|
Genome Assembly
In bioinformatics, sequence assembly refers to aligning and merging fragments from a longer DNA sequence in order to reconstruct the original sequence. This is needed as DNA sequencing technology might not be able to 'read' whole genomes in one go, but rather reads small pieces of between 20 and 30,000 bases, depending on the technology used. Typically, the short fragments (reads) result from shotgun sequencing genomic DNA, or gene transcript ( ESTs). The problem of sequence assembly can be compared to taking many copies of a book, passing each of them through a shredder with a different cutter, and piecing the text of the book back together just by looking at the shredded pieces. Besides the obvious difficulty of this task, there are some extra practical issues: the original may have many repeated paragraphs, and some shreds may be modified during shredding to have typos. Excerpts from another book may also be added in, and some shreds may be completely unrecognizable. Typ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bioinformatics
Bioinformatics () is an interdisciplinary field of science that develops methods and Bioinformatics software, software tools for understanding biological data, especially when the data sets are large and complex. Bioinformatics uses biology, chemistry, physics, computer science, data science, computer programming, information engineering, mathematics and statistics to analyze and interpret biological data. The process of analyzing and interpreting data can sometimes be referred to as computational biology, however this distinction between the two terms is often disputed. To some, the term ''computational biology'' refers to building and using models of biological systems. Computational, statistical, and computer programming techniques have been used for In silico, computer simulation analyses of biological queries. They include reused specific analysis "pipelines", particularly in the field of genomics, such as by the identification of genes and single nucleotide polymorphis ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
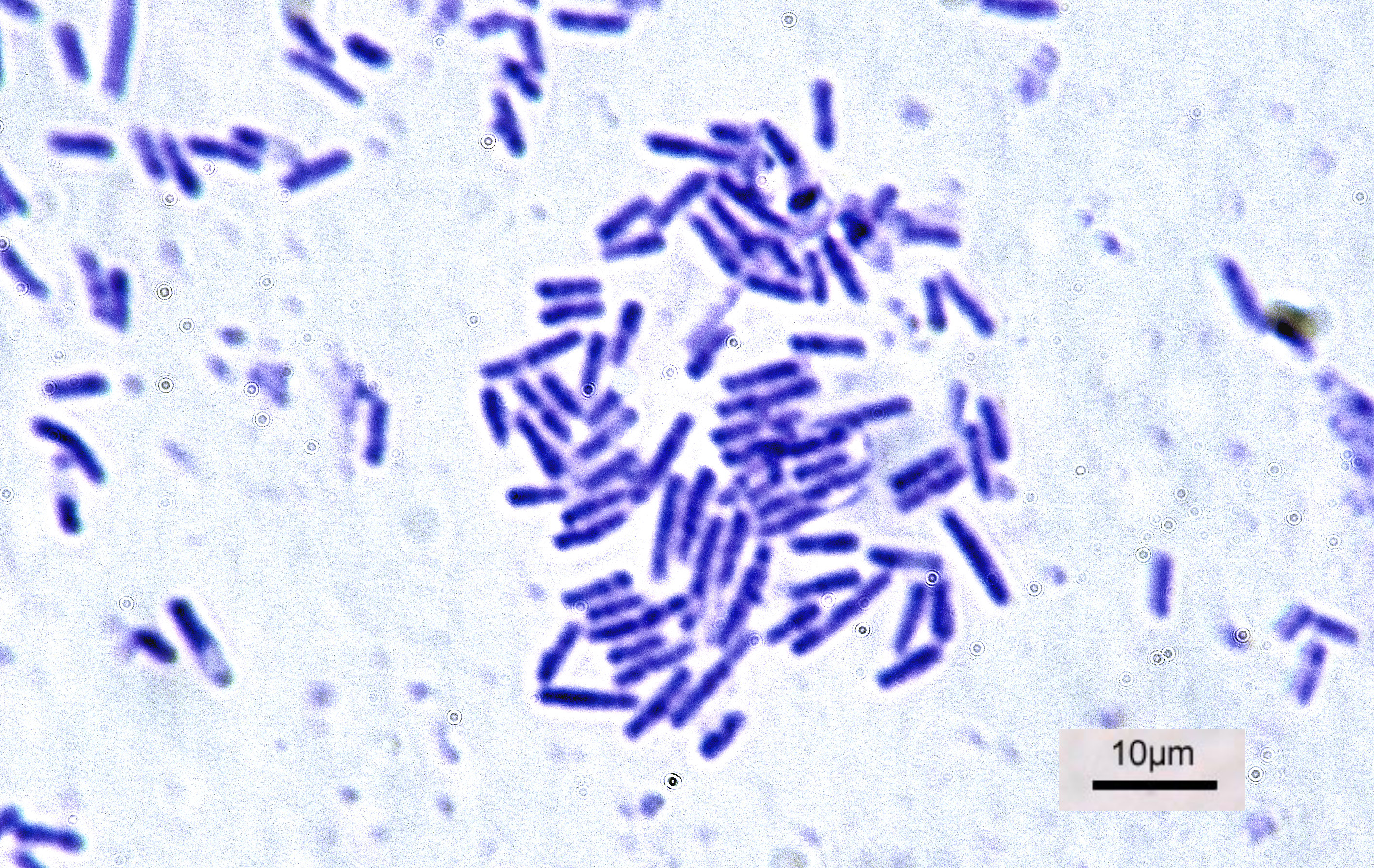
Bacteria
Bacteria (; : bacterium) are ubiquitous, mostly free-living organisms often consisting of one Cell (biology), biological cell. They constitute a large domain (biology), domain of Prokaryote, prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit the air, soil, water, Hot spring, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria play a vital role in many stages of the nutrient cycle by recycling nutrients and the nitrogen fixation, fixation of nitrogen from the Earth's atmosphere, atmosphere. The nutrient cycle includes the decomposition of cadaver, dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, suc ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Trans-splicing
''Trans''-splicing is a special form of RNA processing where exons from two different primary RNA transcripts are joined end to end and ligated. It is usually found in eukaryotes and mediated by the spliceosome, although some bacteria and archaea also have "half-genes" for tRNAs. Genic ''trans''-splicing Whereas "normal" (''cis''-)splicing processes a single molecule, ''trans''-splicing generates a single RNA transcript from multiple separate pre-mRNAs. This phenomenon can be exploited for molecular therapy to address mutated gene products. Genic trans-splicing allows variability in RNA diversity and increases proteome complexity. Oncogenesis While some fusion transcripts occur via ''trans''-splicing in normal human cells, ''trans''-splicing can also be the mechanism behind certain oncogenic fusion transcripts. SL ''trans''-splicing Spliced leader (SL) ''trans''-splicing is used by certain microorganisms, notably protists of the Kinetoplastea class to express genes. I ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
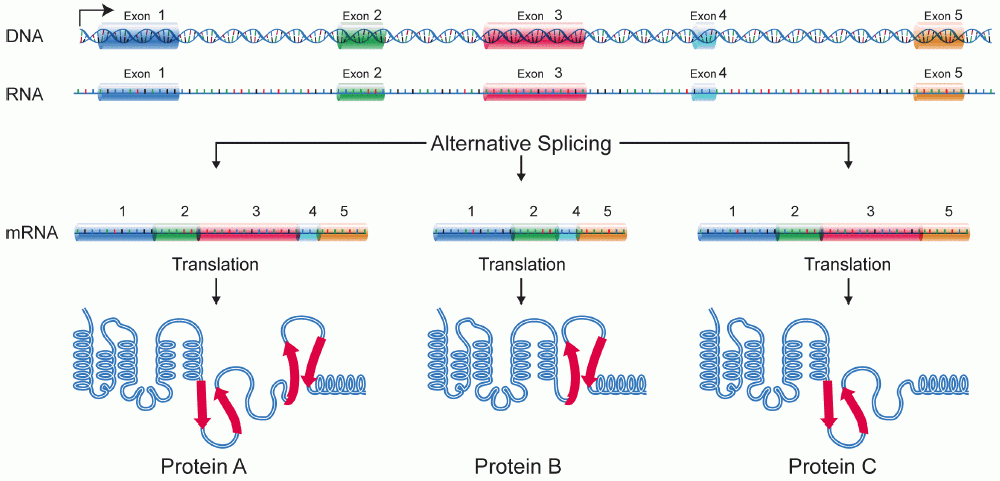
Alternative Splicing
Alternative splicing, alternative RNA splicing, or differential splicing, is an alternative RNA splicing, splicing process during gene expression that allows a single gene to produce different splice variants. For example, some exons of a gene may be included within or excluded from the final RNA product of the gene. This means the exons are joined in different combinations, leading to different splice variants. In the case of protein-coding genes, the proteins translated from these splice variants may contain differences in their amino acid sequence and in their biological functions (see Figure). Biologically relevant alternative splicing occurs as a normal phenomenon in eukaryotes, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most of the observed splice variants ar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Housekeeping Gene
In molecular biology, housekeeping genes are typically constitutive genes that are required for the maintenance of basic cellular function, and are expressed in all cells of an organism under normal and patho-physiological conditions. Although some housekeeping genes are expressed at relatively constant rates in most non-pathological situations, the expression of other housekeeping genes may vary depending on experimental conditions. The origin of the term "housekeeping gene" remains obscure. Literature from 1976 used the term to describe specifically tRNA and rRNA. For experimental purposes, the expression of one or multiple housekeeping genes is used as a reference point for the analysis of expression levels of other genes. The key criterion for the use of a housekeeping gene in this manner is that the chosen housekeeping gene is uniformly expressed with low variance under both control and experimental conditions. Validation of housekeeping genes should be performed before t ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Messenger RNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme (RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and the ribosome creates the protein utilizing amino acids carried by transfer RNA (tRNA). This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, genetic inf ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Expressed Sequence Tag
In genetics, an expressed sequence tag (EST) is a short sub-sequence of a cDNA sequence. ESTs may be used to identify gene transcripts, and were instrumental in gene discovery and in gene-sequence determination. The identification of ESTs has proceeded rapidly, with approximately 74.2 million ESTs now available in public databases (e.g. GenBank 1 January 2013, all species). EST approaches have largely been superseded by whole genome and transcriptome sequencing and metagenome sequencing. An EST results from one-shot sequencing of a cloned cDNA. The cDNAs used for EST generation are typically individual clones from a cDNA library. The resulting sequence is a relatively low-quality fragment whose length is limited by current technology to approximately 500 to 800 nucleotides. Because these clones consist of DNA that is complementary to mRNA, the ESTs represent portions of expressed genes. They may be represented in databases as either cDNA/mRNA sequence or as the reverse compleme ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Open-source Software
Open-source software (OSS) is Software, computer software that is released under a Open-source license, license in which the copyright holder grants users the rights to use, study, change, and Software distribution, distribute the software and its source code to anyone and for any purpose. Open-source software may be developed in a collaborative, public manner. Open-source software is a prominent example of open collaboration, meaning any capable user is able to online collaboration, participate online in development, making the number of possible contributors indefinite. The ability to examine the code facilitates public trust in the software. Open-source software development can bring in diverse perspectives beyond those of a single company. A 2024 estimate of the value of open-source software to firms is $8.8 trillion, as firms would need to spend 3.5 times the amount they currently do without the use of open source software. Open-source code can be used for studying and a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Drosophila Melanogaster
''Drosophila melanogaster'' is a species of fly (an insect of the Order (biology), order Diptera) in the family Drosophilidae. The species is often referred to as the fruit fly or lesser fruit fly, or less commonly the "vinegar fly", "pomace fly", or "banana fly". In the wild, ''D. melanogaster'' are attracted to rotting fruit and fermenting beverages, and are often found in orchards, kitchens and pubs. Starting with Charles W. Woodworth's 1901 proposal of the use of this species as a model organism, ''D. melanogaster'' continues to be widely used for biological research in genetics, physiology, microbial pathogenesis, and Life history theory, life history evolution. ''D. melanogaster'' was the first animal to be Fruit flies in space, launched into space in 1947. As of 2017, six Nobel Prizes have been awarded to drosophilists for their work using the insect. ''Drosophila melanogaster'' is typically used in research owing to its rapid life cycle, relatively simple genetics with on ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Read Errors
Deoxyribonucleic acid (; DNA) is a polymer composed of two polynucleotide chains that coil around each other to form a double helix. The polymer carries genetic instructions for the development, functioning, growth and reproduction of all known organisms and many viruses. DNA and ribonucleic acid (RNA) are nucleic acids. Alongside proteins, lipids and complex carbohydrates (polysaccharides), nucleic acids are one of the four major types of macromolecules that are essential for all known forms of life. The two DNA strands are known as polynucleotides as they are composed of simpler monomeric units called nucleotides. Each nucleotide is composed of one of four nitrogen-containing nucleobases (cytosine guanine adenine or thymine , a sugar called deoxyribose, and a phosphate group. The nucleotides are joined to one another in a chain by covalent bonds (known as the phosphodiester linkage) between the sugar of one nucleotide and the phosphate of the next, resulting in an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cluster Computing
A computer cluster is a set of computers that work together so that they can be viewed as a single system. Unlike Grid computing, grid computers, computer clusters have each Node (networking), node set to perform the same task, controlled and scheduled by software. The newest manifestation of cluster computing is cloud computing. The components of a cluster are usually connected to each other through fast local area networks, with each Node (networking), node (computer used as a server) running its own instance of an operating system. In most circumstances, all of the nodes use the same hardware and the same operating system, although in some setups (e.g. using Open Source Cluster Application Resources (OSCAR)), different operating systems can be used on each computer, or different hardware. Clusters are usually deployed to improve performance and availability over that of a single computer, while typically being much more cost-effective than single computers of comparable s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |