|
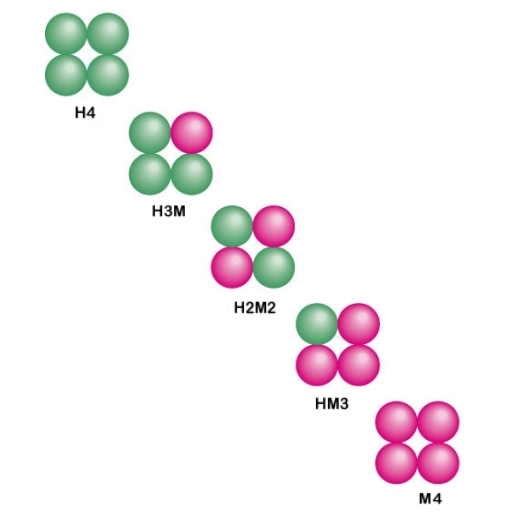
Subfunctionalization
Subfunctionalization was proposed by Stoltzfus (1999) and Force et al. (1999) as one of the possible outcomes of functional divergence that occurs after a gene duplication event, in which pairs of genes that originate from duplication, or paralogs, take on separate functions. Subfunctionalization is a neutral mutation process of constructive neutral evolution; meaning that no new adaptations are formed. During the process of gene duplication paralogs simply undergo a division of labor by retaining different parts (subfunctions) of their original ancestral function. This partitioning event occurs because of segmental gene silencing leading to the formation of paralogs that are no longer duplicates, because each gene only retains a single function. It is important to note that the ancestral gene was capable of performing both functions and the descendant duplicate genes can now only perform one of the original ancestral functions. Alternative Hypothesis Subfunctionalization after gen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Subfunctionalization
Subfunctionalization was proposed by Stoltzfus (1999) and Force et al. (1999) as one of the possible outcomes of functional divergence that occurs after a gene duplication event, in which pairs of genes that originate from duplication, or paralogs, take on separate functions. Subfunctionalization is a neutral mutation process of constructive neutral evolution; meaning that no new adaptations are formed. During the process of gene duplication paralogs simply undergo a division of labor by retaining different parts (subfunctions) of their original ancestral function. This partitioning event occurs because of segmental gene silencing leading to the formation of paralogs that are no longer duplicates, because each gene only retains a single function. It is important to note that the ancestral gene was capable of performing both functions and the descendant duplicate genes can now only perform one of the original ancestral functions. Alternative Hypothesis Subfunctionalization after gen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Gene Duplication
Gene duplication (or chromosomal duplication or gene amplification) is a major mechanism through which new genetic material is generated during molecular evolution. It can be defined as any duplication of a region of DNA that contains a gene. Gene duplications can arise as products of several types of errors in DNA replication and repair machinery as well as through fortuitous capture by selfish genetic elements. Common sources of gene duplications include ectopic recombination, retrotransposition event, aneuploidy, polyploidy, and replication slippage. Mechanisms of duplication Ectopic recombination Duplications arise from an event termed unequal crossing-over that occurs during meiosis between misaligned homologous chromosomes. The chance of it happening is a function of the degree of sharing of repetitive elements between two chromosomes. The products of this recombination are a duplication at the site of the exchange and a reciprocal deletion. Ectopic recombination is ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Constructive Neutral Evolution
Constructive neutral evolution (CNE) is a theory that seeks to explain how complex systems can evolve through neutral transitions and spread through a population by chance fixation (genetic drift). Constructive neutral evolution is a competitor for both adaptationist explanations for the emergence of complex traits and hypotheses positing that a complex trait emerged as a response to a deleterious development in an organism. Constructive neutral evolution often leads to irreversible or "irremediable" complexity and produces systems which, instead of being finely adapted for performing a task, represent an excess complexity that has been described with terms such as "runaway bureaucracy" or even a "Rube Goldberg machine". The groundworks for the concept of CNE were laid by two papers in the 1990s, although first explicitly proposed by Arlin Stoltzfus in 1999. The first proposals for the role CNE was in the evolutionary origins of complex macromolecular machines such as the spliceosom ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Neofunctionalization
Neofunctionalization, one of the possible outcomes of functional divergence, occurs when one gene copy, or paralog, takes on a totally new function after a gene duplication event. Neofunctionalization is an adaptive mutation process; meaning one of the gene copies must mutate to develop a function that was not present in the ancestral gene. In other words, one of the duplicates retains its original function, while the other accumulates molecular changes such that, in time, it can perform a different task. The process The process of Neofunctionalization begins with a gene duplication event, which is thought to occur as a defense mechanism against the accumulation of deleterious mutations. Following the gene duplication event there are two identical copies of the ancestral gene performing exactly the same function. This redundancy allows one the copies to take on a new function. In the event that the new function is advantageous, natural selection positively selects for it and the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Constructive Neutral Evolution
Constructive neutral evolution (CNE) is a theory that seeks to explain how complex systems can evolve through neutral transitions and spread through a population by chance fixation (genetic drift). Constructive neutral evolution is a competitor for both adaptationist explanations for the emergence of complex traits and hypotheses positing that a complex trait emerged as a response to a deleterious development in an organism. Constructive neutral evolution often leads to irreversible or "irremediable" complexity and produces systems which, instead of being finely adapted for performing a task, represent an excess complexity that has been described with terms such as "runaway bureaucracy" or even a "Rube Goldberg machine". The groundworks for the concept of CNE were laid by two papers in the 1990s, although first explicitly proposed by Arlin Stoltzfus in 1999. The first proposals for the role CNE was in the evolutionary origins of complex macromolecular machines such as the spliceosom ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Functional Divergence
Functional divergence is the process by which genes, after gene duplication, shift in function from an ancestral function. Functional divergence can result in either subfunctionalization, where a paralog specializes one of several ancestral functions, or neofunctionalization, where a totally new functional capability evolves. It is thought that this process of gene duplication and functional divergence is a major originator of molecular novelty and has produced the many large protein families that exist today. Functional divergence is just one possible outcome of gene duplication events. Other fates include nonfunctionalization where one of the paralogs acquires deleterious mutations and becomes a pseudogene and superfunctionalization (reinforcement), where both paralogs maintain original function. While gene, chromosome, or whole genome duplication events are considered the canonical sources of functional divergence of paralogs, orthologs (genes descended from speciation events) ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Paralogs
Sequence homology is the biological homology between DNA, RNA, or protein sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry because of three phenomena: either a speciation event (orthologs), or a duplication event (paralogs), or else a horizontal (or lateral) gene transfer event (xenologs). Homology among DNA, RNA, or proteins is typically inferred from their nucleotide or amino acid sequence similarity. Significant similarity is strong evidence that two sequences are related by evolutionary changes from a common ancestral sequence. Alignments of multiple sequences are used to indicate which regions of each sequence are homologous. Identity, similarity, and conservation The term "percent homology" is often used to mean "sequence similarity”, that is the percentage of identical residues (''percent identity''), or the percentage of residues conserved with similar physicochemical properties ('' ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Neutral Mutation
Neutral mutations are changes in DNA sequence that are neither beneficial nor detrimental to the ability of an organism to survive and reproduce. In population genetics, mutations in which natural selection does not affect the spread of the mutation in a species are termed neutral mutations. Neutral mutations that are inheritable and not linked to any genes under selection will be lost or will replace all other alleles of the gene. That loss or fixation of the gene proceeds based on random sampling known as genetic drift. A neutral mutation that is in linkage disequilibrium with other alleles that are under selection may proceed to loss or fixation via genetic hitchhiking and/or background selection. While many mutations in a genome may decrease an organism’s ability to survive and reproduce, also known as fitness, those mutations are selected against and are not passed on to future generations. The most commonly-observed mutations that are detectable as variation in the geneti ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Parsimonious
Occam's razor, Ockham's razor, or Ocham's razor ( la, novacula Occami), also known as the principle of parsimony or the law of parsimony ( la, lex parsimoniae), is the problem-solving principle that "entities should not be multiplied beyond necessity". It is generally understood in the sense that with competing theories or explanations, the simpler one, for example a model with fewer parameters, is to be preferred. The idea is frequently attributed to English Franciscan friar William of Ockham (), a scholastic philosopher and theologian, although he never used these exact words. This philosophical razor advocates that when presented with competing hypotheses about the same prediction, one should select the solution with the fewest assumptions, and that this is not meant to be a way of choosing between hypotheses that make different predictions. Similarly, in science, Occam's razor is used as an abductive heuristic in the development of theoretical models rather than as a rigorou ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Isozymes
In biochemistry, isozymes (also known as isoenzymes or more generally as multiple forms of enzymes) are enzymes that differ in amino acid sequence but catalyze the same chemical reaction. Isozymes usually have different kinetic parameters (e.g. different ''K''M values), or are regulated differently. They permit the fine-tuning of metabolism to meet the particular needs of a given tissue or developmental stage. In many cases, isozymes are encoded by homologous genes that have diverged over time. Strictly speaking, enzymes with different amino acid sequences that catalyse the same reaction are isozymes if encoded by different genes, or allozymes if encoded by different alleles of the same gene; the two terms are often used interchangeably. Introduction Isozymes were first described by R. L. Hunter and Clement Markert (1957) who defined them as ''different variants of the same enzyme having identical functions and present in the same individual''. This definition encompasses (1) ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hemoglobin
Hemoglobin (haemoglobin BrE) (from the Greek word αἷμα, ''haîma'' 'blood' + Latin ''globus'' 'ball, sphere' + ''-in'') (), abbreviated Hb or Hgb, is the iron-containing oxygen-transport metalloprotein present in red blood cells (erythrocytes) of almost all vertebrates (the exception being the fish family Channichthyidae) as well as the tissues of some invertebrates. Hemoglobin in blood carries oxygen from the respiratory organs (''e.g.'' lungs or gills) to the rest of the body (''i.e.'' tissues). There it releases the oxygen to permit aerobic respiration to provide energy to power functions of an organism in the process called metabolism. A healthy individual human has 12to 20grams of hemoglobin in every 100mL of blood. In mammals, the chromoprotein makes up about 96% of the red blood cells' dry content (by weight), and around 35% of the total content (including water). Hemoglobin has an oxygen-binding capacity of 1.34mL O2 per gram, which increases the total blood oxygen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genomics
Genomics is an interdisciplinary field of biology focusing on the structure, function, evolution, mapping, and editing of genomes. A genome is an organism's complete set of DNA, including all of its genes as well as its hierarchical, three-dimensional structural configuration. In contrast to genetics, which refers to the study of ''individual'' genes and their roles in inheritance, genomics aims at the collective characterization and quantification of ''all'' of an organism's genes, their interrelations and influence on the organism. Genes may direct the production of proteins with the assistance of enzymes and messenger molecules. In turn, proteins make up body structures such as organs and tissues as well as control chemical reactions and carry signals between cells. Genomics also involves the sequencing and analysis of genomes through uses of high throughput DNA sequencing and bioinformatics to assemble and analyze the function and structure of entire genomes. Advances in ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |