|
Shuttle Vector
A shuttle vector is a vector (usually a plasmid) constructed so that it can propagate in two different host species. Therefore, DNA inserted into a shuttle vector can be tested or manipulated in two different cell types. The main advantage of these vectors is they can be manipulated in ''E. coli'', then used in a system which is more difficult or slower to use (e.g. yeast). Shuttle vectors include plasmids that can propagate in eukaryotes and prokaryotes (e.g. both ''Saccharomyces cerevisiae'' and ''Escherichia coli'') or in different species of bacteria (e.g. both ''E. coli'' and ''Rhodococcus erythropolis''). There are also adenovirus shuttle vectors, which can propagate in ''E. coli'' and mammals. Shuttle vectors are frequently used to quickly make multiple copies of the gene in '' E. coli'' (amplification). They can also be used for ''in vitro'' experiments and modifications (e.g. mutagenesis, PCR). One of the most common types of shuttle vectors is the yeast shuttle ve ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Vector (molecular Biology)
In molecular cloning, a vector is any particle (e.g., plasmids, cosmids, Lambda phages) used as a vehicle to artificially carry a foreign nucleic sequence – usually DNA – into another cell, where it can be replicated and/or expressed. A vector containing foreign DNA is termed recombinant DNA. The four major types of vectors are plasmids, viral vectors, cosmids, and artificial chromosomes. Of these, the most commonly used vectors are plasmids. Common to all engineered vectors have an origin of replication, a multicloning site, and a selectable marker. The vector itself generally carries a DNA sequence that consists of an insert (in this case the transgene) and a larger sequence that serves as the "backbone" of the vector. The purpose of a vector which transfers genetic information to another cell is typically to isolate, multiply, or express the insert in the target cell. All vectors may be used for cloning and are therefore cloning vectors, but there are also vect ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Yeast
Yeasts are eukaryotic, single-celled microorganisms classified as members of the fungus kingdom. The first yeast originated hundreds of millions of years ago, and at least 1,500 species are currently recognized. They are estimated to constitute 1% of all described fungal species. Yeasts are unicellular organisms that evolved from multicellular ancestors, with some species having the ability to develop multicellular characteristics by forming strings of connected budding cells known as pseudohyphae or false hyphae. Yeast sizes vary greatly, depending on species and environment, typically measuring 3–4 µm in diameter, although some yeasts can grow to 40 µm in size. Most yeasts reproduce asexually by mitosis, and many do so by the asymmetric division process known as budding. With their single-celled growth habit, yeasts can be contrasted with molds, which grow hyphae. Fungal species that can take both forms (depending on temperature or other conditions) a ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Uracil
Uracil () (symbol U or Ura) is one of the four nucleobases in the nucleic acid RNA. The others are adenine (A), cytosine (C), and guanine (G). In RNA, uracil binds to adenine via two hydrogen bonds. In DNA, the uracil nucleobase is replaced by thymine (T). Uracil is a demethylated form of thymine. Uracil is a common and naturally occurring pyrimidine derivative. The name "uracil" was coined in 1885 by the German chemist Robert Behrend, who was attempting to synthesize derivatives of uric acid. Originally discovered in 1900 by Alberto Ascoli, it was isolated by hydrolysis of yeast nuclein; it was also found in bovine thymus and spleen, herring sperm, and wheat germ. It is a planar, unsaturated compound that has the ability to absorb light. Based on 12C/13C isotopic ratios of organic compounds found in the Murchison meteorite, it is believed that uracil, xanthine, and related molecules can also be formed extraterrestrially. Data from the Cassini mission, orbiting in th ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
URA3
URA3 is a gene on chromosome V in ''Saccharomyces cerevisiae'' (yeast). Its systematic name is YEL021W. URA3 is often used in yeast research as a "marker gene", that is, a gene to label chromosomes or plasmids. URA3 encodes Orotidine 5'-phosphate decarboxylase (ODCase), which is an enzyme that catalyzes one reaction in the synthesis of pyrimidine ribonucleotides (a component of RNA). Use in yeast research Loss of ODCase activity leads to a lack of cell growth unless uracil or uridine is added to the media. The presence of the URA3 gene in yeast restores ODCase activity, facilitating growth on media not supplemented with uracil or uridine, thereby allowing selection for yeast carrying the gene. In contrast, if 5-FOA (5-Fluoroorotic acid) is added to the media, the active ODCase will convert 5-FOA into the toxic compound (a suicide inhibitor) 5-fluorouracil causing cell death, which allows for selection against yeast carrying the gene. Since URA3 allows for both positive and ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Selectable Marker
A selectable marker is a gene introduced into a cell, especially a bacterium or to cells in culture, that confers a trait suitable for artificial selection. They are a type of reporter gene used in laboratory microbiology, molecular biology, and genetic engineering to indicate the success of a transfection or other procedure meant to introduce foreign DNA into a cell. Selectable markers are often antibiotic resistance genes (''An antibiotic resistance marker is a gene that produces a protein that provides cells expressing this protein with resistance to an antibiotic.''). Bacteria that have been subjected to a procedure to introduce foreign DNA are grown on a medium containing an antibiotic, and those bacterial colonies that can grow have successfully taken up and expressed the introduced genetic material. Normally the genes encoding resistance to antibiotics such as ampicillin, chloramphenicol, tetracycline or kanamycin, etc., are considered useful selectable markers for ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Centromere
The centromere links a pair of sister chromatids together during cell division. This constricted region of chromosome connects the sister chromatids, creating a short arm (p) and a long arm (q) on the chromatids. During mitosis, spindle fibers attach to the centromere via the kinetochore. The physical role of the centromere is to act as the site of assembly of the kinetochores – a highly complex multiprotein structure that is responsible for the actual events of chromosome segregation – i.e. binding microtubules and signaling to the cell cycle machinery when all chromosomes have adopted correct attachments to the spindle, so that it is safe for cell division to proceed to completion and for cells to enter anaphase. There are, broadly speaking, two types of centromeres. "Point centromeres" bind to specific proteins that recognize particular DNA sequences with high efficiency. Any piece of DNA with the point centromere DNA sequence on it will typically form a centromere ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Autonomously Replicating Sequence
An autonomously replicating sequence (ARS) contains the origin of replication in the yeast genome. It contains four regions (A, B1, B2, and B3), named in order of their effect on plasmid stability. The A-Domain is highly conserved, any mutation abolishes origin function. Mutations on B1, B2, and B3 will diminish, but not prevent functioning of the origin. Element A is highly conserved, consisting of the consensus sequence: (where ''Y'' is either pyrimidine and ''R'' is either purine). When this element is mutated, the ARS loses all activity. As seen above the ARS are considerably A-T rich which makes it easy for replicative proteins to disrupt the H-bonding in that area. ORC protein complex (origin recognition complex) is bound at the ARS throughout the cell cycle, allowing replicative proteins access to the ARS. Mutational analysis for the yeast ARS elements have shown that any mutation in the B1, B2 and B3 regions result in a reduction of function of the ARS element. A muta ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta Lactamase
Beta-lactamases, (β-lactamases) are enzymes () produced by bacteria that provide multi-resistance to beta-lactam antibiotics such as penicillins, cephalosporins, cephamycins, monobactams and carbapenems (ertapenem), although carbapenems are relatively resistant to beta-lactamase. Beta-lactamase provides antibiotic resistance by breaking the antibiotics' structure. These antibiotics all have a common element in their molecular structure: a four-atom ring known as a beta-lactam (β-lactam) ring. Through hydrolysis, the enzyme lactamase breaks the β-lactam ring open, deactivating the molecule's antibacterial properties. Beta-lactam antibiotics are typically used to target a broad spectrum of gram-positive and gram-negative bacteria. Beta-lactamases produced by gram-negative bacteria are usually secreted, especially when antibiotics are present in the environment. Structure The structure of a ''Streptomyces'' serine β-lactamase (SBLs) is given by . The alpha-beta ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Antimicrobial Resistance
Antimicrobial resistance (AMR) occurs when microbes evolve mechanisms that protect them from the effects of antimicrobials. All classes of microbes can evolve resistance. Fungi evolve antifungal resistance. Viruses evolve antiviral resistance. Protozoa evolve antiprotozoal resistance, and bacteria evolve antibiotic resistance. Those bacteria that are considered extensively drug resistant (XDR) or totally drug-resistant (TDR) are sometimes called "superbugs".A.-P. Magiorakos, A. Srinivasan, R. B. Carey, Y. Carmeli, M. E. Falagas, C. G. Giske, S. Harbarth, J. F. Hinndler ''et al''Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria... Clinical Microbiology and Infection, Vol 8, Iss. 3 first published 27 July 2011 ia Wiley Online Library Retrieved 28 August 2020 Although antimicrobial resistance is a naturally-occurring process, it is often the result of improper usage of the drugs and management of the infections. Antibiotic resistance is a major subset of ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Origin Of Replication
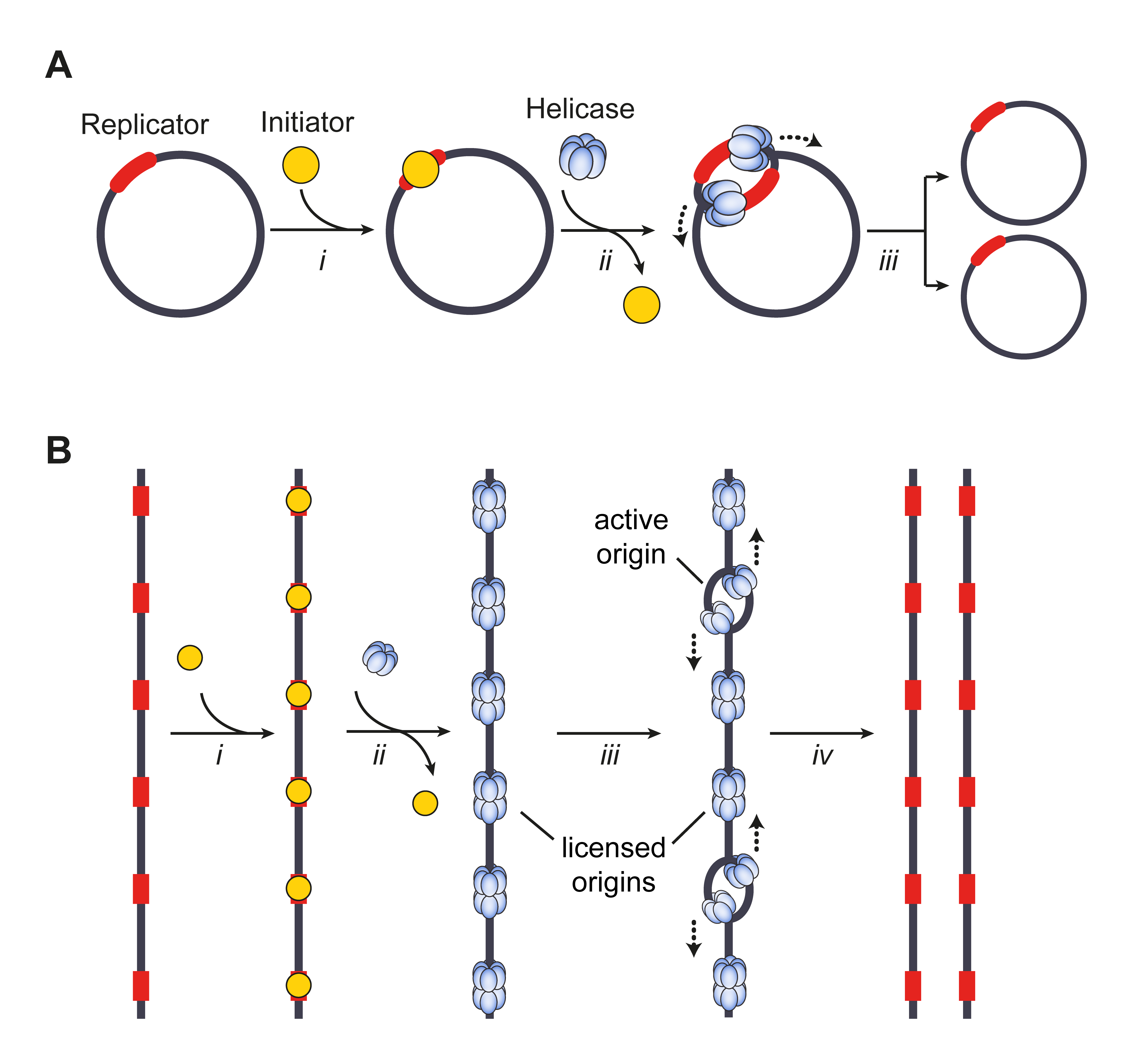
The origin of replication (also called the replication origin) is a particular sequence in a genome at which replication is initiated. Propagation of the genetic material between generations requires timely and accurate duplication of DNA by semiconservative replication prior to cell division to ensure each daughter cell receives the full complement of chromosomes. Material was copied from this source, which is available under Creative Commons Attribution 4.0 International License This can either involve the replication of DNA in living organisms such as prokaryotes and eukaryotes, or that of DNA or RNA in viruses, such as double-stranded RNA viruses. Synthesis of daughter strands starts at discrete sites, termed replication origins, and proceeds in a bidirectional manner until all genomic DNA is replicated. Despite the fundamental nature of these events, organisms have evolved surprisingly divergent strategies that control replication onset. Although the specific replicatio ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Polymerase Chain Reaction
The polymerase chain reaction (PCR) is a method widely used to rapidly make millions to billions of copies (complete or partial) of a specific DNA sample, allowing scientists to take a very small sample of DNA and amplify it (or a part of it) to a large enough amount to study in detail. PCR was invented in 1983 by the American biochemist Kary Mullis at Cetus Corporation; Mullis and biochemist Michael Smith, who had developed other essential ways of manipulating DNA, were jointly awarded the Nobel Prize in Chemistry in 1993. PCR is fundamental to many of the procedures used in genetic testing and research, including analysis of ancient samples of DNA and identification of infectious agents. Using PCR, copies of very small amounts of DNA sequences are exponentially amplified in a series of cycles of temperature changes. PCR is now a common and often indispensable technique used in medical laboratory research for a broad variety of applications including biomedical research ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Plasmid
A plasmid is a small, extrachromosomal DNA molecule within a cell that is physically separated from chromosomal DNA and can replicate independently. They are most commonly found as small circular, double-stranded DNA molecules in bacteria; however, plasmids are sometimes present in archaea and eukaryotic organisms. In nature, plasmids often carry genes that benefit the survival of the organism and confer selective advantage such as antibiotic resistance. While chromosomes are large and contain all the essential genetic information for living under normal conditions, plasmids are usually very small and contain only additional genes that may be useful in certain situations or conditions. Artificial plasmids are widely used as vectors in molecular cloning, serving to drive the replication of recombinant DNA sequences within host organisms. In the laboratory, plasmids may be introduced into a cell via transformation. Synthetic plasmids are available for procurement over the int ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |