Functional genomics is a field of
molecular biology
Molecular biology is a branch of biology that seeks to understand the molecule, molecular basis of biological activity in and between Cell (biology), cells, including biomolecule, biomolecular synthesis, modification, mechanisms, and interactio ...
that attempts to describe
gene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protei ...
(and
protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme catalysis, catalysing metab ...
) functions and interactions. Functional genomics make use of the vast data generated by
genomic and
transcriptomic projects (such as
genome sequencing projects and
RNA sequencing
RNA-Seq (named as an abbreviation of RNA sequencing) is a technique that uses next-generation sequencing to reveal the presence and quantity of RNA molecules in a biological sample, providing a snapshot of gene expression in the sample, also kn ...
). Functional genomics focuses on the dynamic aspects such as gene
transcription,
translation
Translation is the communication of the semantics, meaning of a #Source and target languages, source-language text by means of an Dynamic and formal equivalence, equivalent #Source and target languages, target-language text. The English la ...
,
regulation of gene expression
Regulation of gene expression, or gene regulation, includes a wide range of mechanisms that are used by cells to increase or decrease the production of specific gene products (protein or RNA). Sophisticated programs of gene expression are wide ...
and
protein–protein interaction
Protein–protein interactions (PPIs) are physical contacts of high specificity established between two or more protein molecules as a result of biochemical events steered by interactions that include electrostatic forces, hydrogen bonding and t ...
s, as opposed to the static aspects of the genomic information such as
DNA sequence
A nucleic acid sequence is a succession of bases within the nucleotides forming alleles within a DNA (using GACT) or RNA (GACU) molecule. This succession is denoted by a series of a set of five different letters that indicate the order of the nu ...
or structures. A key characteristic of functional genomics studies is their genome-wide approach to these questions, generally involving high-throughput methods rather than a more traditional "candidate-gene" approach.
Definition and goals
In order to understand functional genomics it is important to first define function. In their paper Graur et al. define function in two possible ways. These are "selected effect" and "causal role". The "selected effect" function refers to the function for which a trait (DNA, RNA, protein etc.) is selected for. The "causal role" function refers to the function that a trait is sufficient and necessary for. Functional genomics usually tests the "causal role" definition of function.
The goal of functional genomics is to understand the function of genes or proteins, eventually all components of a genome. The term functional genomics is often used to refer to the many technical approaches to study an organism's genes and proteins, including the "biochemical, cellular, and/or physiological properties of each and every gene product"
while some authors include the study of nongenic elements in their definition.
Functional genomics may also include studies of natural genetic variation over time (such as an organism's development) or space (such as its body regions), as well as functional disruptions such as mutations.
The promise of functional genomics is to generate and synthesize genomic and proteomic knowledge into an understanding of the dynamic properties of an organism. This could potentially provide a more complete picture of how the genome specifies function compared to studies of single genes. Integration of functional genomics data is often a part of
systems biology approaches.
Techniques and applications
Functional genomics includes function-related aspects of the genome itself such as
mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA or viral replication, ...
and
polymorphism (such as
single nucleotide polymorphism (SNP) analysis), as well as the measurement of molecular activities. The latter comprise a number of "-
omics
Omics is the collective characterization and quantification of entire sets of biological molecules and the investigation of how they translate into the structure, function, and dynamics of an organism or group of organisms. The branches of scien ...
" such as
transcriptomics (
gene expression
Gene expression is the process (including its Regulation of gene expression, regulation) by which information from a gene is used in the synthesis of a functional gene product that enables it to produce end products, proteins or non-coding RNA, ...
),
proteomics
Proteomics is the large-scale study of proteins. Proteins are vital macromolecules of all living organisms, with many functions such as the formation of structural fibers of muscle tissue, enzymatic digestion of food, or synthesis and replicatio ...
(
protein production), and
metabolomics
Metabolomics is the scientific study of chemical processes involving metabolites, the small molecule substrates, intermediates, and products of cell metabolism. Specifically, metabolomics is the "systematic study of the unique chemical fingerpri ...
. Functional genomics uses mostly
multiplex
Multiplex may refer to:
Science and technology
* Multiplex communication, combining many signals into one transmission circuit or channel
** Multiplex (television), a group of digital television or radio channels that are combined for broadcast
* ...
techniques to measure the abundance of many or all gene products such as
mRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of Protein biosynthesis, synthesizing a protein.
mRNA is ...
s or
protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme catalysis, catalysing metab ...
s within a
biological sample. A more focused functional genomics approach might test the function of all variants of one gene and quantify the effects of mutants by using sequencing as a readout of activity. Together these measurement modalities endeavor to quantitate the various biological processes and improve our understanding of gene and protein functions and interactions.
At the DNA level
Genetic interaction mapping
Systematic pairwise deletion of genes or inhibition of gene expression can be used to identify genes with related function, even if they do not interact physically.
Epistasis refers to the fact that effects for two different gene knockouts may not be additive; that is, the phenotype that results when two genes are inhibited may be different from the sum of the effects of single knockouts.
DNA/Protein interactions
Proteins formed by the translation of the mRNA (messenger RNA, a coded information from DNA for protein synthesis) play a major role in regulating gene expression. To understand how they regulate gene expression it is necessary to identify DNA sequences that they interact with. Techniques have been developed to identify sites of DNA-protein interactions. These include
ChIP-sequencing,
CUT&RUN sequencing and Calling Cards.
DNA accessibility assays
Assays have been developed to identify regions of the genome that are accessible. These regions of accessible chromatin are candidate regulatory regions. These assays include
ATAC-seq,
DNase-Seq and
FAIRE-Seq.
At the RNA level
Microarrays

Microarrays measure the amount of mRNA in a sample that corresponds to a given gene or probe DNA sequence. Probe sequences are immobilized on a solid surface and allowed to
hybridize with fluorescently labeled "target" mRNA. The intensity of fluorescence of a spot is proportional to the amount of target sequence that has hybridized to that spot and therefore to the abundance of that mRNA sequence in the sample. Microarrays allow for the identification of candidate genes involved in a given process based on variation between transcript levels for different conditions and shared expression patterns with genes of known function.
SAGE
Serial analysis of gene expression (SAGE) is an alternate method of analysis based on RNA sequencing rather than hybridization. SAGE relies on the sequencing of 10–17 base pair tags which are unique to each gene. These tags are produced from
poly-A mRNA and ligated end-to-end before sequencing. SAGE gives an unbiased measurement of the number of transcripts per cell, since it does not depend on prior knowledge of what transcripts to study (as microarrays do).
RNA sequencing
RNA sequencing has taken over microarray and SAGE technology in recent years, as noted in 2016, and has become the most efficient way to study transcription and gene expression. This is typically done by
next-generation sequencing.
A subset of sequenced RNAs are small RNAs, a class of non-coding RNA molecules that are key regulators of transcriptional and post-transcriptional gene silencing, or
RNA silencing
RNA silencing or RNA interference refers to a family of gene silencing effects by which gene expression is negatively regulated by non-coding RNAs such as microRNAs. RNA silencing may also be defined as sequence-specific regulation of gene expressi ...
. Next-generation sequencing is the gold standard tool for
non-coding RNA
A non-coding RNA (ncRNA) is a functional RNA molecule that is not Translation (genetics), translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally imp ...
discovery, profiling and expression analysis.
Massively Parallel Reporter Assays (MPRAs)
Massively parallel reporter assays is a technology to test the cis-regulatory activity of DNA sequences. MPRAs use a
plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria and ...
with a synthetic cis-regulatory element upstream of a promoter driving a synthetic gene such as Green Fluorescent Protein. A library of cis-regulatory elements is usually tested using MPRAs, a library can contain from hundreds to thousands of cis-regulatory elements. The cis-regulatory activity of the elements is assayed by using the downstream reporter activity. The activity of all the library members is assayed in parallel using barcodes for each cis-regulatory element. One limitation of MPRAs is that the activity is assayed on a plasmid and may not capture all aspects of gene regulation observed in the genome.
STARR-seq
STARR-seq is a technique similar to MPRAs to assay enhancer activity of randomly sheared genomic fragments. In the original publication, randomly sheared fragments of the ''Drosophila'' genome were placed downstream of a minimal promoter. Candidate enhancers amongst the randomly sheared fragments will transcribe themselves using the minimal promoter. By using sequencing as a readout and controlling for input amounts of each sequence the strength of putative enhancers are assayed by this method.
Perturb-seq

Perturb-seq couples CRISPR mediated gene knockdowns with single-cell gene expression. Linear models are used to calculate the effect of the knockdown of a single gene on the expression of multiple genes.
At the protein level
Yeast two-hybrid system
A yeast
two-hybrid screening (Y2H) tests a "bait" protein against many potential interacting proteins ("prey") to identify physical protein–protein interactions. This system is based on a transcription factor, originally GAL4,
whose separate DNA-binding and transcription activation domains are both required in order for the protein to cause transcription of a reporter gene. In a Y2H screen, the "bait" protein is fused to the binding domain of GAL4, and a library of potential "prey" (interacting) proteins is recombinantly expressed in a vector with the activation domain. In vivo interaction of bait and prey proteins in a yeast cell brings the activation and binding domains of GAL4 close enough together to result in expression of a
reporter gene. It is also possible to systematically test a library of bait proteins against a library of prey proteins to identify all possible interactions in a cell.
MS and AP/MS
Mass spectrometry
Mass spectrometry (MS) is an analytical technique that is used to measure the mass-to-charge ratio of ions. The results are presented as a ''mass spectrum'', a plot of intensity as a function of the mass-to-charge ratio. Mass spectrometry is used ...
(MS) can identify proteins and their relative levels, hence it can be used to study protein expression. When used in combination with
affinity purification,
mass spectrometry
Mass spectrometry (MS) is an analytical technique that is used to measure the mass-to-charge ratio of ions. The results are presented as a ''mass spectrum'', a plot of intensity as a function of the mass-to-charge ratio. Mass spectrometry is used ...
(AP/MS) can be used to study protein complexes, that is, which proteins interact with one another in complexes and in which ratios. In order to purify protein complexes, usually a "bait" protein is tagged with a specific protein or peptide that can be used to pull out the complex from a complex mix. The purification is usually done using an antibody or a compound that binds to the fusion part. The proteins are then digested into short
peptide
Peptides are short chains of amino acids linked by peptide bonds. A polypeptide is a longer, continuous, unbranched peptide chain. Polypeptides that have a molecular mass of 10,000 Da or more are called proteins. Chains of fewer than twenty am ...
fragments and mass spectrometry is used to identify the proteins based on the mass-to-charge ratios of those fragments.
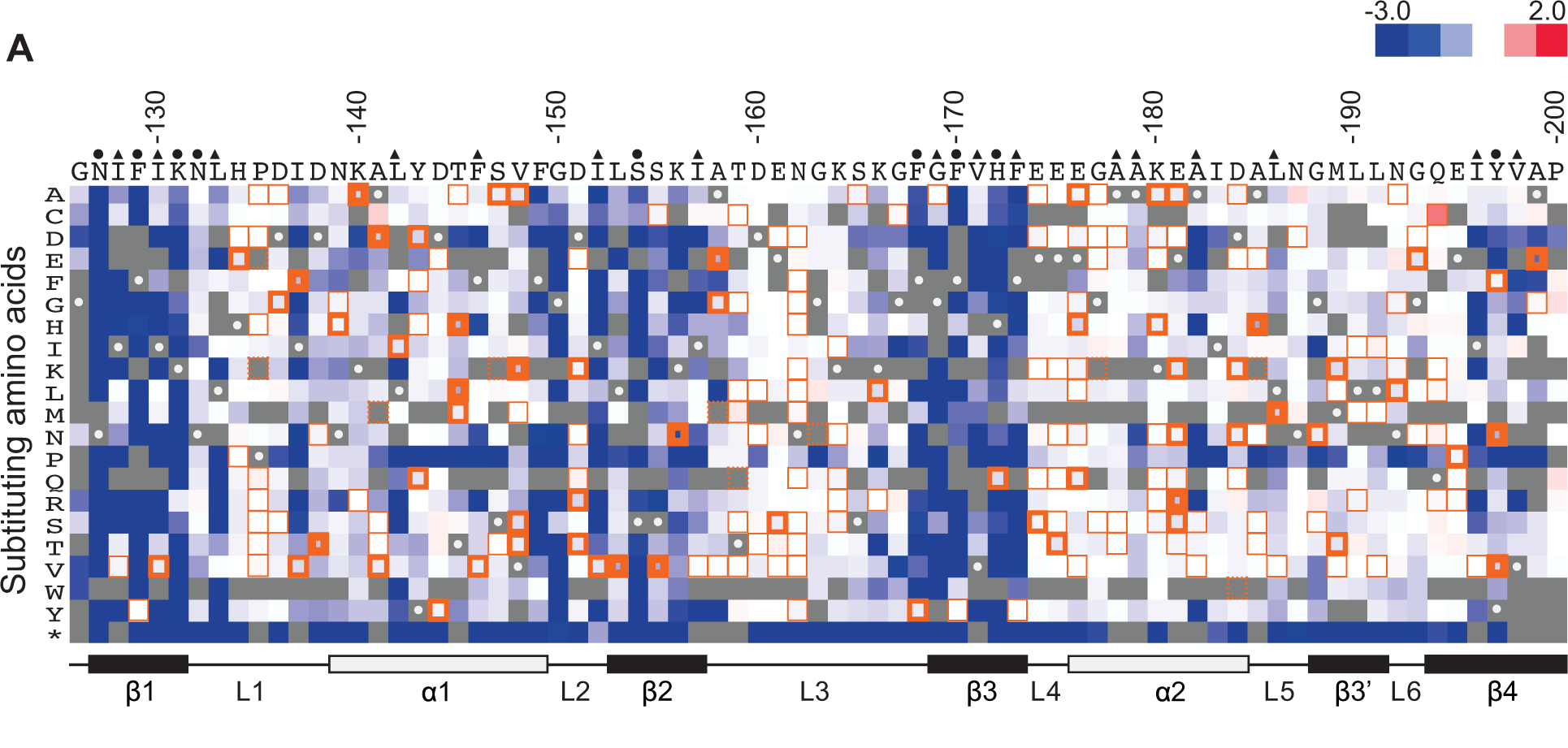
Deep mutational scanning
In deep mutational scanning, every possible amino acid change in a given protein is first synthesized. The activity of each of these protein variants is assayed in parallel using barcodes for each variant. By comparing the activity to the wild-type protein, the effect of each mutation is identified. While it is possible to assay every possible single amino-acid change due to combinatorics two or more concurrent mutations are hard to test. Deep mutational scanning experiments have also been used to infer protein structure and protein-protein interactions. Deep Mutational Scanning is an example of a multiplexed assays of variant effect (MAVEs), a family of methods that involve mutagenesis of a DNA-encoded protein or regulatory element followed by a multiplexed assay for some aspect of function. MAVEs enable the generation of ‘variant effect maps’ characterizing aspects of the function of every possible single nucleotide change in a gene or functional element of interest.
Mutagenesis and phenotyping
An important functional feature of genes is the phenotype caused by mutations. Mutants can be produced by random mutations or by directed mutagenesis, including site-directed mutagenesis, deleting complete genes, or other techniques.
Knock-outs (gene deletions)
Gene function can be investigated by systematically "knocking out" genes one by one. This is done by either
deletion or disruption of function (such as by
insertional mutagenesis) and the resulting organisms are screened for phenotypes that provide clues to the function of the disrupted gene. Knock-outs have been produced for whole genomes, i.e. by deleting all genes in a genome. For
essential genes, this is not possible, so other techniques are used, e.g. deleting a gene while expressing the gene from a
plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria and ...
, using an inducible promoter, so that the level of gene product can be changed at will (and thus a "functional" deletion achieved).
Site-directed mutagenesis
Site-directed mutagenesis is used to mutate specific bases (and thus
amino acid
Amino acids are organic compounds that contain both amino and carboxylic acid functional groups. Although over 500 amino acids exist in nature, by far the most important are the 22 α-amino acids incorporated into proteins. Only these 22 a ...
s). This is critical to investigate the function of specific amino acids in a protein, e.g. in the active site of an
enzyme
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different mol ...
.
RNAi
RNA interference
RNA interference (RNAi) is a biological process in which RNA molecules are involved in sequence-specific suppression of gene expression by double-stranded RNA, through translational or transcriptional repression. Historically, RNAi was known by ...
(RNAi) methods can be used to transiently silence or knockdown gene expression using ~20 base-pair double-stranded RNA typically delivered by transfection of synthetic ~20-mer short-interfering RNA molecules (siRNAs) or by virally encoded short-hairpin RNAs (shRNAs). RNAi screens, typically performed in cell culture-based assays or experimental organisms (such as ''C. elegans'') can be used to systematically disrupt nearly every gene in a genome or subsets of genes (sub-genomes); possible functions of disrupted genes can be assigned based on observed
phenotype
In genetics, the phenotype () is the set of observable characteristics or traits of an organism. The term covers the organism's morphology (physical form and structure), its developmental processes, its biochemical and physiological propert ...
s.
CRISPR screens
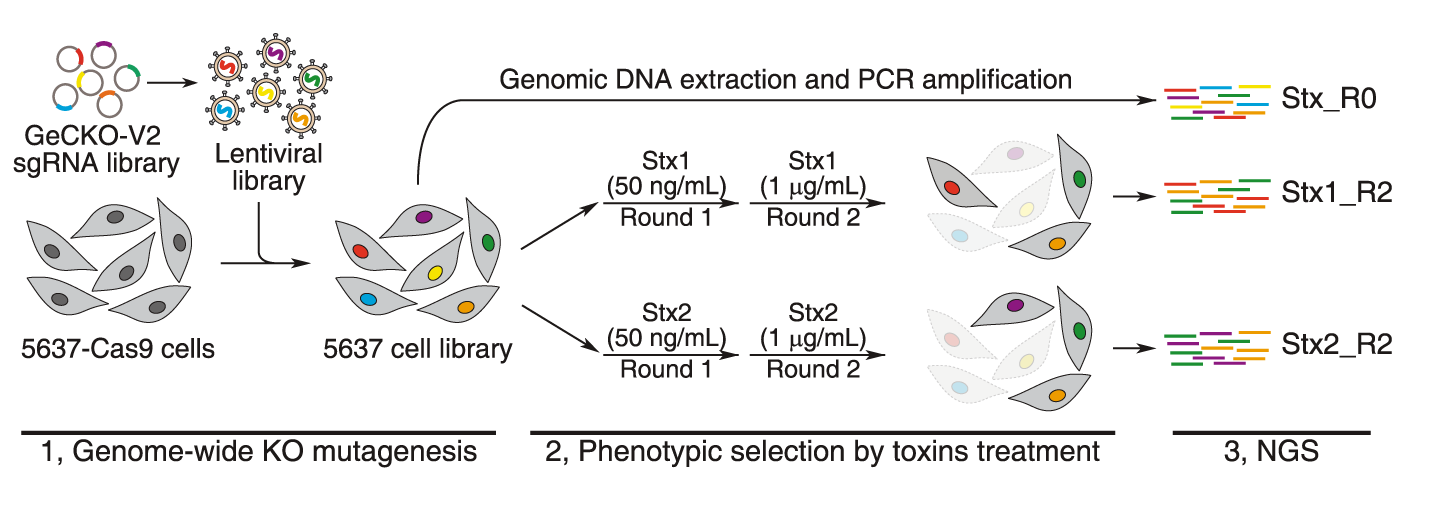
CRISPR-Cas9 has been used to delete genes in a multiplexed manner in cell-lines. Quantifying the amount of guide-RNAs for each gene before and after the experiment can point towards essential genes. If a guide-RNA disrupts an essential gene it will lead to the loss of that cell and hence there will be a depletion of that particular guide-RNA after the screen. In a recent CRISPR-cas9 experiment in mammalian cell-lines, around 2000 genes were found to be essential in multiple cell-lines. Some of these genes were essential in only one cell-line. Most of genes are part of multi-protein complexes. This approach can be used to identify synthetic lethality by using the appropriate genetic background. CRISPRi and CRISPRa enable loss-of-function and gain-of-function screens in a similar manner. CRISPRi identified ~2100 essential genes in the K562 cell-line. CRISPR deletion screens have also been used to identify potential regulatory elements of a gene. For example, a technique called ScanDel was published which attempted this approach. The authors deleted regions outside a gene of interest(HPRT1 involved in a Mendelian disorder) in an attempt to identify regulatory elements of this gene. Gassperini et al. did not identify any distal regulatory elements for HPRT1 using this approach, however such approaches can be extended to other genes of interest.
Functional annotations for genes
Genome annotation
Putative genes can be identified by scanning a genome for regions likely to encode proteins, based on characteristics such as long
open reading frames, transcriptional initiation sequences, and
polyadenylation
Polyadenylation is the addition of a poly(A) tail to an RNA transcript, typically a messenger RNA (mRNA). The poly(A) tail consists of multiple adenosine monophosphates; in other words, it is a stretch of RNA that has only adenine bases. In euka ...
sites. A sequence identified as a putative gene must be confirmed by further evidence, such as similarity to cDNA or EST sequences from the same organism, similarity of the predicted protein sequence to known proteins, association with promoter sequences, or evidence that mutating the sequence produces an observable phenotype.
Rosetta stone approach
The Rosetta stone approach is a computational method for de-novo protein function prediction. It is based on the hypothesis that some proteins involved in a given physiological process may exist as two separate genes in one organism and as a single gene in another. Genomes are scanned for sequences that are independent in one organism and in a single open reading frame in another. If two genes have fused, it is predicted that they have similar biological functions that make such co-regulation advantageous.
Bioinformatics methods for Functional genomics
Because of the large quantity of data produced by these techniques and the desire to find biologically meaningful patterns,
bioinformatics
Bioinformatics () is an interdisciplinary field of science that develops methods and Bioinformatics software, software tools for understanding biological data, especially when the data sets are large and complex. Bioinformatics uses biology, ...
is crucial to analysis of functional genomics data. Examples of techniques in this class are
data clustering
Cluster analysis or clustering is the data analyzing technique in which task of grouping a set of objects in such a way that objects in the same group (called a cluster) are more similar (in some specific sense defined by the analyst) to each o ...
or
principal component analysis
Principal component analysis (PCA) is a linear dimensionality reduction technique with applications in exploratory data analysis, visualization and data preprocessing.
The data is linearly transformed onto a new coordinate system such that th ...
for unsupervised
machine learning
Machine learning (ML) is a field of study in artificial intelligence concerned with the development and study of Computational statistics, statistical algorithms that can learn from data and generalise to unseen data, and thus perform Task ( ...
(class detection) as well as
artificial neural network
In machine learning, a neural network (also artificial neural network or neural net, abbreviated ANN or NN) is a computational model inspired by the structure and functions of biological neural networks.
A neural network consists of connected ...
s or
support vector machines for supervised machine learning (class prediction,
classification
Classification is the activity of assigning objects to some pre-existing classes or categories. This is distinct from the task of establishing the classes themselves (for example through cluster analysis). Examples include diagnostic tests, identif ...
). Functional enrichment analysis is used to determine the extent of over- or under-expression (positive- or negative- regulators in case of RNAi screens) of functional categories relative to a background sets.
Gene ontology based enrichment analysis are provided by
DAVID
David (; , "beloved one") was a king of ancient Israel and Judah and the third king of the United Monarchy, according to the Hebrew Bible and Old Testament.
The Tel Dan stele, an Aramaic-inscribed stone erected by a king of Aram-Dam ...
and
gene set enrichment analysis (GSEA), pathway based analysis by Ingenuity and Pathway studio and protein complex based analysis by COMPLEAT.

New computational methods have been developed for understanding the results of a deep mutational scanning experiment. 'phydms' compares the result of a deep mutational scanning experiment to a phylogenetic tree.
This allows the user to infer if the selection process in nature applies similar constraints on a protein as the results of the deep mutational scan indicate. This may allow an experimenter to choose between different experimental conditions based on how well they reflect nature. Deep mutational scanning has also been used to infer protein-protein interactions. The authors used a thermodynamic model to predict the effects of mutations in different parts of a dimer. Deep mutational structure can also be used to infer protein structure. Strong positive epistasis between two mutations in a deep mutational scan can be indicative of two parts of the protein that are close to each other in 3-D space. This information can then be used to infer protein structure. A proof of principle of this approach was shown by two groups using the protein GB1.
Results from MPRA experiments have required machine learning approaches to interpret the data. A gapped k-mer SVM model has been used to infer the kmers that are enriched within cis-regulatory sequences with high activity compared to sequences with lower activity. These models provide high predictive power. Deep learning and random forest approaches have also been used to interpret the results of these high-dimensional experiments. These models are beginning to help develop a better understanding of
non-coding DNA
Non-coding DNA (ncDNA) sequences are components of an organism's DNA that do not encode protein sequences. Some non-coding DNA is transcribed into functional non-coding RNA molecules (e.g. transfer RNA, microRNA, piRNA, ribosomal RNA, and reg ...
function towards gene-regulation.
Consortium projects
The ENCODE project
The
ENCODE (Encyclopedia of DNA elements) project is an in-depth analysis of the human genome whose goal is to identify all the functional elements of genomic DNA, in both coding and non-coding regions. Important results include evidence from genomic tiling arrays that most nucleotides are transcribed as coding transcripts, non-coding RNAs, or random transcripts, the discovery of additional transcriptional regulatory sites, further elucidation of chromatin-modifying mechanisms.
The Genotype-Tissue Expression (GTEx) project
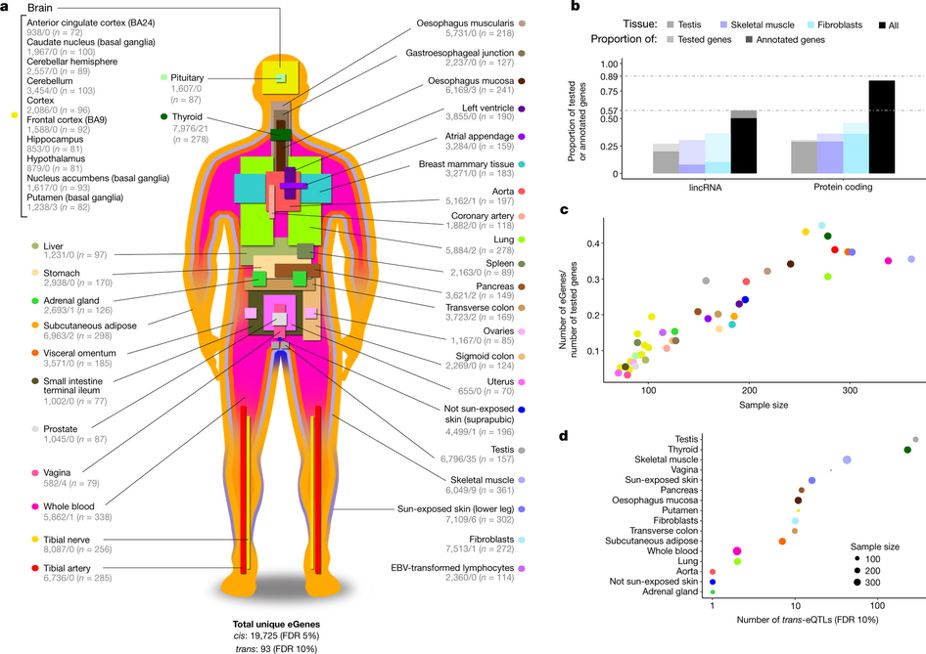
The GTEx project is a human genetics project aimed at understanding the role of genetic variation in shaping variation in the transcriptome across tissues. The project has collected a variety of tissue samples (> 50 different tissues) from more than 700 post-mortem donors. This has resulted in the collection of >11,000 samples. GTEx has helped understand the tissue-sharing and tissue-specificity of
eQTLs. The genomic resource was developed to "enrich our understanding of how differences in our DNA sequence contribute to health and disease."
The Atlas of Variant Effects Alliance The Atlas of Variant Effects Alliance
(AVE),
See also
*
Systems biology
*
Structural genomics
*
Comparative genomics
Comparative genomics is a branch of biological research that examines genome sequences across a spectrum of species, spanning from humans and mice to a diverse array of organisms from bacteria to chimpanzees. This large-scale holistic approach c ...
*
Pharmacogenomics
*
MGED Society
*
Epigenetics
In biology, epigenetics is the study of changes in gene expression that happen without changes to the DNA sequence. The Greek prefix ''epi-'' (ἐπι- "over, outside of, around") in ''epigenetics'' implies features that are "on top of" or "in ...
*
Bioinformatics
Bioinformatics () is an interdisciplinary field of science that develops methods and Bioinformatics software, software tools for understanding biological data, especially when the data sets are large and complex. Bioinformatics uses biology, ...
*
Epistasis and functional genomics
*
Synthetic viability
*
Protein function prediction
References
External links
European Science Foundation Programme on Frontiers of Functional GenomicsMUGEN NoE— Integrated Functional Genomics in Mutant Mouse Models
Nature insights: functional genomicsENCODE
{{DEFAULTSORT:Functional Genomics
Molecular biology
Genomics
 Functional genomics is a field of
Functional genomics is a field of  Perturb-seq couples CRISPR mediated gene knockdowns with single-cell gene expression. Linear models are used to calculate the effect of the knockdown of a single gene on the expression of multiple genes.
Perturb-seq couples CRISPR mediated gene knockdowns with single-cell gene expression. Linear models are used to calculate the effect of the knockdown of a single gene on the expression of multiple genes.
 CRISPR-Cas9 has been used to delete genes in a multiplexed manner in cell-lines. Quantifying the amount of guide-RNAs for each gene before and after the experiment can point towards essential genes. If a guide-RNA disrupts an essential gene it will lead to the loss of that cell and hence there will be a depletion of that particular guide-RNA after the screen. In a recent CRISPR-cas9 experiment in mammalian cell-lines, around 2000 genes were found to be essential in multiple cell-lines. Some of these genes were essential in only one cell-line. Most of genes are part of multi-protein complexes. This approach can be used to identify synthetic lethality by using the appropriate genetic background. CRISPRi and CRISPRa enable loss-of-function and gain-of-function screens in a similar manner. CRISPRi identified ~2100 essential genes in the K562 cell-line. CRISPR deletion screens have also been used to identify potential regulatory elements of a gene. For example, a technique called ScanDel was published which attempted this approach. The authors deleted regions outside a gene of interest(HPRT1 involved in a Mendelian disorder) in an attempt to identify regulatory elements of this gene. Gassperini et al. did not identify any distal regulatory elements for HPRT1 using this approach, however such approaches can be extended to other genes of interest.
CRISPR-Cas9 has been used to delete genes in a multiplexed manner in cell-lines. Quantifying the amount of guide-RNAs for each gene before and after the experiment can point towards essential genes. If a guide-RNA disrupts an essential gene it will lead to the loss of that cell and hence there will be a depletion of that particular guide-RNA after the screen. In a recent CRISPR-cas9 experiment in mammalian cell-lines, around 2000 genes were found to be essential in multiple cell-lines. Some of these genes were essential in only one cell-line. Most of genes are part of multi-protein complexes. This approach can be used to identify synthetic lethality by using the appropriate genetic background. CRISPRi and CRISPRa enable loss-of-function and gain-of-function screens in a similar manner. CRISPRi identified ~2100 essential genes in the K562 cell-line. CRISPR deletion screens have also been used to identify potential regulatory elements of a gene. For example, a technique called ScanDel was published which attempted this approach. The authors deleted regions outside a gene of interest(HPRT1 involved in a Mendelian disorder) in an attempt to identify regulatory elements of this gene. Gassperini et al. did not identify any distal regulatory elements for HPRT1 using this approach, however such approaches can be extended to other genes of interest.
 New computational methods have been developed for understanding the results of a deep mutational scanning experiment. 'phydms' compares the result of a deep mutational scanning experiment to a phylogenetic tree. This allows the user to infer if the selection process in nature applies similar constraints on a protein as the results of the deep mutational scan indicate. This may allow an experimenter to choose between different experimental conditions based on how well they reflect nature. Deep mutational scanning has also been used to infer protein-protein interactions. The authors used a thermodynamic model to predict the effects of mutations in different parts of a dimer. Deep mutational structure can also be used to infer protein structure. Strong positive epistasis between two mutations in a deep mutational scan can be indicative of two parts of the protein that are close to each other in 3-D space. This information can then be used to infer protein structure. A proof of principle of this approach was shown by two groups using the protein GB1.
Results from MPRA experiments have required machine learning approaches to interpret the data. A gapped k-mer SVM model has been used to infer the kmers that are enriched within cis-regulatory sequences with high activity compared to sequences with lower activity. These models provide high predictive power. Deep learning and random forest approaches have also been used to interpret the results of these high-dimensional experiments. These models are beginning to help develop a better understanding of
New computational methods have been developed for understanding the results of a deep mutational scanning experiment. 'phydms' compares the result of a deep mutational scanning experiment to a phylogenetic tree. This allows the user to infer if the selection process in nature applies similar constraints on a protein as the results of the deep mutational scan indicate. This may allow an experimenter to choose between different experimental conditions based on how well they reflect nature. Deep mutational scanning has also been used to infer protein-protein interactions. The authors used a thermodynamic model to predict the effects of mutations in different parts of a dimer. Deep mutational structure can also be used to infer protein structure. Strong positive epistasis between two mutations in a deep mutational scan can be indicative of two parts of the protein that are close to each other in 3-D space. This information can then be used to infer protein structure. A proof of principle of this approach was shown by two groups using the protein GB1.
Results from MPRA experiments have required machine learning approaches to interpret the data. A gapped k-mer SVM model has been used to infer the kmers that are enriched within cis-regulatory sequences with high activity compared to sequences with lower activity. These models provide high predictive power. Deep learning and random forest approaches have also been used to interpret the results of these high-dimensional experiments. These models are beginning to help develop a better understanding of  The GTEx project is a human genetics project aimed at understanding the role of genetic variation in shaping variation in the transcriptome across tissues. The project has collected a variety of tissue samples (> 50 different tissues) from more than 700 post-mortem donors. This has resulted in the collection of >11,000 samples. GTEx has helped understand the tissue-sharing and tissue-specificity of eQTLs. The genomic resource was developed to "enrich our understanding of how differences in our DNA sequence contribute to health and disease."
The GTEx project is a human genetics project aimed at understanding the role of genetic variation in shaping variation in the transcriptome across tissues. The project has collected a variety of tissue samples (> 50 different tissues) from more than 700 post-mortem donors. This has resulted in the collection of >11,000 samples. GTEx has helped understand the tissue-sharing and tissue-specificity of eQTLs. The genomic resource was developed to "enrich our understanding of how differences in our DNA sequence contribute to health and disease."