|
Allosteric
In the fields of biochemistry and pharmacology an allosteric regulator (or allosteric modulator) is a substance that binds to a site on an enzyme or receptor distinct from the active site, resulting in a conformational change that alters the protein's activity, either enhancing or inhibiting its function. In contrast, substances that bind directly to an enzyme's active site or the binding site of the endogenous ligand of a receptor are called orthosteric regulators or modulators. The site to which the effector binds is termed the ''allosteric site'' or ''regulatory site''. Allosteric sites allow effectors to bind to the protein, often resulting in a conformational change and/or a change in protein dynamics. Effectors that enhance the protein's activity are referred to as ''allosteric activators'', whereas those that decrease the protein's activity are called ''allosteric inhibitors''. Allosteric regulations are a natural example of control loops, such as feedback from downs ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Allosteric Regulation
In the fields of biochemistry and pharmacology an allosteric regulator (or allosteric modulator) is a substance that binds to a site on an enzyme or receptor distinct from the active site, resulting in a conformational change that alters the protein's activity, either enhancing or inhibiting its function. In contrast, substances that bind directly to an enzyme's active site or the binding site of the endogenous ligand of a receptor are called orthosteric regulators or modulators. The site to which the effector binds is termed the ''allosteric site'' or ''regulatory site''. Allosteric sites allow effectors to bind to the protein, often resulting in a conformational change and/or a change in protein dynamics. Effectors that enhance the protein's activity are referred to as ''allosteric activators'', whereas those that decrease the protein's activity are called ''allosteric inhibitors''. Allosteric regulations are a natural example of control loops, such as feedback from do ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Jean-Pierre Changeux
Jean-Pierre Changeux (; born 6 April 1936) is a French neuroscientist known for his research in several fields of biology, from the structure and function of proteins (with a focus on the allosteric proteins), to the early development of the nervous system up to cognitive functions. Although being famous in biological sciences for the MWC model, the identification and purification of the nicotinic acetylcholine receptor, the theory of epigenesis by synapse selection and the global neuronal workspace theory for conscious processing are also notable scientific achievements. Changeux is known by the non-scientific public for his ideas regarding the connection between mind and physical brain. As put forth in his book, ''Conversations on Mind, Matter and Mathematics'', Changeux strongly supports the view that the nervous system functions in a projective rather than reactive style and that interaction with the environment, rather than being instructive, results in the selection amongst ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Morpheein
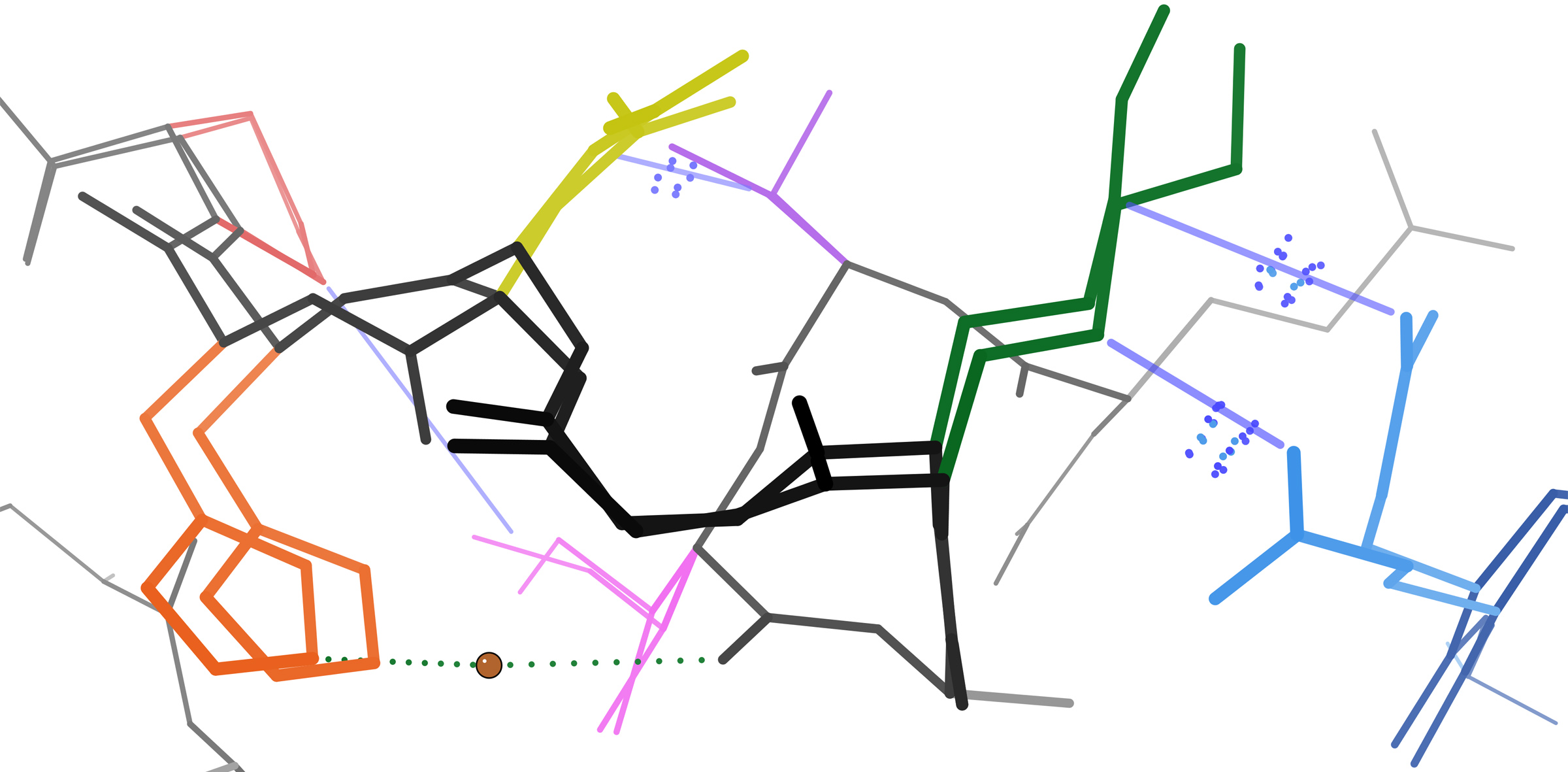
Morpheeins are proteins that can form two or more different homo-oligomers (morpheein forms), but must come apart and change shape to convert between forms. The alternate shape may reassemble to a different oligomer. The shape of the subunit dictates which oligomer is formed. Each oligomer has a finite number of subunits (stoichiometry). Morpheeins can interconvert between forms under physiological conditions and can exist as an equilibrium of different oligomers. These oligomers are physiologically relevant and are not misfolded protein; this distinguishes morpheeins from prions and amyloid. The different oligomers have distinct functionality. Interconversion of morpheein forms can be a structural basis for allosteric regulation, an idea noted many years ago, and later revived. A mutation that shifts the normal equilibrium of morpheein forms can serve as the basis for a conformational disease. Features of morpheeins can be exploited for drug discovery. The dice image (Fig 1) r ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Porphobilinogen Synthase
Aminolevulinic acid dehydratase (porphobilinogen synthase, or ALA dehydratase, or aminolevulinate dehydratase) is an enzyme () that in humans is encoded by the ''ALAD'' gene. Porphobilinogen synthase (or ALA dehydratase, or aminolevulinate dehydratase) synthesizes porphobilinogen through the asymmetric condensation of two molecules of aminolevulinic acid. All natural tetrapyrroles, including hemes, chlorophylls and vitamin B12, share porphobilinogen as a common precursor. Porphobilinogen synthase is the prototype morpheein. Function It catalyzes the following reaction, the second step of the biosynthesis of porphyrin: :2 5- Aminolevulinic acid \rightleftharpoons porphobilinogen + 2 H2O It therefore catalyzes the condensation of 2 molecules of 5-aminolevulinate to form porphobilinogen (a precursor of heme, cytochromes and other hemoproteins). This reaction is the first common step in the biosynthesis of all biological tetrapyrroles. Zinc is essential for enzymatic activity. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzyme Model
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. Almost all metabolic processes in the cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts include catalytic RNA molecules, also called ribozymes. They are sometimes described as a ''type'' of enzyme rather than being ''like'' an enzyme, but even in the decades ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzyme
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different molecules known as product (chemistry), products. Almost all metabolism, metabolic processes in the cell (biology), cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme, pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts include Ribozyme, catalytic RNA molecules, also called ribozymes. They are sometimes descr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Dynamics
In molecular biology, proteins are generally thought to adopt unique structures determined by their amino acid sequences. However, proteins are not strictly static objects, but rather populate ensembles of (sometimes similar) conformations. Transitions between these states occur on a variety of length scales (tenths of angstroms to nm) and time scales (ns to s), and have been linked to functionally relevant phenomena such as allosteric signaling and enzyme catalysis. The study of protein dynamics is most directly concerned with the transitions between these states, but can also involve the nature and equilibrium populations of the states themselves. These two perspectives— kinetics and thermodynamics, respectively—can be conceptually synthesized in an " energy landscape" paradigm: highly populated states and the kinetics of transitions between them can be described by the depths of energy wells and the heights of energy barriers, respectively. Local flexibility: atoms ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Active Site
In biology and biochemistry, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of amino acid residues that form temporary bonds with the substrate, the ''binding site'', and residues that catalyse a reaction of that substrate, the ''catalytic site''. Although the active site occupies only ~10–20% of the volume of an enzyme, it is the most important part as it directly catalyzes the chemical reaction. It usually consists of three to four amino acids, while other amino acids within the protein are required to maintain the tertiary structure of the enzymes. Each active site is evolved to be optimised to bind a particular substrate and catalyse a particular reaction, resulting in high specificity. This specificity is determined by the arrangement of amino acids within the active site and the structure of the substrates. Sometimes enzymes also need to bind with some cofactors to fulfil their functio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell Signaling
In biology, cell signaling (cell signalling in British English) is the Biological process, process by which a Cell (biology), cell interacts with itself, other cells, and the environment. Cell signaling is a fundamental property of all Cell (biology), cellular life in both prokaryotes and eukaryotes. Typically, the signaling process involves three components: the signal, the receptor, and the effector. In biology, signals are mostly chemical in nature, but can also be physical cues such as pressure, Membrane potential, voltage, temperature, or light. Chemical signals are molecules with the ability to bind and activate a specific Receptor (biochemistry), receptor. These molecules, also referred to as Ligand (biochemistry), ligands, are chemically diverse, including ions (e.g. Na+, K+, Ca2+, etc.), lipids (e.g. steroid, prostaglandin), peptides (e.g. insulin, ACTH), carbohydrates, glycosylated proteins (proteoglycans), nucleic acids, etc. Peptide and lipid ligands are particularly ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Jacques Monod
Jacques Lucien Monod (; 9 February 1910 – 31 May 1976) was a French biochemist who won the Nobel Prize in Physiology or Medicine in 1965, sharing it with François Jacob and André Lwoff "for their discoveries concerning genetic control of enzyme and virus synthesis".'' Chance and Necessity: An Essay on the Natural Philosophy of Modern Biology'' by Jacques Monod, New York, Alfred A. Knopf, 1971, Monod and Jacob became famous for their work on the '' E. coli'' ''lac'' operon, which encodes proteins necessary for the transport and breakdown of the sugar lactose (lac). From their own work and the work of others, they came up with a model for how the levels of some proteins in a cell are controlled. In their model, the manufacture of a set of related proteins, such as the ones encoded within the ''lac'' (lactose) operon, is prevented when a repressor protein, encoded by a regulatory gene, binds to its operator, a specific site in the DNA sequence that is close to the genes en ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequential Model
The sequential model (also known as the KNF model) is a theory that describes cooperativity of protein subunits. Koshland, D.E., Némethy, G. and Filmer, D. (1966) Comparison of experimental binding data and theoretical models in proteins containing subunits. Biochemistry 5, 365–385DOI: 10.1021/bi00865a047/ref> It postulates that a protein's conformation changes with each binding of a ligand, thus sequentially changing its affinity for the ligand at neighboring binding sites. It gives one explanation for cooperative binding. Overview This model for allosteric regulation of enzymes suggests that the subunits of multimeric proteins have two conformational states. The binding of the ligand causes conformational change in the other subunits of the multimeric protein. Although the subunits go through conformational changes independently (as opposed to in the MWC model), the switch of one subunit makes the other subunits more likely to change, by reducing the energy needed for subs ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Endogenous Ligand
In biochemistry and pharmacology, a ligand is a substance that forms a complex with a biomolecule to serve a biological purpose. The etymology stems from Latin ''ligare'', which means 'to bind'. In protein-ligand binding, the ligand is usually a molecule which produces a signal by binding to a site on a target protein. The binding typically results in a change of conformational isomerism (conformation) of the target protein. In DNA-ligand binding studies, the ligand can be a small molecule, ion, or protein which binds to the DNA double helix. The relationship between ligand and binding partner is a function of charge, hydrophobicity, and molecular structure. Binding occurs by intermolecular forces, such as ionic bonds, hydrogen bonds and Van der Waals forces. The association or docking is actually reversible through dissociation. Measurably irreversible covalent bonding between a ligand and target molecule is atypical in biological systems. In contrast to the definition of l ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |