|
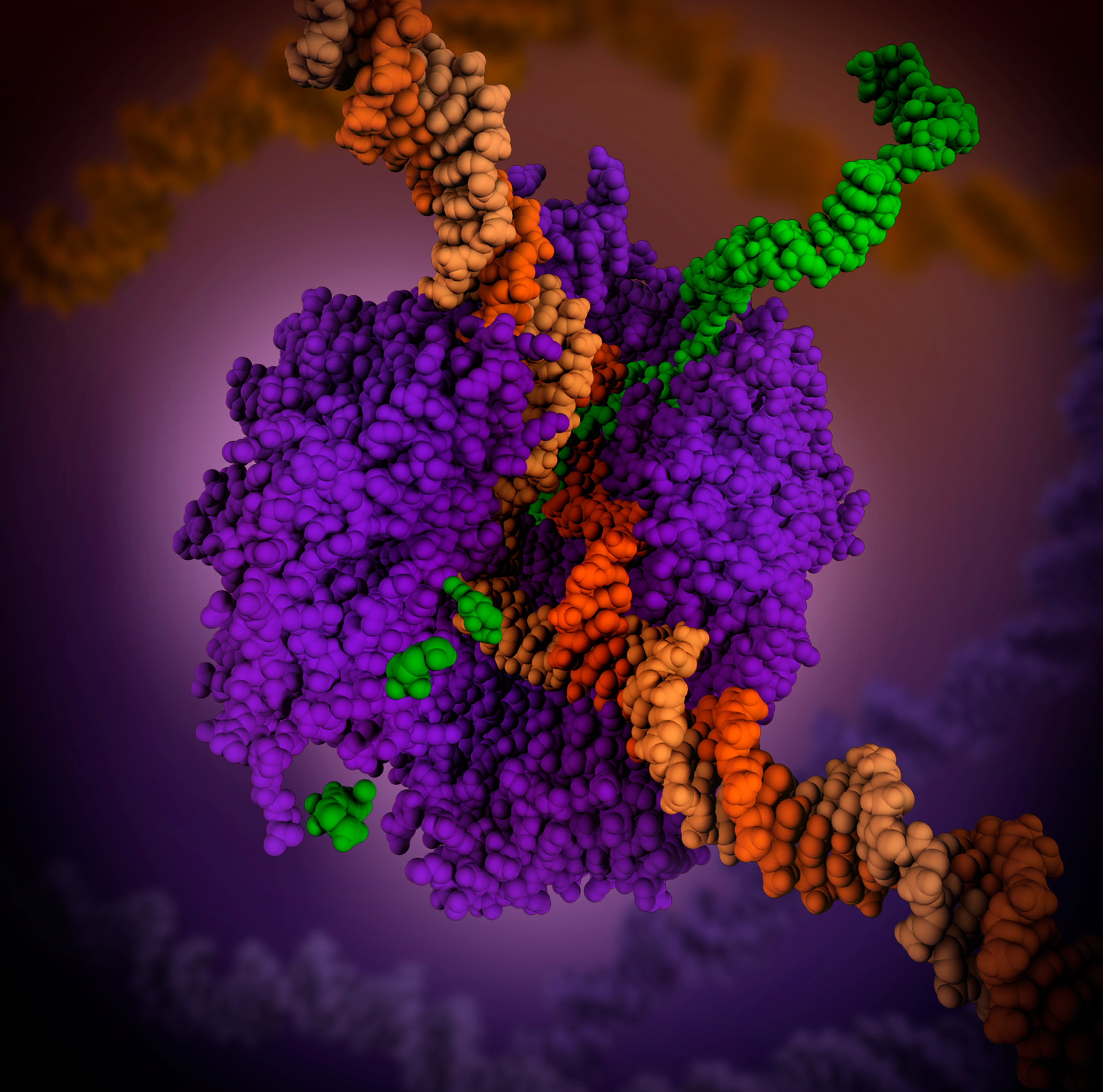
RNA-binding Proteins
RNA-binding proteins (often abbreviated as RBPs) are proteins that bind to the double or single stranded RNA in cells and participate in forming ribonucleoprotein complexes. RBPs contain various structural motifs, such as RNA recognition motif (RRM), dsRNA binding domain, zinc finger and others. They are cytoplasmic and nuclear proteins. However, since most mature RNA is exported from the nucleus relatively quickly, most RBPs in the nucleus exist as complexes of protein and pre-mRNA called heterogeneous ribonucleoprotein particles (hnRNPs). RBPs have crucial roles in various cellular processes such as: cellular function, transport and localization. They especially play a major role in post-transcriptional control of RNAs, such as: splicing, polyadenylation, mRNA stabilization, mRNA localization and translation. Eukaryotic cells express diverse RBPs with unique RNA-binding activity and protein–protein interaction. According to the Eukaryotic RBP Database (EuRBPDB), there are ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Retinol Binding Protein
Retinol-binding proteins (RBP) are a family of proteins with diverse functions. They are carrier proteins that bind retinol. Assessment of retinol-binding protein is used to determine visceral protein mass in health-related nutritional studies. Retinol and retinoic acid play crucial roles in the modulation of gene expression and overall development of an embryo. However, deficit or excess of either one of these substances can cause early embryo mortality or developmental malformations. Regulation of transport and metabolism of retinol necessary for a successful pregnancy is accomplished via RBP. Retinol-binding proteins have been identified within the uterus, embryo, and extraembryonic tissue of the bovine, ovine, and porcine, clearly indicating that RBP plays a role in proper retinol exposure to the embryo and successful transport at the maternal-fetal interface. Further research is necessary to determine the exact effects of poor RBP expression on pregnancy and threshold levels f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Eukaryote
The eukaryotes ( ) constitute the Domain (biology), domain of Eukaryota or Eukarya, organisms whose Cell (biology), cells have a membrane-bound cell nucleus, nucleus. All animals, plants, Fungus, fungi, seaweeds, and many unicellular organisms are eukaryotes. They constitute a major group of Outline of life forms, life forms alongside the two groups of prokaryotes: the Bacteria and the Archaea. Eukaryotes represent a small minority of the number of organisms, but given their generally much larger size, their collective global biomass is much larger than that of prokaryotes. The eukaryotes emerged within the archaeal Kingdom (biology), kingdom Asgard (Archaea), Promethearchaeati and its sole phylum Promethearchaeota. This implies that there are only Two-domain system, two domains of life, Bacteria and Archaea, with eukaryotes incorporated among the Archaea. Eukaryotes first emerged during the Paleoproterozoic, likely as Flagellated cell, flagellated cells. The leading evolutiona ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Exon
An exon is any part of a gene that will form a part of the final mature RNA produced by that gene after introns have been removed by RNA splicing. The term ''exon'' refers to both the DNA sequence within a gene and to the corresponding sequence in RNA transcripts. In RNA splicing, introns are removed and exons are covalently joined to one another as part of generating the mature RNA. Just as the entire set of genes for a species constitutes the genome, the entire set of exons constitutes the exome. History The term ''exon'' is a shortening of the phrase ''expressed region'' and was coined by American biochemist Walter Gilbert in 1978: "The notion of the cistron... must be replaced by that of a transcription unit containing regions which will be lost from the mature messengerwhich I suggest we call introns (for intragenic regions)alternating with regions which will be expressedexons." This definition was originally made for protein-coding transcripts that are spliced before b ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene
In biology, the word gene has two meanings. The Mendelian gene is a basic unit of heredity. The molecular gene is a sequence of nucleotides in DNA that is transcribed to produce a functional RNA. There are two types of molecular genes: protein-coding genes and non-coding genes. During gene expression (the synthesis of Gene product, RNA or protein from a gene), DNA is first transcription (biology), copied into RNA. RNA can be non-coding RNA, directly functional or be the intermediate protein biosynthesis, template for the synthesis of a protein. The transmission of genes to an organism's offspring, is the basis of the inheritance of phenotypic traits from one generation to the next. These genes make up different DNA sequences, together called a genotype, that is specific to every given individual, within the gene pool of the population (biology), population of a given species. The genotype, along with environmental and developmental factors, ultimately determines the phenotype ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
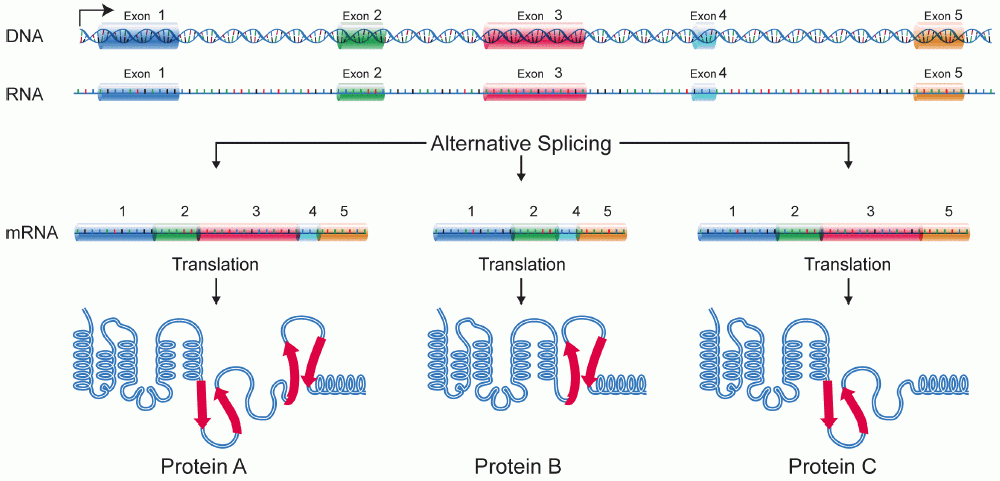
Alternative Splicing
Alternative splicing, alternative RNA splicing, or differential splicing, is an alternative RNA splicing, splicing process during gene expression that allows a single gene to produce different splice variants. For example, some exons of a gene may be included within or excluded from the final RNA product of the gene. This means the exons are joined in different combinations, leading to different splice variants. In the case of protein-coding genes, the proteins translated from these splice variants may contain differences in their amino acid sequence and in their biological functions (see Figure). Biologically relevant alternative splicing occurs as a normal phenomenon in eukaryotes, where it increases the number of proteins that can be encoded by the genome. In humans, it is widely believed that ~95% of multi-exonic genes are alternatively spliced to produce functional alternative products from the same gene but many scientists believe that most of the observed splice variants ar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Nuclear RNA
Small nuclear RNA (snRNA) is a class of small RNA molecules that are found within the Cell nucleus#Splicing speckles, splicing speckles and Cajal body, Cajal bodies of the cell nucleus in eukaryotic cells. The length of an average snRNA is approximately 150 nucleotides. They are transcribed by either RNA polymerase II or RNA polymerase III. Their primary function is in the processing of pre-messenger RNA (hnRNA) in the nucleus. They have also been shown to aid in the regulation of transcription factors (7SK RNA) or RNA polymerase II (B2 RNA), and maintaining the telomeres. snRNA are always associated with a set of specific proteins, and the complexes are referred to as small nuclear ribonucleoproteins (snRNP, often pronounced "snurps"). Each snRNP particle is composed of a snRNA component and several snRNP-specific proteins (including LSm, Sm proteins, a family of nuclear proteins). The most common human snRNA components of these complexes are known, respectively, as: U1 spliceos ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Small Interfering RNA
Small interfering RNA (siRNA), sometimes known as short interfering RNA or silencing RNA, is a class of double-stranded RNA, double-stranded non-coding RNA, non-coding RNA, RNA molecules, typically 20–24 base pairs in length, similar to microRNA (miRNA), and operating within the RNA interference (RNAi) pathway. It interferes with the gene expression, expression of specific genes with complementary nucleotide sequences by degrading messenger RNA (mRNA) after Transcription (biology), transcription, preventing translation (biology), translation. Text was copied from this source, which is available under Creative Commons Attribution 4.0 International License It was discovered in 1998 by Andrew Fire at the Carnegie Institution for Science in Washington, D.C. and Craig Mello at the University of Massachusetts in Worcester. Structure Naturally occurring siRNAs have a well-defined structure that is a short (usually 20 to 24-base pair, bp) double-stranded RNA (dsRNA) with phosph ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MicroRNAs
Micro ribonucleic acid (microRNA, miRNA, μRNA) are small, single-stranded, non-coding RNA molecules containing 21–23 nucleotides. Found in plants, animals, and even some viruses, miRNAs are involved in RNA silencing and post-transcriptional regulation of gene expression. miRNAs base-pair to complementary sequences in messenger RNA (mRNA) molecules, then silence said mRNA molecules by one or more of the following processes: * Cleaving the mRNA strand into two pieces. * Destabilizing the mRNA by shortening its poly(A) tail. * Reducing translation of the mRNA into proteins. In cells of humans and other animals, miRNAs primarily act by destabilizing the mRNA. miRNAs resemble the small interfering RNAs (siRNAs) of the RNA interference (RNAi) pathway, except miRNAs derive from regions of RNA transcripts that fold back on themselves to form short stem-loops (hairpins), whereas siRNAs derive from longer regions of double-stranded RNA. The human genome may encode over 1 ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Non-coding RNA
A non-coding RNA (ncRNA) is a functional RNA molecule that is not Translation (genetics), translated into a protein. The DNA sequence from which a functional non-coding RNA is transcribed is often called an RNA gene. Abundant and functionally important list of RNAs, types of non-coding RNAs include transfer RNAs (tRNAs) and ribosomal RNAs (rRNAs), as well as small RNAs such as microRNAs, siRNAs, piRNAs, snoRNAs, snRNAs, Extracellular RNA, exRNAs, scaRNAs and the long noncoding RNA, long ncRNAs such as Xist and HOTAIR. The number of non-coding RNAs within the human genome is unknown; however, recent Transcriptomics, transcriptomic and Bioinformatics, bioinformatic studies suggest that there are thousands of non-coding transcripts. Many of the newly identified ncRNAs have unknown functions, if any. There is no consensus on how much of non-coding transcription is functional: some believe most ncRNAs to be non-functional "junk RNA", spurious transcriptions, while others expect that ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
RNA Polymerase
In molecular biology, RNA polymerase (abbreviated RNAP or RNApol), or more specifically DNA-directed/dependent RNA polymerase (DdRP), is an enzyme that catalyzes the chemical reactions that synthesize RNA from a DNA template. Using the enzyme helicase, RNAP locally opens the double-stranded DNA so that one strand of the exposed nucleotides can be used as a template for the synthesis of RNA, a process called transcription. A transcription factor and its associated transcription mediator complex must be attached to a DNA binding site called a promoter region before RNAP can initiate the DNA unwinding at that position. RNAP not only initiates RNA transcription, it also guides the nucleotides into position, facilitates attachment and elongation, has intrinsic proofreading and replacement capabilities, and termination recognition capability. In eukaryotes, RNAP can build chains as long as 2.4 million nucleotides. RNAP produces RNA that, functionally, is either for protei ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Introns
An intron is any Nucleic acid sequence, nucleotide sequence within a gene that is not expressed or operative in the final RNA product. The word ''intron'' is derived from the term ''intragenic region'', i.e., a region inside a gene."The notion of the cistron [i.e., gene] ... must be replaced by that of a transcription unit containing regions which will be lost from the mature messenger – which I suggest we call introns (for intragenic regions) – alternating with regions which will be expressed – exons." (Gilbert 1978) The term ''intron'' refers to both the DNA sequence within a gene and the corresponding RNA sequence in RNA Transcription (genetics), transcripts. The non-intron sequences that become joined by this RNA processing to form the mature RNA are called exons. Introns are found in the genes of most eukaryotes and many eukaryotic viruses, and they can be located in both protein-coding genes and genes that function as RNA (Non-coding RNA, noncoding genes). There are f ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Evolution
Evolution is the change in the heritable Phenotypic trait, characteristics of biological populations over successive generations. It occurs when evolutionary processes such as natural selection and genetic drift act on genetic variation, resulting in certain characteristics becoming more or less common within a population over successive generations. The process of evolution has given rise to biodiversity at every level of biological organisation. The scientific theory of evolution by natural selection was conceived independently by two British naturalists, Charles Darwin and Alfred Russel Wallace, in the mid-19th century as an explanation for why organisms are adapted to their physical and biological environments. The theory was first set out in detail in Darwin's book ''On the Origin of Species''. Evolution by natural selection is established by observable facts about living organisms: (1) more offspring are often produced than can possibly survive; (2) phenotypic variatio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |