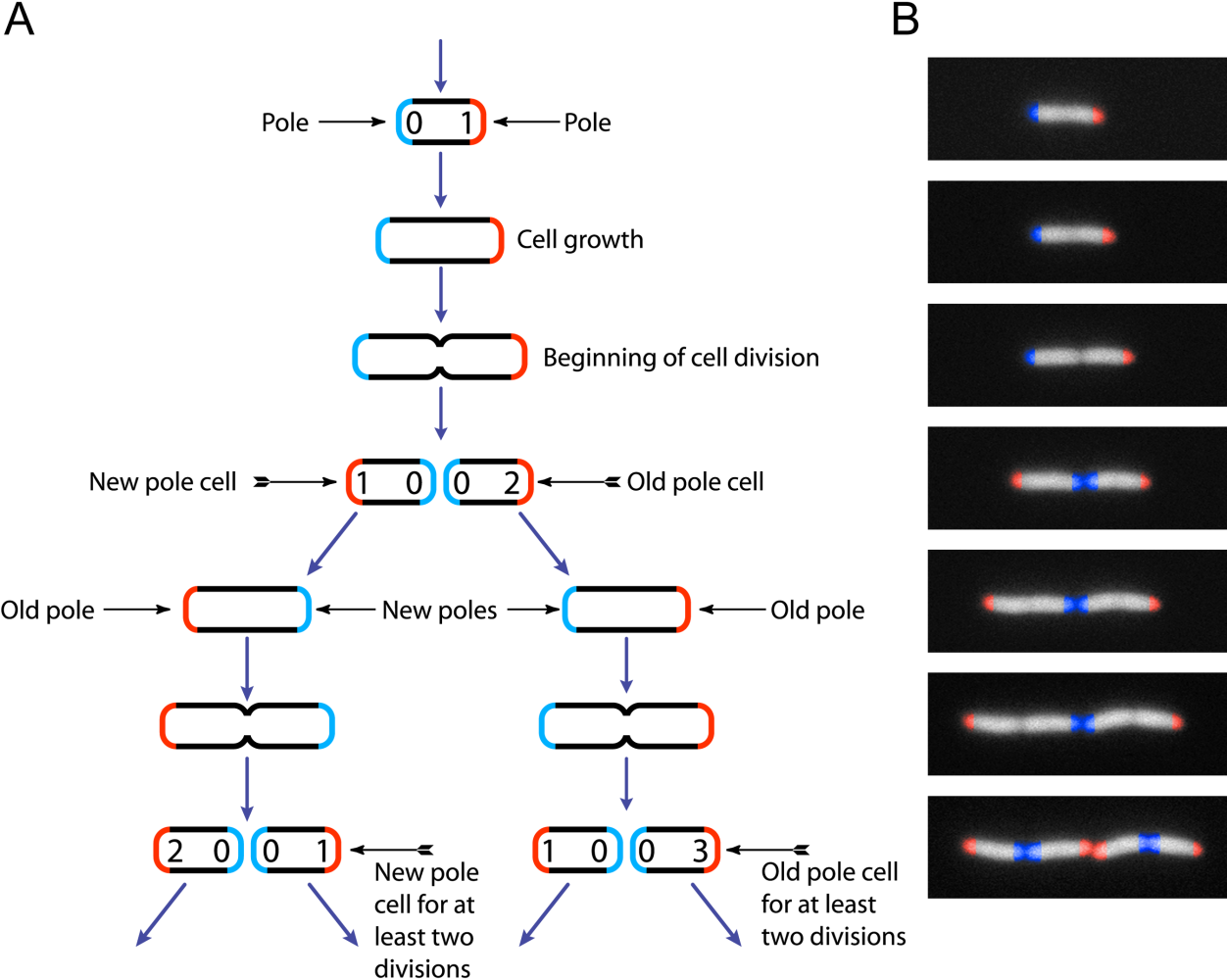
Escherichia Coli
''Escherichia coli'' ( )Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. is a gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus '' Escherichia'' that is commonly found in the lower intestine of warm-blooded organisms. Most ''E. coli'' strains are part of the normal microbiota of the gut, where they constitute about 0.1%, along with other facultative anaerobes. These bacteria are mostly harmless or even beneficial to humans. For example, some strains of ''E. coli'' benefit their hosts by producing vitamin K2 or by preventing the colonization of the intestine by harmful pathogenic bacteria. These mutually beneficial relationships between ''E. coli'' and humans are a type of mutualistic biological relationship—where both the humans and the ''E. coli'' are benefitting each other. ''E. coli'' is expelled into the environment within fecal matter. The bacterium grows massi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleophile
In chemistry, a nucleophile is a chemical species that forms bonds by donating an electron pair. All molecules and ions with a free pair of electrons or at least one pi bond can act as nucleophiles. Because nucleophiles donate electrons, they are Lewis bases. ''Nucleophilic'' describes the affinity of a nucleophile to bond with positively charged Atomic nucleus, atomic nuclei. Nucleophilicity, sometimes referred to as nucleophile strength, refers to a substance's nucleophilic character and is often used to compare the affinity of atoms. Neutral nucleophilic reactions with solvents such as Alcohol (chemistry), alcohols and water are named solvolysis. Nucleophiles may take part in nucleophilic substitution, whereby a nucleophile becomes attracted to a full or partial positive charge, and nucleophilic addition. Nucleophilicity is closely related to basicity. The difference between the two is, that basicity is a thermodynamic property (i.e. relates to an equilibrium state), but nucleop ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hydroxy Group
In chemistry, a hydroxy or hydroxyl group is a functional group with the chemical formula and composed of one oxygen atom covalently bonded to one hydrogen atom. In organic chemistry, alcohols and carboxylic acids contain one or more hydroxy groups. Both the negatively charged anion , called hydroxide, and the neutral radical , known as the hydroxyl radical, consist of an unbonded hydroxy group. According to IUPAC definitions, the term ''hydroxyl'' refers to the hydroxyl radical () only, while the functional group is called a ''hydroxy group''. Properties Water, alcohols, carboxylic acids, and many other hydroxy-containing compounds can be readily deprotonated due to a large difference between the electronegativity of oxygen (3.5) and that of hydrogen (2.1). Hydroxy-containing compounds engage in intermolecular hydrogen bonding increasing the electrostatic attraction between molecules and thus to higher boiling and melting points than found for compounds that lack thi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hydrophobic Effect
The hydrophobic effect is the observed tendency of nonpolar substances to aggregate in an aqueous solution and to be excluded by water. The word hydrophobic literally means "water-fearing", and it describes the segregation of water and nonpolar substances, which maximizes the entropy of water and minimizes the area of contact between water and nonpolar molecules. In terms of thermodynamics, the hydrophobic effect is the free energy change of water surrounding a solute. A positive free energy change of the surrounding solvent indicates hydrophobicity, whereas a negative free energy change implies hydrophilicity. The hydrophobic effect is responsible for the separation of a mixture of oil and water into its two components. It is also responsible for effects related to biology, including: cell membrane and vesicle formation, protein folding, insertion of membrane proteins into the nonpolar lipid environment and protein-small molecule associations. Hence the hydrophobic effect i ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transfer RNA
Transfer ribonucleic acid (tRNA), formerly referred to as soluble ribonucleic acid (sRNA), is an adaptor molecule composed of RNA, typically 76 to 90 nucleotides in length (in eukaryotes). In a cell, it provides the physical link between the genetic code in messenger RNA (mRNA) and the amino acid sequence of proteins, carrying the correct sequence of amino acids to be combined by the protein-synthesizing machinery, the ribosome. Each three-nucleotide codon in mRNA is complemented by a three-nucleotide anticodon in tRNA. As such, tRNAs are a necessary component of translation, the biological synthesis of new proteins in accordance with the genetic code. Overview The process of translation starts with the information stored in the nucleotide sequence of DNA. This is first transformed into mRNA, then tRNA specifies which three-nucleotide codon from the genetic code corresponds to which amino acid. Each mRNA codon is recognized by a particular type of tRNA, which docks to it along ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ribonuclease III
Ribonuclease III (RNase III or RNase C) (BREND3.1.26.3 is a type of ribonuclease that recognizes dsRNA and cleaves it at specific targeted locations to transform them into mature RNAs. These enzymes are a group of endoribonucleases that are characterized by their ribonuclease domain, which is labelled the RNase III domain. They are ubiquitous compounds in the cell and play a major role in pathways such as RNA precursor synthesis, RNA Silencing, and the ''pnp'' autoregulatory mechanism. Types of RNase III The RNase III superfamily is divided into four known classes: 1, 2, 3, and 4. Each class is defined by its domain structure.Liang Y-H, Lavoie M, Comeau M-A, Elela SA, Ji X. Structure of a Eukaryotic RNase III Post-Cleavage Complex Reveals a Double- Ruler Mechanism for Substrate Selection. Molecular cell. 2014;54(3):431-444. doi:10.1016/j.molcel.2014.03.006. Class 1 RNase III *Class 1 RNase III enzymes have a homodimeric structure whose function is to cleave dsRNA into multiple ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
23S Ribosomal RNA
The 23S rRNA is a 2,904 nucleotide long (in ''E. coli'') component of the large subunit (50S) of the bacterial/archean ribosome and makes up the peptidyl transferase center (PTC). The 23S rRNA is divided into six secondary structural domains titled I-VI, with the corresponding 5S rRNA being considered domain VII. The ribosomal peptidyl transferase activity resides in domain V of this rRNA, which is also the most common binding site for antibiotics that inhibit translation, making it a target for ribosomal engineering. A well-known member of this antibiotic class, chloramphenicol, acts by inhibiting peptide bond formation, with recent 3D-structural studies showing two different binding sites depending on the species of ribosome. Numerous mutations in domains of the 23S rRNA with Peptidyl transferase activity have resulted in antibiotic resistance. 23S rRNA genes typically have higher sequence variations, including insertions and/or deletions, compared to other rRNAs. The eukary ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |