Phage-assisted Continuous Evolution on:
[Wikipedia]
[Google]
[Amazon]
Phage-assisted continuous evolution (PACE) is a



 PACE was also used to create a more catalytically active deoxyadenosine deaminase. Deoxyadenosine deaminase is used in base editors to perform the single nucleotide edit A-->T. This was done by placing
PACE was also used to create a more catalytically active deoxyadenosine deaminase. Deoxyadenosine deaminase is used in base editors to perform the single nucleotide edit A-->T. This was done by placing 
phage
A bacteriophage (), also known informally as a phage (), is a virus that infects and replicates within bacteria. The term is derived . Bacteriophages are composed of proteins that encapsulate a DNA or RNA genome, and may have structures tha ...
-based technique for the automated directed evolution
Directed evolution (DE) is a method used in protein engineering that mimics the process of natural selection to steer proteins or nucleic acids toward a user-defined goal. It consists of subjecting a gene to iterative rounds of mutagenesis (cre ...
of proteins. It relies on relating the desired activity of a target protein with the fitness of an infectious bacteriophage which carries the protein's corresponding gene. Proteins with greater desired activity hence confer greater infectivity to their carrier phage. More infectious phage propagate more effectively, selecting for advantageous mutations. Genetic variation
Genetic variation is the difference in DNA among individuals or the differences between populations among the same species. The multiple sources of genetic variation include mutation and genetic recombination. Mutations are the ultimate sources ...
is generated using error-prone polymerases
In biochemistry, a polymerase is an enzyme ( EC 2.7.7.6/7/19/48/49) that synthesizes long chains of polymers or nucleic acids. DNA polymerase and RNA polymerase are used to assemble DNA and RNA molecules, respectively, by copying a DNA template s ...
on the phage vectors, and over time the protein accumulates beneficial mutations. This technique is notable for performing hundreds of rounds of selection with minimal human intervention.
Principle
The central component of PACE is a fixed-volume vessel known as the “lagoon”. The lagoon containsM13 bacteriophage
M13 is one of the Ff phages (fd and f1 are others), a member of the family filamentous bacteriophage ( inovirus). Ff phages are composed of circular single-stranded DNA (ssDNA), which in the case of the m13 phage is 6407 nucleotides long and i ...
vectors carrying the gene of interest (known as the selection plasmid, or SP), as well as host ''E. coli
''Escherichia coli'' ( )Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. is a gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus ''Escherichia'' that is commonly foun ...
'' cells that allow the phage to replicate. The lagoon is constantly diluted via the addition and draining of liquid media containing ''E. coli'' cells. The liquid flow rate is set such that the dilution rate is faster than the rate of ''E. coli'' reproduction but slower than the rate of phage reproduction. Hence, a fresh supply of ''E. coli'' cells is constantly present in the lagoon, but phage can only be retained via sufficiently fast replication.
Phage replication requires ''E. coli'' infection, which, for M13 phage, relies on protein III (pIII). When using PACE, the phage vectors lack the gene to produce pIII. Instead, the production of pIII is tied with the activity of the protein of interest via a mechanism that varies per use case, oftentimes involves an extra plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria and ...
containing the pIII-expressing gene III (gIII) known as the accessory plasmid, or AP. Notably, production of infectious phage scales with the production of pIII. Hence, the better the activity of the protein, the higher the rate of pIII production, and the more infectious phage are generated for that particular gene.
Using error-prone polymerases (encoded on the mutagenesis plasmid, or MP), genetic variation is introduced into the protein gene portion of the phage vectors. Due to the selective pressures applied by the constant draining of the lagoon, only phages that can replicate fast enough can be retained in the lagoon, so over time beneficial mutations accumulate in phage replicating in the lagoon. In this manner, rounds of evolution are continuously performed, allowing hundreds of rounds to elapse with little human intervention.

Applications
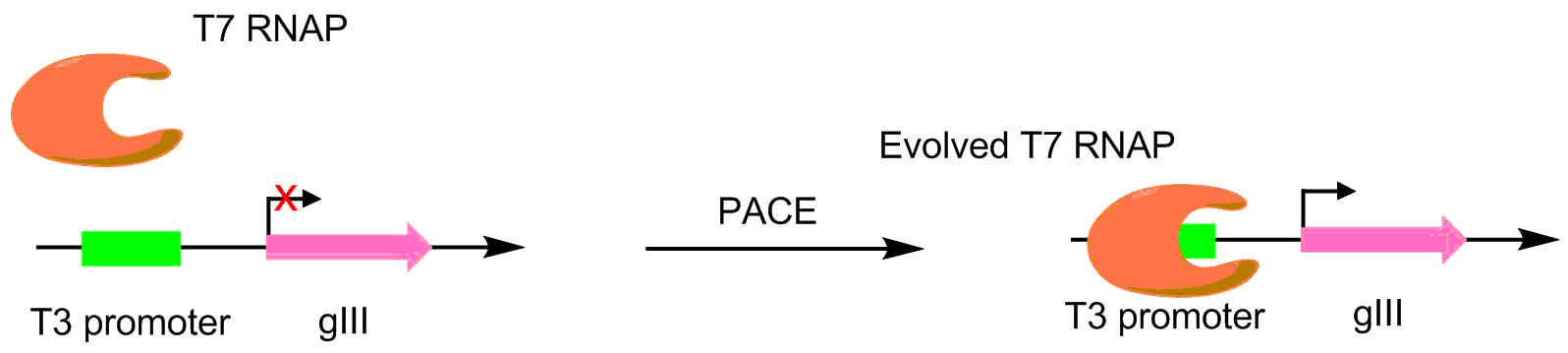
Polymerase promoter specificity
In the initial paper pioneering this technique,T7 RNA polymerase
T7 RNA Polymerase is an RNA polymerase from the T7 bacteriophage that catalyzes the formation of RNA from DNA in the 5'→ 3' direction.
Activity
T7 polymerase is extremely promoter-specific and transcribes only DNA downstream of a T7 promo ...
s were evolved to recognize different promoters, such as the T3 or SP6 promoters. This was done by making the target promoter the sole promoter for gIII. Hence, mutant polymerases with greater specificity for the desired promoter caused greater pIII production. This resulted in polymerases with ~3-4 orders of magnitude greater activity for the target promoter than the original T3 promoter.
While this original PACE system only performed positive selection, a variant was developed that allowed for negative selection as well. This is done by linking undesired activity to the production of non-functional pIII, which decreases the amount of infectious phage made.

Protease substrate specificity
Proteases
A protease (also called a peptidase, proteinase, or proteolytic enzyme) is an enzyme that catalyzes proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do ...
have been evolved to cut different peptides using PACE. In these systems, the desired protease cut site is used to link a T7 RNA polymerase and a T7 lysozyme
Lysozyme (, muramidase, ''N''-acetylmuramide glycanhydrolase; systematic name peptidoglycan ''N''-acetylmuramoylhydrolase) is an antimicrobial enzyme produced by animals that forms part of the innate immune system. It is a glycoside hydrolase ...
. The T7 lysozyme prevents the T7 polymerase from transcribing gIII. When the peptide linker is cleaved, the T7 polymerase is activated, allowing for the transcription of the pIII gene. This method was used to create a TEV protease
TEV protease (, ''Tobacco Etch Virus nuclear-inclusion-a endopeptidase'') is a highly sequence-specific cysteine protease from Tobacco Etch Virus (TEV). It is a member of the PA clan of chymotrypsin-like proteases. Due to its high sequence sp ...
with a significantly different peptide substrate.

Orthogonal Aminoacyl-tRNA Synthetases
Using PACE,aminoacyl-tRNA synthetases
An aminoacyl-tRNA synthetase (aaRS or ARS), also called tRNA-ligase, is an enzyme that attaches the appropriate amino acid onto its corresponding tRNA. It does so by catalyzing the transesterification of a specific cognate amino acid or its pre ...
(aaRSs) were evolved for noncanonical amino acids as well. Activity of an aaRS is linked to pIII production by the addition of a TAG stop codon in the middle of gIII. Synthetases that aminoacylate the TAG codon's suppressor tRNA prevents stop codon
In molecular biology, a stop codon (or termination codon) is a codon (nucleotide triplet within messenger RNA) that signals the termination of the translation process of the current protein. Most codons in messenger RNA correspond to the additio ...
activity, allowing for production of functional pIII. Using this system, aaRSs were evolved that utilize non-canonical amino acids ''p''-nitro-phenyalanine, iodophenylalanine, and Boc-lysine.

Protein-Protein Interactions
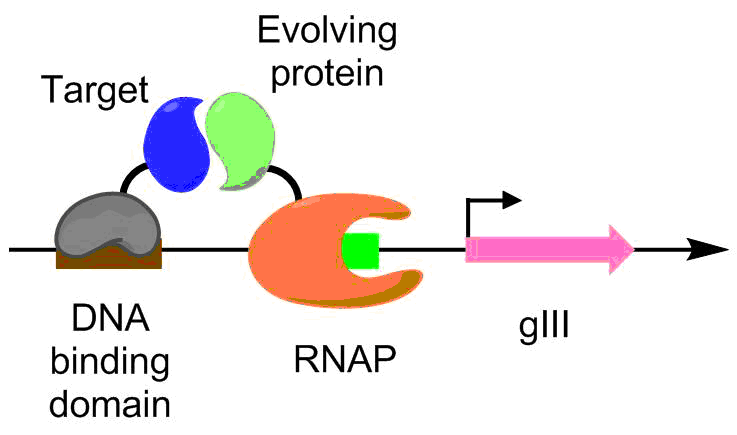
Protein-protein interactions have been evolved using PACE as well. Under this scheme, the target protein is fused with a DNA binding protein, which binds to a target sequence placed upstream of the gIII promoter. The protein undergoing evolution is fused with an RNA polymerase. The better the protein-protein interaction, the more transcription of pIII occurs, allowing the evolution of the protein-protein interaction under PACE conditions. This method was used to evolve ''Bacillus thuringiensis
''Bacillus thuringiensis'' (or Bt) is a gram-positive bacteria, gram-positive, soil-dwelling bacterium, the most commonly used biological pesticide worldwide. ''B. thuringiensis'' also occurs naturally in the gut of caterpillars of various types ...
'' endotoxin
Lipopolysaccharide (LPS), now more commonly known as endotoxin, is a collective term for components of the outermost membrane of the cell envelope of gram-negative bacteria, such as '' E. coli'' and ''Salmonella'' with a common structural archit ...
variants that can overcome insect toxin resistance.

Base Editors
PACE was used to evolveAPOBEC1
Apolipoprotein B mRNA editing enzyme, catalytic polypeptide 1 also known as C->U-editing enzyme APOBEC-1 is a protein that in humans is encoded by the ''APOBEC1'' gene.
This gene encodes a member of the APOBEC, APOBEC protein family and the cyti ...
for greater soluble expression. APOBEC1 is a cytidine deaminase
Cytidine deaminase is an enzyme that in humans is encoded by the ''CDA'' gene.
This gene encodes an enzyme involved in pyrimidine salvaging. The encoded protein forms a homotetramer that catalyzes the irreversible hydrolytic deamination of cytidi ...
that has found use in base editors to catalyze the single nucleotide edit C-->T. In ''E. coli'', APOBEC1 usually falls out of solution into the insoluble fraction. To evolve APOBEC1 for better soluble expression, the N-terminus of a T7 polymerase was fused to APOBEC1, with the remaining portion of the polymerase separately expressed. The T7 polymerase can only function when the N-terminus portion can bind to the rest of the polymerase. Since APOBEC1 must be properly folded for the N-terminus portion to be exposed properly, T7 polymerase activity is correlated to APOBEC1 folding. As follows, pIII transcription and production is linked with APOBEC1 soluble expression via the T7 polymerase. Using this approach, the soluble expression of APOBEC1 was increased by 4 fold with no change in function.
 PACE was also used to create a more catalytically active deoxyadenosine deaminase. Deoxyadenosine deaminase is used in base editors to perform the single nucleotide edit A-->T. This was done by placing
PACE was also used to create a more catalytically active deoxyadenosine deaminase. Deoxyadenosine deaminase is used in base editors to perform the single nucleotide edit A-->T. This was done by placing adenosine
Adenosine (symbol A) is an organic compound that occurs widely in nature in the form of diverse derivatives. The molecule consists of an adenine attached to a ribose via a β-N9- glycosidic bond. Adenosine is one of the four nucleoside build ...
-containing stop codons in the gene for T7 polymerase. If the base editor is able to correct the error, functional T7 polymerase is produced, allowing production of pIII. Using this system, they evolved a deoxyadenosine deaminase with 590 fold activity compared to wild type.

References
{{reflist Evolutionary biology