|
ChIP-sequencing
ChIP-sequencing, also known as ChIP-seq, is a method used to analyze protein interactions with DNA. ChIP-seq combines chromatin immunoprecipitation (ChIP) with massively parallel DNA sequencing to identify the binding sites of DNA-associated proteins. It can be used to map global binding sites precisely for any protein of interest. Previously, ChIP-on-chip was the most common technique utilized to study these protein–DNA relations. Uses ChIP-seq is primarily used to determine how transcription factors and other chromatin-associated proteins influence phenotype-affecting mechanisms. Determining how proteins interact with DNA to regulate gene expression is essential for fully understanding many biological processes and disease states. This epigenetic information is complementary to genotype and expression analysis. ChIP-seq technology is currently seen primarily as an alternative to ChIP-chip which requires a hybridization array. This introduces some bias, as an array is restr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein
Proteins are large biomolecules and macromolecules that comprise one or more long chains of amino acid residue (biochemistry), residues. Proteins perform a vast array of functions within organisms, including Enzyme catalysis, catalysing metabolic reactions, DNA replication, Cell signaling, responding to stimuli, providing Cytoskeleton, structure to cells and Fibrous protein, organisms, and Intracellular transport, transporting molecules from one location to another. Proteins differ from one another primarily in their sequence of amino acids, which is dictated by the Nucleic acid sequence, nucleotide sequence of their genes, and which usually results in protein folding into a specific Protein structure, 3D structure that determines its activity. A linear chain of amino acid residues is called a polypeptide. A protein contains at least one long polypeptide. Short polypeptides, containing less than 20–30 residues, are rarely considered to be proteins and are commonly called pep ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Modification
In molecular biology, post-translational modification (PTM) is the covalent process of changing proteins following protein biosynthesis. PTMs may involve enzymes or occur spontaneously. Proteins are created by ribosomes, which translate mRNA into polypeptide chains, which may then change to form the mature protein product. PTMs are important components in cell signalling, as for example when prohormones are converted to hormones. Post-translational modifications can occur on the amino acid side chains or at the protein's C- or N- termini. They can expand the chemical set of the 22 amino acids by changing an existing functional group or adding a new one such as phosphate. Phosphorylation is highly effective for controlling the enzyme activity and is the most common change after translation. Many eukaryotic and prokaryotic proteins also have carbohydrate molecules attached to them in a process called glycosylation, which can promote protein folding and improve stability as well ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tiling Array
Tiling arrays are a subtype of microarray chips. Like traditional microarrays, they function by hybridizing labeled DNA or RNA target molecules to probes fixed onto a solid surface. Tiling arrays differ from traditional microarrays in the nature of the probes. Instead of probing for sequences of known or predicted genes that may be dispersed throughout the genome, tiling arrays probe intensively for sequences which are known to exist in a contiguous region. This is useful for characterizing regions that are sequenced, but whose local functions are largely unknown. Tiling arrays aid in transcriptome mapping as well as in discovering sites of DNA/protein interaction (ChIP-chip, DamID), of DNA methylation (MeDIP-chip) and of sensitivity to DNase (DNase Chip) and array CGH. In addition to detecting previously unidentified genes and regulatory sequences, improved quantification of transcription products is possible. Specific probes are present in millions of copies (as opposed to only ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Massive Parallel Sequencing
Massive parallel sequencing or massively parallel sequencing is any of several high-throughput approaches to DNA sequencing using the concept of massively parallel processing; it is also called next-generation sequencing (NGS) or second-generation sequencing. Some of these technologies emerged between 1993 and 1998 and have been commercially available since 2005. These technologies use miniaturized and parallelized platforms for sequencing of 1 million to 43 billion short reads (50 to 400 bases each) per instrument run. Many NGS platforms differ in engineering configurations and sequencing chemistry. They share the technical paradigm of massive parallel sequencing via spatially separated, clonally amplified DNA templates or single DNA molecules in a flow cell. This design is very different from that of Sanger sequencing—also known as capillary sequencing or first-generation sequencing—which is based on electrophoretic separation of chain-termination products produced in ind ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Oligonucleotide
Oligonucleotides are short DNA or RNA molecules, oligomers, that have a wide range of applications in genetic testing, Recombinant DNA, research, and Forensic DNA, forensics. Commonly made in the laboratory by Oligonucleotide synthesis, solid-phase chemical synthesis, these small fragments of nucleic acids can be manufactured as single-stranded molecules with any user-specified sequence, and so are vital for artificial gene synthesis, polymerase chain reaction (PCR), DNA sequencing, molecular cloning and as Fluorescence in situ hybridization, molecular probes. In nature, oligonucleotides are usually found as small RNA molecules that function in the regulation of gene expression (e.g. microRNA), or are degradation intermediates derived from the breakdown of larger nucleic acid molecules. Oligonucleotides are characterized by the Nucleic acid sequence, sequence of nucleotide residues that make up the entire molecule. The length of the oligonucleotide is usually denoted by "Monomer, -m ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
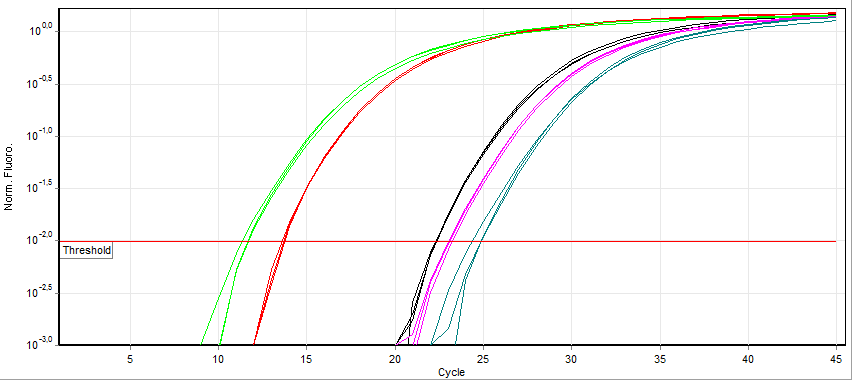
Real-time Polymerase Chain Reaction
A real-time polymerase chain reaction (real-time PCR, or qPCR when used quantitatively) is a laboratory technique of molecular biology based on the polymerase chain reaction (PCR). It monitors the amplification of a targeted DNA molecule during the PCR (i.e., in real time), not at its end, as in conventional PCR. Real-time PCR can be used quantitatively and semi-quantitatively (i.e., above/below a certain amount of DNA molecules). Two common methods for the detection of PCR products in real-time PCR are (1) non-specific fluorescent dyes that Intercalation (biochemistry), intercalate with any double-stranded DNA and (2) sequence-specific DNA probes consisting of oligonucleotides that are labelled with a fluorescence, fluorescent reporter, which permits detection only after nucleic acid hybridisation, hybridization of the probe with its complementary sequence. The Minimum Information for Publication of Quantitative Real-Time PCR Experiments (MIQE) guidelines propose that the abb ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Antibody
An antibody (Ab) or immunoglobulin (Ig) is a large, Y-shaped protein belonging to the immunoglobulin superfamily which is used by the immune system to identify and neutralize antigens such as pathogenic bacteria, bacteria and viruses, including those that cause disease. Each individual antibody recognizes one or more specific antigens, and antigens of virtually any size and chemical composition can be recognized. Antigen literally means "antibody generator", as it is the presence of an antigen that drives the formation of an antigen-specific antibody. Each of the branching chains comprising the "Y" of an antibody contains a paratope that specifically binds to one particular epitope on an antigen, allowing the two molecules to bind together with precision. Using this mechanism, antibodies can effectively "tag" the antigen (or a microbe or an infected cell bearing such an antigen) for attack by cells of the immune system, or can neutralize it directly (for example, by blocking a p ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Crosslinking Of DNA
In genetics, crosslinking of DNA occurs when various exogenous or endogenous agents react with two nucleotides of DNA, forming a covalent linkage between them. This crosslink can occur within the same strand (intrastrand) or between opposite strands of double-stranded DNA (interstrand). These adducts interfere with cellular metabolism, such as DNA replication and Transcription (biology), transcription, triggering cell death. These crosslinks can, however, be DNA repair, repaired through excision or recombination pathways. DNA crosslinking also has useful merit in chemotherapy and targeting cancerous cells for apoptosis, as well as in understanding how proteins interact with DNA. Crosslinking agents Many characterized crosslinking agents have two independently reactive groups within the same molecule, each of which is able to bind with a nucleotide residue of DNA. These agents are separated based upon their source of origin and labeled either as exogenous or endogenous. Exogenous c ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Cell (biology)
The cell is the basic structural and functional unit of all life, forms of life. Every cell consists of cytoplasm enclosed within a Cell membrane, membrane; many cells contain organelles, each with a specific function. The term comes from the Latin word meaning 'small room'. Most cells are only visible under a light microscope, microscope. Cells Abiogenesis, emerged on Earth about 4 billion years ago. All cells are capable of Self-replication, replication, protein synthesis, and cell motility, motility. Cells are broadly categorized into two types: eukaryotic cells, which possess a Cell nucleus, nucleus, and prokaryotic, prokaryotic cells, which lack a nucleus but have a nucleoid region. Prokaryotes are single-celled organisms such as bacteria, whereas eukaryotes can be either single-celled, such as amoebae, or multicellular organism, multicellular, such as some algae, plants, animals, and fungi. Eukaryotic cells contain organelles including Mitochondrion, mitochondria, which ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
ChIP
Chip may refer to: Food * Chip (snack), thinly sliced and deep-fried gastro item ** Potato chips (US) or crisp (UK) * Chips (fried potato strips) (UK) or french fries (US) (common as a takeout side) * Game chips, thin chip/French fries * Chocolate chip Sports and gaming * Chip (association football), a type of football shot * Chip (golf), a type of golf shot * Casino token, often referred to as a chip * Chip, one of the mascots of the University of Colorado at Boulder * Chip, the mascot of Rhode Island FC People and fictional characters * Chip (name), a list of people and fictional characters with either the given name or nickname * Chip (rapper), English hip hop recording artist born Jahmaal Noel Fyffe in 1990 * King Chip (born 1986), stage name of American hip hop rapper Charles Jawanzaa Worth, formerly known as Chip tha Ripper * Chip Douglas, American songwriter, musician, and record producer Douglas Hatlelid (born 1942) * Chip Taylor, stage name of American songwriter Ja ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatin Immunoprecipitation Sequencing
Chromatin is a complex of DNA and protein found in eukaryote, eukaryotic cells. The primary function is to package long DNA molecules into more compact, denser structures. This prevents the strands from becoming tangled and also plays important roles in reinforcing the DNA during cell division, preventing DNA repair#DNA damage, DNA damage, and regulating gene expression and DNA replication. During mitosis and meiosis, chromatin facilitates proper segregation of the chromosomes in anaphase; the characteristic shapes of chromosomes visible during this stage are the result of DNA being coiled into highly condensed chromatin. The primary protein components of chromatin are histones. An octamer of two sets of four histone cores (Histone H2A, Histone H2B, Histone H3, and Histone H4) bind to DNA and function as "anchors" around which the strands are wound.Maeshima, K., Ide, S., & Babokhov, M. (2019). Dynamic chromatin organization without the 30 nm fiber. ''Current opinion in cell biolog ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
FAIRE-Seq
FAIRE-Seq (Formaldehyde-Assisted Isolation of Regulatory Elements) is a method in molecular biology used for determining the sequences of DNA regions in the genome associated with regulatory activity. The technique was developed in the laboratory of Jason D. Lieb at the University of North Carolina, Chapel Hill. In contrast to DNase-Seq, the FAIRE-Seq protocol doesn't require the permeabilization of cells or isolation of nuclei, and can analyse any cell type. In a study of seven diverse human cell types, DNase-seq and FAIRE-seq produced strong cross-validation, with each cell type having 1-2% of the human genome as open chromatin. Workflow The protocol is based on the fact that the formaldehyde cross-linking is more efficient in nucleosome-bound DNA than it is in nucleosome-depleted regions of the genome. This method then segregates the non cross-linked DNA that is usually found in open chromatin, which is then sequenced. The protocol consists of cross linking, phenol extraction an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |