|
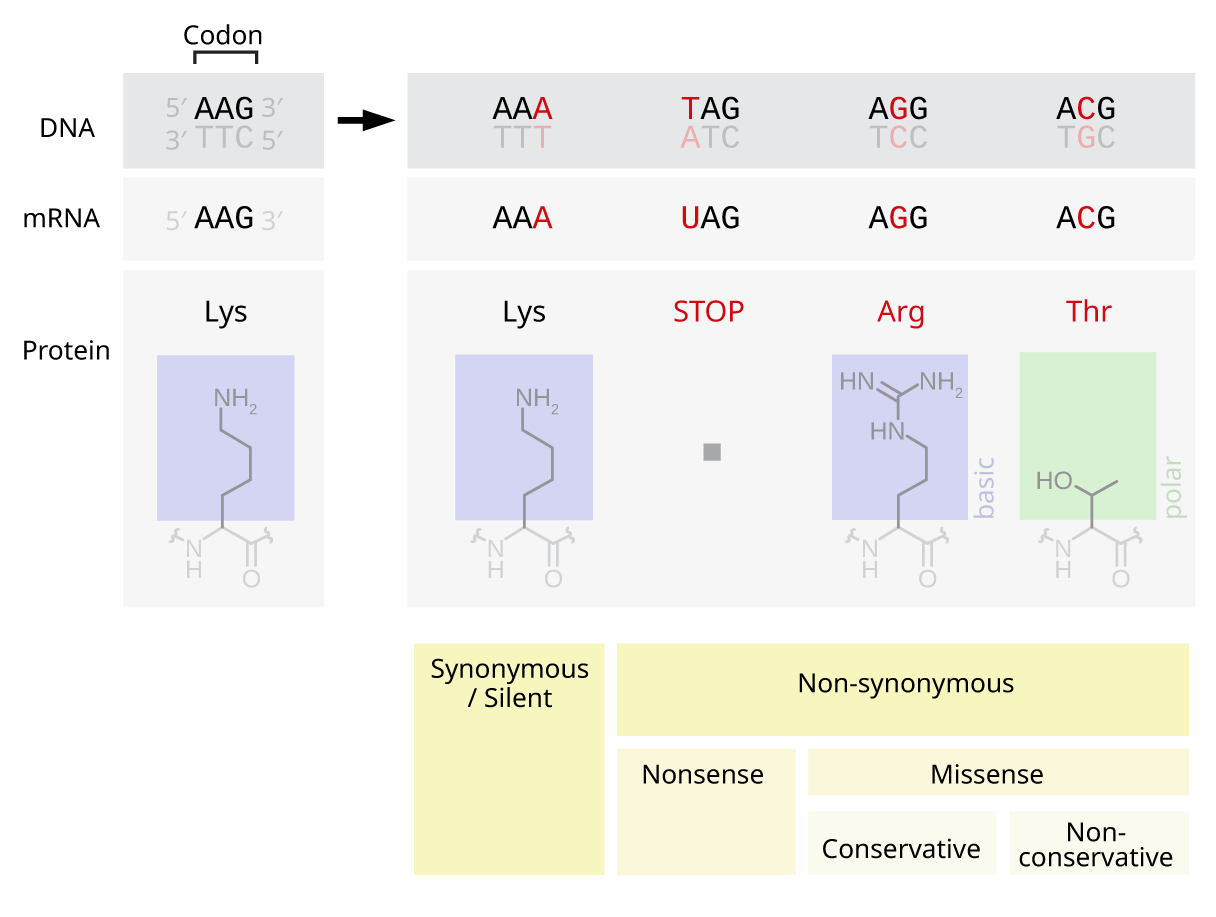
Nonsynonymous Variant
A nonsynonymous substitution is a nucleotide mutation that alters the amino acid sequence of a protein. Nonsynonymous substitutions differ from synonymous substitutions, which do not alter amino acid sequences and are (sometimes) silent mutations. As nonsynonymous substitutions result in a biological change in the organism, they are subject to natural selection. Nonsynonymous substitutions at a certain locus can be compared to the synonymous substitutions at the same locus to obtain the Ka/Ks ratio. This ratio is used to measure the evolutionary rate of gene sequences. If a gene has lower levels of nonsynonymous than synonymous nucleotide substitution, then it can be inferred to be functional because a Ka/Ks ratio < 1 is a hallmark of sequences that are being constrained to code for proteins. Nonsynonymous substitutions are also referred to as replacement mutations. Types There are several common types of nonsynonymous substitutions. ...[...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nucleotide
Nucleotides are organic molecules consisting of a nucleoside and a phosphate. They serve as monomeric units of the nucleic acid polymers – deoxyribonucleic acid (DNA) and ribonucleic acid (RNA), both of which are essential biomolecules within all life-forms on Earth. Nucleotides are obtained in the diet and are also synthesized from common nutrients by the liver. Nucleotides are composed of three subunit molecules: a nucleobase, a five-carbon sugar ( ribose or deoxyribose), and a phosphate group consisting of one to three phosphates. The four nucleobases in DNA are guanine, adenine, cytosine and thymine; in RNA, uracil is used in place of thymine. Nucleotides also play a central role in metabolism at a fundamental, cellular level. They provide chemical energy—in the form of the nucleoside triphosphates, adenosine triphosphate (ATP), guanosine triphosphate (GTP), cytidine triphosphate (CTP) and uridine triphosphate (UTP)—throughout the cell for the ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Nonsense Mutation
In genetics, a nonsense mutation is a point mutation in a sequence of DNA that results in a premature stop codon, or a ''nonsense codon'' in the transcribed mRNA, and in leading to a truncated, incomplete, and usually nonfunctional protein product. The functional effect of a nonsense mutation depends on the location of the stop codon within the coding DNA. For example, the effect of a nonsense mutation depends on the proximity of the nonsense mutation to the original stop codon, and the degree to which functional subdomains of the protein are affected. As nonsense mutations leads to premature termination of polypeptide chains; they are also called chain termination mutations. Missense mutations differ from nonsense mutations since they are point mutations that exhibit a single nucleotide change to cause substitution of a different amino acid. A nonsense mutation also differs from a nonstop mutation, which is a point mutation that removes a stop codon. About 10% of patients facin ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Neutral Theory Of Molecular Evolution
The neutral theory of molecular evolution holds that most evolutionary changes occur at the molecular level, and most of the variation within and between species are due to random genetic drift of mutant alleles that are selectively neutral. The theory applies only for evolution at the molecular level, and is compatible with phenotypic evolution being shaped by natural selection as postulated by Charles Darwin. The neutral theory allows for the possibility that most mutations are deleterious, but holds that because these are rapidly removed by natural selection, they do not make significant contributions to variation within and between species at the molecular level. A neutral mutation is one that does not affect an organism's ability to survive and reproduce. The neutral theory assumes that most mutations that are not deleterious are neutral rather than beneficial. Because only a fraction of gametes are sampled in each generation of a species, the neutral theory suggests that a m ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
McDonald–Kreitman Test
The McDonald–Kreitman test is a statistical test often used by evolutionary and population biologists to detect and measure the amount of adaptive evolution within a species by determining whether adaptive evolution has occurred, and the proportion of substitutions that resulted from positive selection (also known as directional selection). To do this, the McDonald–Kreitman test compares the amount of variation within a species ( polymorphism) to the divergence between species (substitutions) at two types of sites, neutral and nonneutral. A substitution refers to a nucleotide that is fixed within one species, but a different nucleotide is fixed within a second species at the same base pair of homologous DNA sequences.Futuyma, D. J. 2013. Evolution. Sinauer Associates, Inc.: Sunderland. A site is nonneutral if it is either advantageous or deleterious. The two types of sites can be either synonymous or nonsynonymous within a protein-coding region. In a protein-coding sequence of DN ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Molecular Evolution
Molecular evolution is the process of change in the sequence composition of cellular molecules such as DNA, RNA, and proteins across generations. The field of molecular evolution uses principles of evolutionary biology and population genetics to explain patterns in these changes. Major topics in molecular evolution concern the rates and impacts of single nucleotide changes, neutral evolution vs. natural selection, origins of new genes, the genetic nature of complex traits, the genetic basis of speciation, evolution of development, and ways that evolutionary forces influence genomic and phenotypic changes. History The history of molecular evolution starts in the early 20th century with comparative biochemistry, and the use of "fingerprinting" methods such as immune assays, gel electrophoresis and paper chromatography in the 1950s to explore homologous proteins. The field of molecular evolution came into its own in the 1960s and 1970s, following the rise of molecular ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genetic Drift
Genetic drift, also known as allelic drift or the Wright effect, is the change in the frequency of an existing gene variant (allele) in a population due to random chance. Genetic drift may cause gene variants to disappear completely and thereby reduce genetic variation. It can also cause initially rare alleles to become much more frequent and even fixed. When few copies of an allele exist, the effect of genetic drift is more notable, and when many copies exist, the effect is less notable. In the middle of the 20th century, vigorous debates occurred over the relative importance of natural selection versus neutral processes, including genetic drift. Ronald Fisher, who explained natural selection using Mendelian genetics, held the view that genetic drift plays at most a minor role in evolution, and this remained the dominant view for several decades. In 1968, population geneticist Motoo Kimura rekindled the debate with his neutral theory of molecular evolution, which claims that ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Stop Codon
In molecular biology (specifically protein biosynthesis), a stop codon (or termination codon) is a codon (nucleotide triplet within messenger RNA) that signals the termination of the translation process of the current protein. Most codons in messenger RNA correspond to the addition of an amino acid to a growing polypeptide chain, which may ultimately become a protein; stop codons signal the termination of this process by binding release factors, which cause the ribosomal subunits to disassociate, releasing the amino acid chain. While start codons need nearby sequences or initiation factors to start translation, a stop codon alone is sufficient to initiate termination. Properties Standard codons In the standard genetic code, there are three different termination codons: Alternative stop codons There are variations on the standard genetic code, and alternative stop codons have been found in the mitochondrial genomes of vertebrates, '' Scenedesmus obliquus'', and '' Thrau ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Stop Codon
In molecular biology (specifically protein biosynthesis), a stop codon (or termination codon) is a codon (nucleotide triplet within messenger RNA) that signals the termination of the translation process of the current protein. Most codons in messenger RNA correspond to the addition of an amino acid to a growing polypeptide chain, which may ultimately become a protein; stop codons signal the termination of this process by binding release factors, which cause the ribosomal subunits to disassociate, releasing the amino acid chain. While start codons need nearby sequences or initiation factors to start translation, a stop codon alone is sufficient to initiate termination. Properties Standard codons In the standard genetic code, there are three different termination codons: Alternative stop codons There are variations on the standard genetic code, and alternative stop codons have been found in the mitochondrial genomes of vertebrates, '' Scenedesmus obliquus'', and '' Thrau ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Point Mutation
A point mutation is a genetic mutation where a single nucleotide base is changed, inserted or deleted from a DNA or RNA sequence of an organism's genome. Point mutations have a variety of effects on the downstream protein product—consequences that are moderately predictable based upon the specifics of the mutation. These consequences can range from no effect (e.g. synonymous mutations) to deleterious effects (e.g. frameshift mutations), with regard to protein production, composition, and function. Causes Point mutations usually take place during DNA replication. DNA replication occurs when one double-stranded DNA molecule creates two single strands of DNA, each of which is a template for the creation of the complementary strand. A single point mutation can change the whole DNA sequence. Changing one purine or pyrimidine may change the amino acid that the nucleotides code for. Point mutations may arise from spontaneous mutations that occur during DNA replication. The rate o ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mutation
In biology, a mutation is an alteration in the nucleic acid sequence of the genome of an organism, virus, or extrachromosomal DNA. Viral genomes contain either DNA or RNA. Mutations result from errors during DNA replication, DNA or viral replication, mitosis, or meiosis or other types of DNA repair#DNA damage, damage to DNA (such as pyrimidine dimers caused by exposure to ultraviolet radiation), which then may undergo error-prone repair (especially microhomology-mediated end joining), cause an error during other forms of repair, or cause an error during replication (DNA repair#Translesion synthesis, translesion synthesis). Mutations may also result from Insertion (genetics), insertion or Deletion (genetics), deletion of segments of DNA due to mobile genetic elements. Mutations may or may not produce detectable changes in the observable characteristics (phenotype) of an organism. Mutations play a part in both normal and abnormal biological processes including: evolution, cancer ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Missense Mutation
In genetics, a missense mutation is a point mutation in which a single nucleotide change results in a codon that codes for a different amino acid. It is a type of nonsynonymous substitution. Substitution of protein from DNA mutations Missense mutation refers to a change in one amino acid in a protein, arising from a point mutation in a single nucleotide. Missense mutation is a type of nonsynonymous substitution in a DNA sequence. Two other types of nonsynonymous substitution are the nonsense mutations, in which a codon is changed to a premature stop codon that results in truncation of the resulting protein, and the nonstop mutations, in which a stop codon erasement results in a longer, nonfunctional protein. Missense mutations can render the resulting protein nonfunctional, and such mutations are responsible for human diseases such as Epidermolysis bullosa, sickle-cell disease, SOD1 mediated ALS, and a substantial number of cancers. In the most common variant of sickle-cell ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Ka/Ks Ratio
In genetics, the Ka/Ks ratio, also known as ω or ''d''N/''d''S ratio, is used to estimate the balance between neutral mutations, purifying selection and beneficial mutations acting on a set of homologous protein-coding genes. It is calculated as the ratio of the number of nonsynonymous substitutions per non-synonymous site (Ka), in a given period of time, to the number of synonymous substitutions per synonymous site (Ks), in the same period. The latter are assumed to be neutral, so that the ratio indicates the net balance between deleterious and beneficial mutations. Values of Ka/Ks significantly above 1 are unlikely to occur without at least some of the mutations being advantageous. If beneficial mutations are assumed to make little contribution, then Ka/Ks estimates the degree of evolutionary constraint. Context Selection acts on variation in phenotypes, which are often the result of mutations in protein-coding genes. The genetic code is written in DNA sequences as codons, gro ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |