Sleeping Beauty Transposon System on:
[Wikipedia]
[Google]
[Amazon]
The ''Sleeping Beauty'' transposon system is a synthetic DNA
 The ''Sleeping Beauty'' transposon system is composed of a ''Sleeping Beauty'' (SB)
The ''Sleeping Beauty'' transposon system is composed of a ''Sleeping Beauty'' (SB)
 This resurrected transposase gene was named "''Sleeping Beauty (SB)''", after the story of
This resurrected transposase gene was named "''Sleeping Beauty (SB)''", after the story of  SB10 transposase has been improved over the decade since its construction by increasing the consensus with a greater number of extinct Tc1 transposon sequences and testing various combinations of changes. Further work has shown that the DNA-binding domain consists of two paired sequences, which are homologous to sequence motifs found in certain transcription factors. The paired subdomains in SB transposase were designated PAI and RED. The PAI subdomain plays a dominant role in recognition of the DR sequences in the transposon. The RED subdomain overlaps with the nuclear localization signal, but its function remains unclear. The most recent version of SB transposase, SB100X, has about 100 times the activity of SB10 as determined by transposition assays of antibiotic-resistance genes conducted in tissue cultured human HeLa cells. The International Society for Molecular and Cell Biology and Biotechnology Protocols and Research (ISMCBBPR) named SB100X the molecule of the year for 2009 for recognition of the potential it has in future genome engineering.
The transposon recognized by SB transposase was named T because it was isolated from the genome of another salmond fish, ''Tanichthys albonubes''. The transposon consists of a genetic sequence of interest that is flanked by
SB10 transposase has been improved over the decade since its construction by increasing the consensus with a greater number of extinct Tc1 transposon sequences and testing various combinations of changes. Further work has shown that the DNA-binding domain consists of two paired sequences, which are homologous to sequence motifs found in certain transcription factors. The paired subdomains in SB transposase were designated PAI and RED. The PAI subdomain plays a dominant role in recognition of the DR sequences in the transposon. The RED subdomain overlaps with the nuclear localization signal, but its function remains unclear. The most recent version of SB transposase, SB100X, has about 100 times the activity of SB10 as determined by transposition assays of antibiotic-resistance genes conducted in tissue cultured human HeLa cells. The International Society for Molecular and Cell Biology and Biotechnology Protocols and Research (ISMCBBPR) named SB100X the molecule of the year for 2009 for recognition of the potential it has in future genome engineering.
The transposon recognized by SB transposase was named T because it was isolated from the genome of another salmond fish, ''Tanichthys albonubes''. The transposon consists of a genetic sequence of interest that is flanked by
 Over the past decade, SB transposons have been developed as non-viral vectors for introduction of genes into genomes of vertebrate animals and for
Over the past decade, SB transposons have been developed as non-viral vectors for introduction of genes into genomes of vertebrate animals and for
transposon
A transposable element (TE), also transposon, or jumping gene, is a type of mobile genetic element, a nucleic acid sequence in DNA that can change its position within a genome.
The discovery of mobile genetic elements earned Barbara McClinto ...
designed to introduce precisely defined DNA
Deoxyribonucleic acid (; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a double helix. The polymer carries genetic instructions for the development, functioning, growth and reproduction of al ...
sequences into the chromosomes
A chromosome is a package of DNA containing part or all of the genetic material of an organism. In most chromosomes, the very long thin DNA fibers are coated with nucleosome-forming packaging proteins; in eukaryotic cells, the most importa ...
of vertebrate
Vertebrates () are animals with a vertebral column (backbone or spine), and a cranium, or skull. The vertebral column surrounds and protects the spinal cord, while the cranium protects the brain.
The vertebrates make up the subphylum Vertebra ...
animals for the purposes of introducing new traits
Trait may refer to:
* Phenotypic trait in biology, which involve genes and characteristics of organisms
* Genotypic trait, sometimes but not always presenting as a phenotypic trait
* Personality, traits that predict an individual's behavior.
** ...
and to discover new genes
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protei ...
and their functions. It is a Tc1/mariner-type system, with the transposase resurrected from multiple inactive fish sequences.
Mechanism of action
 The ''Sleeping Beauty'' transposon system is composed of a ''Sleeping Beauty'' (SB)
The ''Sleeping Beauty'' transposon system is composed of a ''Sleeping Beauty'' (SB) transposase
A transposase is any of a class of enzymes capable of binding to the end of a transposon and catalysing its movement to another part of a genome, typically by a cut-and-paste mechanism or a replicative mechanism, in a process known as transpositio ...
and a transposon
A transposable element (TE), also transposon, or jumping gene, is a type of mobile genetic element, a nucleic acid sequence in DNA that can change its position within a genome.
The discovery of mobile genetic elements earned Barbara McClinto ...
that was designed in 1997 to insert specific sequences of DNA
Deoxyribonucleic acid (; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a double helix. The polymer carries genetic instructions for the development, functioning, growth and reproduction of al ...
into genomes of vertebrate animals. DNA transposons translocate from one DNA site to another in a simple, cut-and-paste manner (Fig. 1). Transposition is a precise process in which a defined DNA segment is excised from one DNA molecule and moved to another site in the same or different DNA molecule or genome
A genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as ...
.
As do all other Tc1/mariner-type transposases, SB transposase inserts a transposon into a TA dinucleotide base pair
A base pair (bp) is a fundamental unit of double-stranded nucleic acids consisting of two nucleobases bound to each other by hydrogen bonds. They form the building blocks of the DNA double helix and contribute to the folded structure of both DNA ...
in a recipient DNA sequence. The insertion site can be elsewhere in the same DNA molecule, or in another DNA molecule (or chromosome). In mammalian genomes, including humans, there are approximately 200 million TA sites. The TA insertion site is duplicated in the process of transposon integration. This duplication of the TA sequence is a hallmark of transposition and used to ascertain the mechanism in some experiments. However, a recent study indicated that SB also integrates into non-TA dinucleotides at a low frequency. The transposase can be encoded either within the transposon (e.g., the putative transposon shown in Fig. 2) or the transposase can be supplied by another source, in which case the transposon becomes a non-autonomous element. Non-autonomous transposons (e.g., Fig. 1) are most useful as genetic tools because after insertion they cannot independently continue to excise and re-insert. All of the DNA transposons identified in the human genome and other mammalian genomes are non-autonomous because even though they contain transposase genes, the genes are non-functional and unable to generate a transposase that can mobilize the transposon.
Construction
 This resurrected transposase gene was named "''Sleeping Beauty (SB)''", after the story of
This resurrected transposase gene was named "''Sleeping Beauty (SB)''", after the story of Sleeping Beauty
"Sleeping Beauty" (, or ''The Beauty Sleeping in the Wood''; , or ''Little Briar Rose''), also titled in English as ''The Sleeping Beauty in the Woods'', is a fairy tale about a princess curse, cursed by an evil fairy to suspended animation in fi ...
, because it was "awakened from a long evolutionary sleep". The SB transposon system is synthetic in that the SB transposase was re-constructed from extinct (fossil) transposase sequences belonging to the Tc1/mariner class of transposons found in the genomes of salmonid fish. As in humans, where about 20,000 inactivated Tc1/mariner-type transposons comprise almost 3% of the human genome
The human genome is a complete set of nucleic acid sequences for humans, encoded as the DNA within each of the 23 distinct chromosomes in the cell nucleus. A small DNA molecule is found within individual Mitochondrial DNA, mitochondria. These ar ...
, the transposase genes found in fish have been inactive for more than 10 million years due to accumulated mutations. The reconstruction of SB transposase was based on the concept that there was a primordial Tc1-like transposon that was the ancestor to the sequences found in fish genomes. Although there were many sequences that looked like Tc-1 transposons in all the fish genomes studied, the transposon sequences were all inactive due to mutations. By assuming that the variations in sequences were due to independent mutations that accumulated in the different transposons, a putative ancestral transposon (Fig. 2) was postulated.
The construction for the transposase began by fusing portions of two inactive transposon sequences from Atlantic salmon
The Atlantic salmon (''Salmo salar'') is a species of ray-finned fish in the family Salmonidae. It is the third largest of the Salmonidae, behind Hucho taimen, Siberian taimen and Pacific Chinook salmon, growing up to a meter in length. Atlan ...
(''Salmo salar'') and one inactive transposon sequence from rainbow trout
The rainbow trout (''Oncorhynchus mykiss'') is a species of trout native to cold-water tributary, tributaries of the Pacific Ocean in North America and Asia. The steelhead (sometimes called steelhead trout) is an Fish migration#Classification, ...
(''Oncorhynchus mykiss'') and then repairing small deficits in the functional domains of the transposase enzyme
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different mol ...
(Fig. 3). Each amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although over 500 amino acids exist in nature, by far the most important are the 22 α-amino acids incorporated into proteins. Only these 22 a ...
in the first completed transposase, called SB10, was determined by a “majority-rule consensus sequence” based on 12 partial genes found in eight fish species. The first steps (1->3 in Fig. 3) were to restore a complete protein by filling in gaps in the sequence and reversing termination codons that would keep the putative 360-amino acid polypeptide from being synthesized. The next step (4 in Fig. 3) was to reverse mutations in the nuclear localization signal
A nuclear localization signal ''or'' sequence (NLS) is an amino acid sequence that 'tags' a protein for import into the cell nucleus by nuclear transport. Typically, this signal consists of one or more short sequences of positively charged lysin ...
(NLS) that is required to import the transposase enzyme from the cytoplasm
The cytoplasm describes all the material within a eukaryotic or prokaryotic cell, enclosed by the cell membrane, including the organelles and excluding the nucleus in eukaryotic cells. The material inside the nucleus of a eukaryotic cell a ...
where it is made to the nucleus
Nucleus (: nuclei) is a Latin word for the seed inside a fruit. It most often refers to:
*Atomic nucleus, the very dense central region of an atom
*Cell nucleus, a central organelle of a eukaryotic cell, containing most of the cell's DNA
Nucleu ...
where it acts. The amino-terminus of the transposase, which contains the DNA-binding motifs for recognition of the direct repeats (DRs), was restored in steps 5->8. The last two steps restored the catalytic domain, which features conserved aspartic acid
Aspartic acid (symbol Asp or D; the ionic form is known as aspartate), is an α-amino acid that is used in the biosynthesis of proteins. The L-isomer of aspartic acid is one of the 22 proteinogenic amino acids, i.e., the building blocks of protei ...
(D) and glutamic acid
Glutamic acid (symbol Glu or E; known as glutamate in its anionic form) is an α- amino acid that is used by almost all living beings in the biosynthesis of proteins. It is a non-essential nutrient for humans, meaning that the human body can ...
(E) amino acids with specific spacing that are found in integrases
Retroviral integrase (IN) is an enzyme produced by a retrovirus (such as HIV) that Retroviral integration, integrates (forms Covalent bond, covalent links between) its genetic information into that of the host Cell (biology), cell it infects. Re ...
and recombinases
Recombinases are genetic recombination enzymes.
Site specific recombinases
DNA recombinases are widely used in multicellular organisms to manipulate the structure of genomes, and to control gene expression. These enzymes, derived from bacteria ( ...
. The result was SB10, which contains all of the motifs required for function.
 SB10 transposase has been improved over the decade since its construction by increasing the consensus with a greater number of extinct Tc1 transposon sequences and testing various combinations of changes. Further work has shown that the DNA-binding domain consists of two paired sequences, which are homologous to sequence motifs found in certain transcription factors. The paired subdomains in SB transposase were designated PAI and RED. The PAI subdomain plays a dominant role in recognition of the DR sequences in the transposon. The RED subdomain overlaps with the nuclear localization signal, but its function remains unclear. The most recent version of SB transposase, SB100X, has about 100 times the activity of SB10 as determined by transposition assays of antibiotic-resistance genes conducted in tissue cultured human HeLa cells. The International Society for Molecular and Cell Biology and Biotechnology Protocols and Research (ISMCBBPR) named SB100X the molecule of the year for 2009 for recognition of the potential it has in future genome engineering.
The transposon recognized by SB transposase was named T because it was isolated from the genome of another salmond fish, ''Tanichthys albonubes''. The transposon consists of a genetic sequence of interest that is flanked by
SB10 transposase has been improved over the decade since its construction by increasing the consensus with a greater number of extinct Tc1 transposon sequences and testing various combinations of changes. Further work has shown that the DNA-binding domain consists of two paired sequences, which are homologous to sequence motifs found in certain transcription factors. The paired subdomains in SB transposase were designated PAI and RED. The PAI subdomain plays a dominant role in recognition of the DR sequences in the transposon. The RED subdomain overlaps with the nuclear localization signal, but its function remains unclear. The most recent version of SB transposase, SB100X, has about 100 times the activity of SB10 as determined by transposition assays of antibiotic-resistance genes conducted in tissue cultured human HeLa cells. The International Society for Molecular and Cell Biology and Biotechnology Protocols and Research (ISMCBBPR) named SB100X the molecule of the year for 2009 for recognition of the potential it has in future genome engineering.
The transposon recognized by SB transposase was named T because it was isolated from the genome of another salmond fish, ''Tanichthys albonubes''. The transposon consists of a genetic sequence of interest that is flanked by inverted repeats
An inverted repeat (or IR) is a single stranded sequence of nucleotides followed downstream by its reverse complement. The intervening sequence of nucleotides between the initial sequence and the reverse complement can be any length including zero ...
(IRs) that themselves contain short direct repeats (DR) (tandem arrowheads IR-DR in Figs. 1 and 2). T had the closest IR/DR sequence to the consensus sequence for the extinct Tc-1 like transposons in fish. The consensus transposon has IRs of 231 base pairs. The innermost DRs are 29 base pairs long whereas the outermost DRs are 31 base pairs long. The difference in length is critical for maximal transposition rates. The original T transposon component of the SB transposon system has been improved with minor changes to conform to the consensus of many related extinct and active transposons.
Applications
 Over the past decade, SB transposons have been developed as non-viral vectors for introduction of genes into genomes of vertebrate animals and for
Over the past decade, SB transposons have been developed as non-viral vectors for introduction of genes into genomes of vertebrate animals and for gene therapy
Gene therapy is Health technology, medical technology that aims to produce a therapeutic effect through the manipulation of gene expression or through altering the biological properties of living cells.
The first attempt at modifying human DNA ...
. The genetic cargo can be an expression cassette—a transgene
A transgene is a gene that has been transferred naturally, or by any of a number of genetic engineering techniques, from one organism to another. The introduction of a transgene, in a process known as transgenesis, has the potential to change the ...
and associated elements that confer transcriptional regulation
In molecular biology and genetics, transcriptional regulation is the means by which a cell regulates the conversion of DNA to RNA ( transcription), thereby orchestrating gene activity. A single gene can be regulated in a range of ways, from al ...
for expression at a desired level in specific tissue(s). An alternative use of SB transposons is to discover functions of genes, especially those that cause cancer
Cancer is a group of diseases involving Cell growth#Disorders, abnormal cell growth with the potential to Invasion (cancer), invade or Metastasis, spread to other parts of the body. These contrast with benign tumors, which do not spread. Po ...
, by delivering DNA sequences that maximally disrupt expression of genes close to the insertion site. This process is referred to as insertional mutagenesis In molecular biology, insertional mutagenesis is the creation of mutations in DNA by the addition of one or more base pairs. Such insertional mutations can occur naturally, mediated by viruses or transposons, or can be artificially created for resea ...
or transposon mutagenesis Transposon mutagenesis, or transposition mutagenesis, is a biological process that allows genes to be transferred to a host organism's chromosome, interrupting or modifying the function of an extant gene on the chromosome and causing mutation. Trans ...
. When a gene is inactivated by insertion of a transposon (or other mechanism), that gene is “knocked out”. Knockout mice
A knockout mouse, or knock-out mouse, is a genetically modified mouse (''Mus musculus'') in which researchers have inactivated, or " knocked out", an existing gene by replacing it or disrupting it with an artificial piece of DNA. They are importan ...
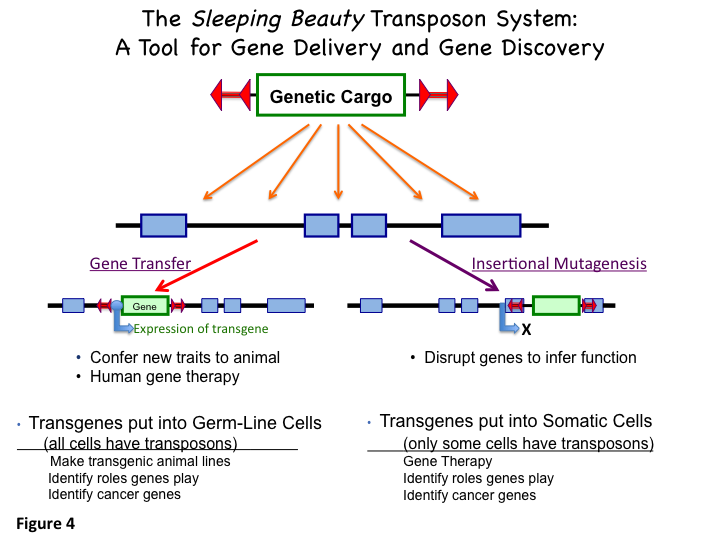
and knockout rats have been made with the SB system. Figure 4 illustrates these two uses of SB transposons.
For either gene delivery or gene disruption, SB transposons combine the advantages of viruses and naked DNA. Viruses have been evolutionarily selected based on their abilities to infect and replicate in new host cells. Simultaneously, cells have evolved major molecular defense mechanisms to protect themselves against viral infections. For some applications of genome engineering such as some forms of gene therapy, avoiding the use of viruses is also important for social and regulatory reasons. The use of non-viral vectors avoids many, but not all, of the defenses that cells employ against vectors.
Plasmids
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria and ...
, the circular DNAs shown in Fig. 1, are generally used for non-viral gene delivery. However, there are two major problems with most methods for delivering DNA to cellular chromosomes using plasmids, the most common form of non-viral gene delivery. First, expression of transgenes from plasmids is brief due to lack of integration and due to cellular responses that turn off expression. Second, uptake of plasmid molecules into cells is difficult and inefficient. The Sleeping Beauty Transposon System was engineered to overcome the first problem. DNA transposons precisely insert defined DNA sequences (Fig. 1) almost randomly into host genomes thereby increasing the longevity of gene expression (even through multiple generations). Moreover, transposition avoids the formation of multiple, tandem integrations, which often results in switching off expression of the transgene. Currently, insertion of transgenes into chromosomes using plasmids is much less efficient than using viruses. However, by using powerful promoters to regulate expression of a transgene, delivery of transposons to a few cells can provide useful levels of secreted gene products for an entire animal.
Arguably the most exciting potential application of ''Sleeping Beauty'' transposons will be for human gene therapy. The widespread human application of gene therapy in first-world nations as well as countries with developing economies can be envisioned if the costs of the vector system are affordable. Because the SB system is composed solely of DNA, the costs of production and delivery are considerably reduced compared to viral vectors. The first clinical trials using SB transposons in genetically modified T cells will test the efficacy of this form of gene therapy in patients at risk of death from advanced malignancies.
See also
* PiggyBac transposon systemReferences
{{Reflist, 2 Mobile genetic elements