|
CHAP Domain
In molecular biology, the CHAP domain is a region between 110 and 140 amino acids that is found in proteins from bacteria, bacteriophages, archaea and eukaryotes of the family Trypanosomidae. The domain is named after the acronym cysteine, histidine-dependent amidohydrolases/peptidases. Many of these proteins are uncharacterised, but it has been proposed that they may function mainly in peptidoglycan hydrolysis. The CHAP domain is found in a wide range of protein architectures; it is commonly associated with bacterial type SH3 domains and with several families of amidase domains. It has been suggested that CHAP domain containing proteins utilise a catalytic cysteine residue in a nucleophilic-attack mechanism. The CHAP domain contains two invariant residues, a cysteine and a histidine. These residues form part of the putative active site of CHAP domain containing proteins. Secondary structure predictions show that the CHAP domain belongs to the alpha + beta structural class, with ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Region
In geography, regions, otherwise referred to as zones, lands or territories, are areas that are broadly divided by physical characteristics (physical geography), human impact characteristics (human geography), and the interaction of humanity and the environment (environmental geography). Geographic regions and sub-regions are mostly described by their imprecisely defined, and sometimes transitory boundaries, except in human geography, where Jurisdiction (area), jurisdiction areas such as national borders are defined in law. Apart from the Earth, global continental regions, there are also hydrosphere, hydrospheric and atmosphere, atmospheric regions that cover the oceans, and discrete climates above the land mass, land and water mass, water masses of the planet. The land and water global regions are divided into subregions geographically bounded by large geological features that influence large-scale ecologies, such as plains and features. As a way of describing spatial areas, the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Active Site
In biology and biochemistry, the active site is the region of an enzyme where substrate molecules bind and undergo a chemical reaction. The active site consists of amino acid residues that form temporary bonds with the substrate ( binding site) and residues that catalyse a reaction of that substrate (catalytic site). Although the active site occupies only ~10–20% of the volume of an enzyme, it is the most important part as it directly catalyzes the chemical reaction. It usually consists of three to four amino acids, while other amino acids within the protein are required to maintain the tertiary structure of the enzymes. Each active site is evolved to be optimised to bind a particular substrate and catalyse a particular reaction, resulting in high specificity. This specificity is determined by the arrangement of amino acids within the active site and the structure of the substrates. Sometimes enzymes also need to bind with some cofactors to fulfil their function. The a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Peptidase
A protease (also called a peptidase, proteinase, or proteolytic enzyme) is an enzyme that catalyzes (increases reaction rate or "speeds up") proteolysis, breaking down proteins into smaller polypeptides or single amino acids, and spurring the formation of new protein products. They do this by cleaving the peptide bonds within proteins by hydrolysis, a reaction where water breaks bonds. Proteases are involved in many biological functions, including digestion of ingested proteins, protein catabolism (breakdown of old proteins), and cell signaling. In the absence of functional accelerants, proteolysis would be very slow, taking hundreds of years. Proteases can be found in all forms of life and viruses. They have independently evolved multiple times, and different classes of protease can perform the same reaction by completely different catalytic mechanisms. Hierarchy of proteases Based on catalytic residue Proteases can be classified into seven broad groups: * Serine pr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MEROPS
MEROPS is an online database for peptidases (also known as proteases, proteinases and proteolytic enzymes) and their inhibitors. The classification scheme for peptidases was published by Rawlings & Barrett in 1993, and that for protein inhibitors by Rawlings ''et al.'' in 2004.Rawlings, N.D., Tolle, D.P. & Barrett, A.J. (2004) "Evolutionary families of peptidase inhibitors." ''Biochem J'' 378, 705-716. The most recent version, MEROPS 12.3, was released in September 2020. Overview The classification is based on similarities at the tertiary and primary structural levels. Comparisons are restricted to that part of the sequence directly involved in the reaction, which in the case of a peptidase must include the active site, and for a protein inhibitor the reactive site. The classification is hierarchical: sequences are assembled into families, and families are assembled into clans. Each peptidase, family, and clan has a unique identifier. Classification Family The families of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chloride
The chloride ion is the anion (negatively charged ion) Cl−. It is formed when the element chlorine (a halogen) gains an electron or when a compound such as hydrogen chloride is dissolved in water or other polar solvents. Chloride salts such as sodium chloride are often very soluble in water.Green, John, and Sadru Damji. "Chapter 3." ''Chemistry''. Camberwell, Vic.: IBID, 2001. Print. It is an essential electrolyte located in all body fluids responsible for maintaining acid/base balance, transmitting nerve impulses and regulating liquid flow in and out of cells. Less frequently, the word ''chloride'' may also form part of the "common" name of chemical compounds in which one or more chlorine atoms are covalently bonded. For example, methyl chloride, with the standard name chloromethane (see IUPAC books) is an organic compound with a covalent C−Cl bond in which the chlorine is not an anion. Electronic properties A chloride ion (diameter 167 pm) is much lar ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
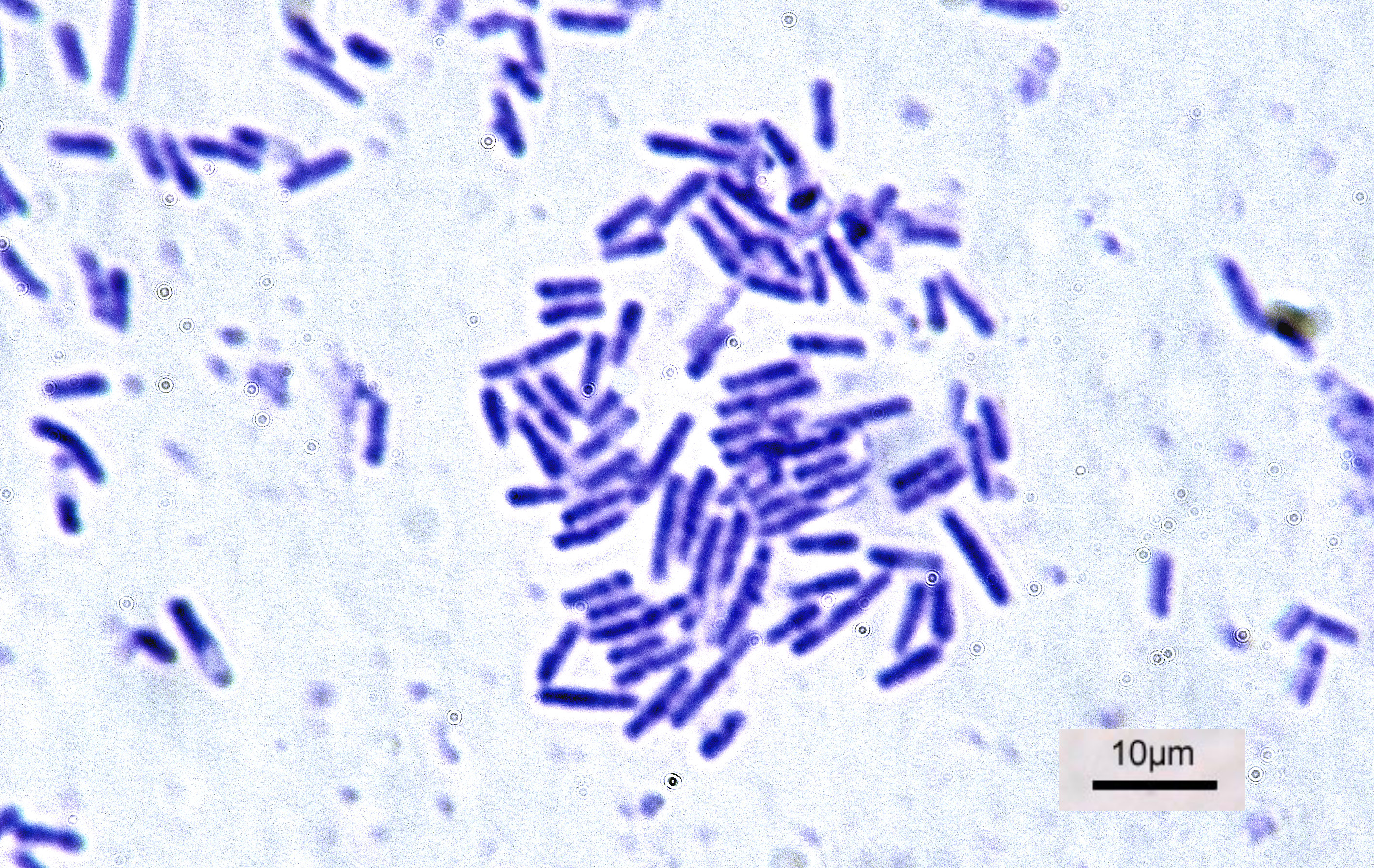
Bacteria
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit soil, water, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria are vital in many stages of the nutrient cycle by recycling nutrients such as the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition of dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. Bacteria also live in symbiotic and parasitic re ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Bacterial
Bacteria (; singular: bacterium) are ubiquitous, mostly free-living organisms often consisting of one biological cell. They constitute a large domain of prokaryotic microorganisms. Typically a few micrometres in length, bacteria were among the first life forms to appear on Earth, and are present in most of its habitats. Bacteria inhabit soil, water, acidic hot springs, radioactive waste, and the deep biosphere of Earth's crust. Bacteria are vital in many stages of the nutrient cycle by recycling nutrients such as the fixation of nitrogen from the atmosphere. The nutrient cycle includes the decomposition of dead bodies; bacteria are responsible for the putrefaction stage in this process. In the biological communities surrounding hydrothermal vents and cold seeps, extremophile bacteria provide the nutrients needed to sustain life by converting dissolved compounds, such as hydrogen sulphide and methane, to energy. Bacteria also live in symbiotic and parasitic relationships ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta Strand
The beta sheet, (β-sheet) (also β-pleated sheet) is a common motif of the regular protein secondary structure. Beta sheets consist of beta strands (β-strands) connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. A β-strand is a stretch of polypeptide chain typically 3 to 10 amino acids long with backbone in an extended conformation. The supramolecular association of β-sheets has been implicated in the formation of the fibrils and protein aggregates observed in amyloidosis, notably Alzheimer's disease. History The first β-sheet structure was proposed by William Astbury in the 1930s. He proposed the idea of hydrogen bonding between the peptide bonds of parallel or antiparallel extended β-strands. However, Astbury did not have the necessary data on the bond geometry of the amino acids in order to build accurate models, especially since he did not then know that the peptide bond was planar. A refined version w ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
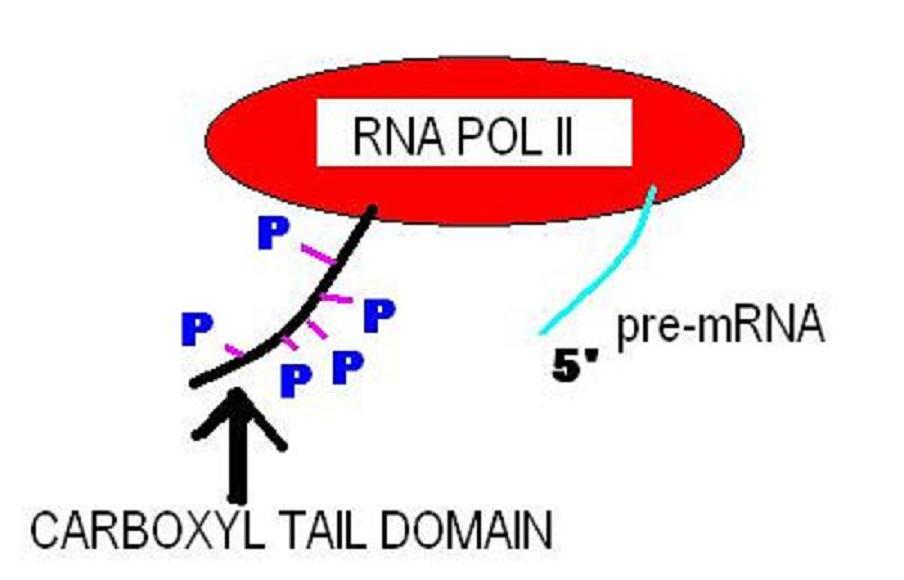
C-terminal
The C-terminus (also known as the carboxyl-terminus, carboxy-terminus, C-terminal tail, C-terminal end, or COOH-terminus) is the end of an amino acid chain (protein or polypeptide), terminated by a free carboxyl group (-COOH). When the protein is translated from messenger RNA, it is created from N-terminus to C-terminus. The convention for writing peptide sequences is to put the C-terminal end on the right and write the sequence from N- to C-terminus. Chemistry Each amino acid has a carboxyl group and an amine group. Amino acids link to one another to form a chain by a dehydration reaction which joins the amine group of one amino acid to the carboxyl group of the next. Thus polypeptide chains have an end with an unbound carboxyl group, the C-terminus, and an end with an unbound amine group, the N-terminus. Proteins are naturally synthesized starting from the N-terminus and ending at the C-terminus. Function C-terminal retention signals While the N-terminus of a protein often con ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Alpha Helix
The alpha helix (α-helix) is a common motif in the secondary structure of proteins and is a right hand-helix conformation in which every backbone N−H group hydrogen bonds to the backbone C=O group of the amino acid located four residues earlier along the protein sequence. The alpha helix is also called a classic Pauling–Corey–Branson α-helix. The name 3.613-helix is also used for this type of helix, denoting the average number of residues per helical turn, with 13 atoms being involved in the ring formed by the hydrogen bond. Among types of local structure in proteins, the α-helix is the most extreme and the most predictable from sequence, as well as the most prevalent. Discovery In the early 1930s, William Astbury showed that there were drastic changes in the X-ray fiber diffraction of moist wool or hair fibers upon significant stretching. The data suggested that the unstretched fibers had a coiled molecular structure with a characteristic repeat of ≈. Astbu ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
N-terminus
The N-terminus (also known as the amino-terminus, NH2-terminus, N-terminal end or amine-terminus) is the start of a protein or polypeptide, referring to the free amine group (-NH2) located at the end of a polypeptide. Within a peptide, the amine group is bonded to the carboxylic group of another amino acid, making it a chain. That leaves a free carboxylic group at one end of the peptide, called the C-terminus, and a free amine group on the other end called the N-terminus. By convention, peptide sequences are written N-terminus to C-terminus, left to right (in LTR writing systems). This correlates the translation direction to the text direction, because when a protein is translated from messenger RNA, it is created from the N-terminus to the C-terminus, as amino acids are added to the carboxyl end of the protein. Chemistry Each amino acid has an amine group and a carboxylic group. Amino acids link to one another by peptide bonds which form through a dehydration reaction ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the three dimensional form of ''local segments'' of proteins. The two most common secondary structural elements are alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein folds into its three dimensional tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the amino hydrogen and carboxyl oxygen atoms in the peptide backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik Linderstrøm-Lang at Stanford in 1952. Other types of biopolymers such as nucleic acids also possess characteristic secondary structures. Types The most common secondary st ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |