Histone H2A on:
[Wikipedia]
[Google]
[Amazon]
Histone H2A is one of the five main 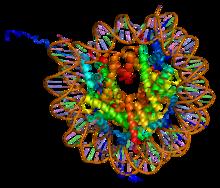
"HistoneDB with Variants" database
Changes in variant composition occur in differentiating cells. This was observed in differentiating neurons during synthesis and turnover; changes in variant composition were seen among the H2A.1 histone. The only variant that remained constant in the neural differentiation was variant H2A.Z. H2A.Z is a variant that exchanges with conventional H2A core protein; this variant is important for gene silencing. Physically, there are small changes on the surface area of the nucleosome that make the histone differ from H2A. Recent research suggests that H2AZ is incorporated into the nucleosome using a Swr1, a Swi2/Snf2- related adenosine triphosphatase. Another H2A variant that has been identified is H2AX. This variant has a
 H2A consists of a main globular domain, an
H2A consists of a main globular domain, an
 DNA Folding:
H2A is important for packaging DNA into chromatin. Since H2A packages DNA molecules into chromatin, the packaging process will affect gene expression. H2A has been correlated with DNA modification and
DNA Folding:
H2A is important for packaging DNA into chromatin. Since H2A packages DNA molecules into chromatin, the packaging process will affect gene expression. H2A has been correlated with DNA modification and
Nextbio
{{Chromo Proteins
histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei and in most Archaeal phyla. They act as spools around which DNA winds to create structural units called nucleosomes ...
proteins involved in the structure of chromatin
Chromatin is a complex of DNA and protein found in eukaryote, eukaryotic cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important r ...
in eukaryotic cells.
The other histone
In biology, histones are highly basic proteins abundant in lysine and arginine residues that are found in eukaryotic cell nuclei and in most Archaeal phyla. They act as spools around which DNA winds to create structural units called nucleosomes ...
proteins
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residues. Proteins perform a vast array of functions within organisms, including catalysing metabolic reactions, DNA replication, re ...
are: H1, H2B, H3 and H4.

Background
Histones are proteins that packageDNA
Deoxyribonucleic acid (; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a double helix. The polymer carries genetic instructions for the development, functioning, growth and reproduction of al ...
into nucleosome
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone, histone proteins and resembles thread wrapped around a bobbin, spool. The nucleosome ...
s. Histones are responsible for maintaining the shape and structure of a nucleosome. One chromatin molecule is composed of at least one of each core histones per 100 base pairs of DNA.
There are five families of histones known to date; these histones are termed H1/H5, H2A, H2B, H3, and H4. H2A is considered a core histone, along with H2B, H3 and H4. Core formation first occurs through the interaction of two H2A molecules. Then, H2A forms a dimer with H2B; the core molecule is complete when H3-H4 also attaches to form a tetramer.
Sequence variants
Histone H2A is composed of non-allelic variants. The term "Histone H2A" is intentionally non-specific and refers to a variety of closely related proteins that vary often by only a few amino acids. Apart from the canonical form, notable variants include H2A.1, H2A.2, H2A.X, and H2A.Z. H2A variants can be explored usin"HistoneDB with Variants" database
Changes in variant composition occur in differentiating cells. This was observed in differentiating neurons during synthesis and turnover; changes in variant composition were seen among the H2A.1 histone. The only variant that remained constant in the neural differentiation was variant H2A.Z. H2A.Z is a variant that exchanges with conventional H2A core protein; this variant is important for gene silencing. Physically, there are small changes on the surface area of the nucleosome that make the histone differ from H2A. Recent research suggests that H2AZ is incorporated into the nucleosome using a Swr1, a Swi2/Snf2- related adenosine triphosphatase. Another H2A variant that has been identified is H2AX. This variant has a
C-terminal
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, carboxy tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When t ...
extension that is utilized for DNA repair. The method of repair this variant employs is non-homologous end joining
Non-homologous end joining (NHEJ) is a pathway that repairs double-strand breaks in DNA. It is called "non-homologous" because the break ends are directly ligated without the need for a homologous template, in contrast to homology directed repair ...
. Direct DNA damage
Direct may refer to:
Mathematics
* Directed set, in order theory
* Direct limit of (pre), sheaves
* Direct sum of modules, a construction in abstract algebra which combines several vector spaces
Computing
* Direct access (disambiguation), a ...
can induce changes to the sequence variants. Experiments performed with ionizing radiation linked γ- phosphorylation of H2AX to double-strand breaks
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. A weakened capacity for DNA repair is a risk factor for the development of cancer. DNA is constantly modified ...
. A large amount of chromatin is involved with each DNA double-strand break; a response to DNA damage is the formation of γ- H2AX.
Lastly, MacroH2A variant is a variant that is similar to H2A; it is encoded by the H2AFY gene. This variant differs from H2A because of the addition of a fold domain in its C-terminal tail. MacroH2A is expressed in the inactive X chromosome in females.
Structure
N-terminal
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amin ...
tail and a C-terminal tail. Both tails are the location of post-translational modification
In molecular biology, post-translational modification (PTM) is the covalent process of changing proteins following protein biosynthesis. PTMs may involve enzymes or occur spontaneously. Proteins are created by ribosomes, which translation (biolog ...
. Thus far, researchers have not identified any secondary structures that arise in the tails. H2A utilizes a protein fold known as the ‘ histone fold’. The histone fold is a three-helix core domain that is connected by two loops. This connection forms a ‘handshake arrangement.’ Most notably, this is termed the helix-turn-helix
Helix-turn-helix is a DNA-binding domain (DBD). The helix-turn-helix (HTH) is a major structural motif capable of binding DNA. Each monomer incorporates two alpha helix, α helices, joined by a short strand of amino acids, that bind to the majo ...
motif, which allows for dimerization with H2B. The ‘histone fold’ is conserved among H2A at the structural level; however the genetic sequence that encodes for this structure differs between variants.
The structure of macroH2A variant was exposed through X-ray crystallography
X-ray crystallography is the experimental science of determining the atomic and molecular structure of a crystal, in which the crystalline structure causes a beam of incident X-rays to Diffraction, diffract in specific directions. By measuring th ...
. The conserved domain contains a DNA binding structure and a peptidase fold. The function of this conserved domain remains unknown. Research suggests that this conserved domain may function as an anchor site for Xist DNA or it may also function as a modifying enzyme.
Function
epigenetics
In biology, epigenetics is the study of changes in gene expression that happen without changes to the DNA sequence. The Greek prefix ''epi-'' (ἐπι- "over, outside of, around") in ''epigenetics'' implies features that are "on top of" or "in ...
. H2A plays a major role in determining the overall structure of chromatin. Inadvertently, H2A has been found to regulate gene expression.
DNA modification by H2A occurs in the cell nucleus
The cell nucleus (; : nuclei) is a membrane-bound organelle found in eukaryote, eukaryotic cell (biology), cells. Eukaryotic cells usually have a single nucleus, but a few cell types, such as mammalian red blood cells, have #Anucleated_cells, ...
. Proteins responsible for nuclear import of H2A protein are karyopherin
Karyopherins are protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme c ...
and importin
Importin is a type of karyopherin that transports protein molecules from the Eukaryotic Cell, cell's cytoplasm to the cell nucleus, nucleus. It does so by binding to specific recognition sequences, called nuclear localization sequences (NLS).
I ...
. Recent studies also show that nucleosome assembly protein 1 is also used to transport of H2A into the nucleus so it can wrap DNA.
Other functions of H2A have been seen in the histone variant H2A.Z. This variant is associated with gene activation, silencing and suppression of antisense RNA
Antisense RNA (asRNA), also referred to as antisense transcript, natural antisense transcript (NAT) or antisense oligonucleotide, is a single stranded RNA that is complementary to a protein coding messenger RNA (mRNA) with which it hybridizes, and ...
. In addition, when H2A.Z was studied in human and yeast cells, it was used to promote RNA polymerase II recruitment.
Antimicrobial peptide:
Histones are conserved eukaryotic cationic proteins present in the cells and are involved in the antimicrobial
activities. In vertebrates and invertebrates, Histone H2A variant is reported to be involved in host immune response by acting as antimicrobial peptides (AMPs). H2A are α-helical molecule, amphipathic protein with hydrophobic and hydrophilic residues on opposing sides that enhances the antimicrobial activity of H2A.
DNA damage response
Site specificubiquitin
Ubiquitin is a small (8.6 kDa) regulatory protein found in most tissues of eukaryotic organisms, i.e., it is found ''ubiquitously''. It was discovered in 1975 by Gideon Goldstein and further characterized throughout the late 1970s and 19 ...
ation of histone H2A has a role in the recruitment of DNA repair
DNA repair is a collection of processes by which a cell (biology), cell identifies and corrects damage to the DNA molecules that encode its genome. A weakened capacity for DNA repair is a risk factor for the development of cancer. DNA is cons ...
proteins to DNA double strand breaks which then may be repaired by either homologous recombination
Homologous recombination is a type of genetic recombination in which genetic information is exchanged between two similar or identical molecules of double-stranded or single-stranded nucleic acids (usually DNA as in Cell (biology), cellular organi ...
or non-homologous end joining
Non-homologous end joining (NHEJ) is a pathway that repairs double-strand breaks in DNA. It is called "non-homologous" because the break ends are directly ligated without the need for a homologous template, in contrast to homology directed repair ...
.Uckelmann M, Sixma TK. Histone ubiquitination in the DNA damage response. DNA Repair (Amst). 2017 Aug;56:92-101. doi: 10.1016/j.dnarep.2017.06.011. Epub 2017 Jun 9. PMID 28624371 In the DNA damage response, it is thought that ubiquitination of H2A by the BRCA1
Breast cancer type 1 susceptibility protein is a protein that in humans is encoded by the ''BRCA1'' () gene. Orthologs are common in other vertebrate species, whereas invertebrate genomes may encode a more distantly related gene. ''BRCA1'' is a ...
/ BARD1 heterodimer promotes homologous recombination, and that ubiquitination of H2A by RNF168 protein promotes non-homologous end joining.
Genetics
H2A is coded by many genes in the human genome, including: H2AFB1, H2AFB2, H2AFB3, H2AFJ, H2AFV,H2AFX
H2A histone family member X (usually abbreviated as H2AX) is a type of histone protein from the histone H2A, H2A family encoded by the ''H2AFX'' gene. An important phosphorylated form is γH2AX (S139), which forms when double-strand breaks appear. ...
, H2AFY, H2AFY2, and H2AFZ
Histone H2A.Z is a protein encoded by the ''H2AZ1'' gene in humans.
Function
Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Nucleosomes consist of approximately 146 ...
. Genetic patterns among the different H2A molecules are mostly conserved among variants. The variability in gene expression
Gene expression is the process (including its Regulation of gene expression, regulation) by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, proteins or non-coding RNA, ...
exists among the regulatory machinery that manages H2A expression. Researchers studied eukaryotic evolutionary lineages of histone proteins and found diversification among the regulatory genes. The greatest differences were observed in core histone gene cis-regulatory sequence motifs and associated protein factors. Variability in gene sequence was seen in bacterial, fungi, plant, and mammalian genes.
One variant of H2A protein is H2ABbd (Barr body
A Barr body (named after discoverer Murray Barr) or X-chromatin is an inactive X chromosome. In species with XY sex-determination (including humans), females typically have two X chromosomes, and one is rendered inactive in a process calle ...
deficient) variant. This variant is composed of a different genetic sequence compared to H2A. The variant functions with transcriptionally active domains. Other variations associated with H2ABbd are located within its C-terminus
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, carboxy tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein
Proteins are large biomolecules and macromolecules that comp ...
. H2ABbd has a shorter C-terminal domain compared to the large C-terminal found on H2A. The two C terminals are about 48% identical. H2ABbd functions with active chromosomes. Thus far, it is missing from Xi chromosomes in fibroblast
A fibroblast is a type of cell (biology), biological cell typically with a spindle shape that synthesizes the extracellular matrix and collagen, produces the structural framework (Stroma (tissue), stroma) for animal Tissue (biology), tissues, and ...
cells. Lastly, it found to be associated with acetylated H4.
Different functions of H2A.Z compared to H2A are correlated with genetic differences between H2A and the variant. Resistance to nucleosomes occurs in H2A.Z by binding to H1 factor. H2A.Z gene is an essential gene in yeast and it is denoted as Htz1. Comparatively, vertebrates have two H2A.Z genes. These genes, H2A.Z1 and H2A.Z2 encode for proteins that differ from H2A.Z by three residues. At first researchers figured that these genes were redundant; however, when a mutant H2A.Z1 was created, it resulted in lethality during mammalian tests. Therefore, H2A.Z1 is an essential gene. On the other hand, researchers have not identified the function of H2A.Z2 variant. It is known that it is transcribed in mammals and this gene expression is conserved among mammalian species. This conservation suggests that the gene is functional.
When studying H2A.Z in plants species, the protein different among residues from species to species. These differences contribute to differences in cell-cycle regulation. This phenomenon was only observed in plants.
Phylogenetic trees were created to show the divergence of variants from their ancestors. The divergence of variant, H2A.X, from H2A occurred at multiple origins in a phylogenetic tree. Acquisition of the phosphorylation motif was consistent with the many origins of H2A that arose from an ancestral H2A.X. Finally, the presence of H2A.X and absence of H2A in fungi leads researchers to believe that H2A.X was the original ancestor of the histone protein H2A
Modification of H2A
H2A modification is under current research. However, modification of H2A does occur. Serinephosphorylation
In biochemistry, phosphorylation is described as the "transfer of a phosphate group" from a donor to an acceptor. A common phosphorylating agent (phosphate donor) is ATP and a common family of acceptor are alcohols:
:
This equation can be writ ...
sites have been identified on H2A. Threonine ''O''-GlcNAc has also been identified on H2A. Large differences exist between the modified residues of H2A variants. For example, H2ABbd lacks modified residues that exist in H2A. The differences in modification change the function of H2ABbd compared to H2A. As previously mentioned, variant H2AX was found to function in DNA repair
DNA repair is a collection of processes by which a cell (biology), cell identifies and corrects damage to the DNA molecules that encode its genome. A weakened capacity for DNA repair is a risk factor for the development of cancer. DNA is cons ...
. This function is dependent upon the phosphorylation of H2AX C-terminal. Once H2AX becomes phosphorylated, it can function in DNA repair. The H2A.X variant differs from H2A through modification. The C-terminal of H2A.X contains an additional motif compared to H2A. The motif that is added is Ser-Gln-(Glu/Asp)- (hydrophobic residue). The motif becomes heavily phosphorylated at the serine residue; if this phosphorylation occurs the variant becomes γH2A.X. Phosphorylation occurs due to dsDNA breaks. Modification on histone proteins can sometimes result in a change in function. Different H2A variants were exploited to have different functions, genetic sequences, and modifications.
See also
*Histone code
The histone code is a hypothesis that the transcription of genetic information encoded in DNA is in part regulated by chemical modifications (known as ''histone marks'') to histone proteins, primarily on their unstructured ends. Together with sim ...
*Chromatin
Chromatin is a complex of DNA and protein found in eukaryote, eukaryotic cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important r ...
*Nucleosome
A nucleosome is the basic structural unit of DNA packaging in eukaryotes. The structure of a nucleosome consists of a segment of DNA wound around eight histone, histone proteins and resembles thread wrapped around a bobbin, spool. The nucleosome ...
References
External links
Nextbio
{{Chromo Proteins