|
DUTPase
In Enzymology, a dUTP diphosphatase () is an enzyme that catalyzes the chemical reaction :dUTP + H2O \rightleftharpoons dUMP + diphosphate Thus, the two substrates of this enzyme are dUTP and H2O, whereas its two products are dUMP and diphosphate. This enzyme belongs to the family of hydrolases, specifically those acting on acid anhydrides in phosphorus-containing anhydrides. The systematic name of this enzyme class is dUTP nucleotidohydrolase. Other names in common use include deoxyuridine-triphosphatase, dUTPase, dUTP pyrophosphatase, desoxyuridine 5'-triphosphate nucleotidohydrolase, and desoxyuridine 5'-triphosphatase. This enzyme participates in pyrimidine metabolism. This enzyme has a dual function: on one hand, it removes dUTP from the deoxynucleotide pool, which reduces the probability of this base being incorporated into DNA by DNA polymerases, while on the other hand, it produces the dTTP precursor dUMP. Lack or inhibition of dUTPase action leads to harmful per ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DUT (gene)
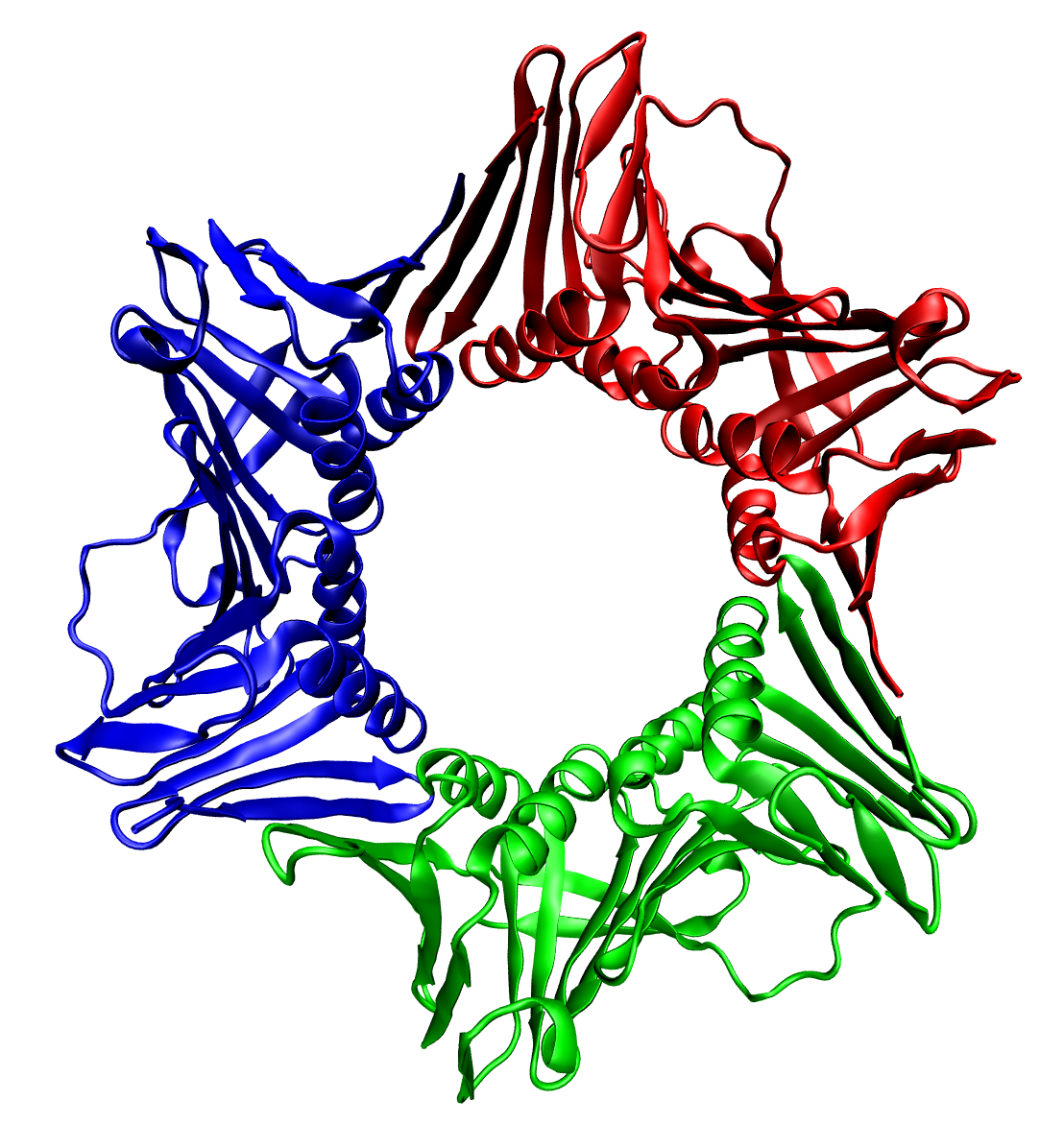
DUTP pyrophosphatase, also known as DUT, is an enzyme which in humans is encoded by the ''DUT'' gene on chromosome 15. This gene encodes an essential enzyme of nucleotide metabolism. The encoded protein forms a ubiquitous, homotrimeric enzyme that hydrolyzes dUTP to dUMP and pyrophosphate. This reaction serves two cellular purposes: providing a precursor (dUMP) for the synthesis of thymine nucleotides needed for DNA replication, and limiting intracellular pools of dUTP. Elevated levels of dUTP lead to increased incorporation of uracil into DNA, which induces extensive excision repair mediated by uracil glycosylase. This repair process, resulting in the removal and reincorporation of dUTP, is self-defeating and leads to DNA fragmentation and cell death. Alternative splicing of this gene leads to different isoforms that localize to either the mitochondrion or nucleus. A related pseudogene is located on chromosome 19. Structure In humans, this gene encodes a homotrimeric enzy ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzymology
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrate (chemistry), substrates, and the enzyme converts the substrates into different molecules known as product (chemistry), products. Almost all metabolism, metabolic processes in the cell (biology), cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme, pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts include Ribozyme, catalytic RNA molecules, also called ribozymes. They are sometimes descr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Tertiary Structure
Protein tertiary structure is the three-dimensional shape of a protein. The tertiary structure will have a single polypeptide chain "backbone" with one or more protein secondary structures, the protein domains. Amino acid side chains and the backbone may interact and bond in a number of ways. The interactions and bonds of side chains within a particular protein determine its tertiary structure. The protein tertiary structure is defined by its atomic coordinates. These coordinates may refer either to a protein domain or to the entire tertiary structure. A number of these structures may bind to each other, forming a quaternary structure. History The science of the tertiary structure of proteins has progressed from one of hypothesis to one of detailed definition. Although Emil Fischer had suggested proteins were made of polypeptide chains and amino acid side chains, it was Dorothy Maud Wrinch who incorporated geometry into the prediction of protein structures. Wrinch demon ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Organism
An organism is any life, living thing that functions as an individual. Such a definition raises more problems than it solves, not least because the concept of an individual is also difficult. Many criteria, few of them widely accepted, have been proposed to define what an organism is. Among the most common is that an organism has autonomous reproduction, Cell growth, growth, and metabolism. This would exclude viruses, despite the fact that they evolution, evolve like organisms. Other problematic cases include colonial organisms; a colony of eusocial insects is organised adaptively, and has Germ-Soma Differentiation, germ-soma specialisation, with some insects reproducing, others not, like cells in an animal's body. The body of a siphonophore, a jelly-like marine animal, is composed of organism-like zooids, but the whole structure looks and functions much like an animal such as a jellyfish, the parts collaborating to provide the functions of the colonial organism. The evolutiona ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Enzymes
An enzyme () is a protein that acts as a biological catalyst by accelerating chemical reactions. The molecules upon which enzymes may act are called substrates, and the enzyme converts the substrates into different molecules known as products. Almost all metabolic processes in the cell need enzyme catalysis in order to occur at rates fast enough to sustain life. Metabolic pathways depend upon enzymes to catalyze individual steps. The study of enzymes is called ''enzymology'' and the field of pseudoenzyme analysis recognizes that during evolution, some enzymes have lost the ability to carry out biological catalysis, which is often reflected in their amino acid sequences and unusual 'pseudocatalytic' properties. Enzymes are known to catalyze more than 5,000 biochemical reaction types. Other biocatalysts include catalytic RNA molecules, also called ribozymes. They are sometimes described as a ''type'' of enzyme rather than being ''like'' an enzyme, but even in the d ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Sequence Homology
Sequence homology is the homology (biology), biological homology between DNA sequence, DNA, RNA sequence, RNA, or Protein primary structure, protein sequences, defined in terms of shared ancestry in the evolutionary history of life. Two segments of DNA can have shared ancestry because of three phenomena: either a speciation event (orthologs), or a Gene duplication, duplication event (paralogs), or else a Horizontal gene transfer, horizontal (or lateral) gene transfer event (xenologs). Homology among DNA, RNA, or proteins is typically inferred from their nucleotide or amino acid sequence similarity. Significant similarity is strong evidence that two sequences are related by evolutionary changes from a common ancestral sequence. Sequence alignment, Alignments of multiple sequences are used to indicate which regions of each sequence are homologous. Identity, similarity, and conservation The term "percent homology" is often used to mean "sequence similarity”, that is the percen ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Escherichia Coli
''Escherichia coli'' ( )Wells, J. C. (2000) Longman Pronunciation Dictionary. Harlow ngland Pearson Education Ltd. is a gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genus '' Escherichia'' that is commonly found in the lower intestine of warm-blooded organisms. Most ''E. coli'' strains are part of the normal microbiota of the gut, where they constitute about 0.1%, along with other facultative anaerobes. These bacteria are mostly harmless or even beneficial to humans. For example, some strains of ''E. coli'' benefit their hosts by producing vitamin K2 or by preventing the colonization of the intestine by harmful pathogenic bacteria. These mutually beneficial relationships between ''E. coli'' and humans are a type of mutualistic biological relationship—where both the humans and the ''E. coli'' are benefitting each other. ''E. coli'' is expelled into the environment within fecal matter. The bacterium grows massi ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Secondary Structure
Protein secondary structure is the local spatial conformation of the polypeptide backbone excluding the side chains. The two most common Protein structure#Secondary structure, secondary structural elements are alpha helix, alpha helices and beta sheets, though beta turns and omega loops occur as well. Secondary structure elements typically spontaneously form as an intermediate before the protein protein folding, folds into its three dimensional protein tertiary structure, tertiary structure. Secondary structure is formally defined by the pattern of hydrogen bonds between the Amine, amino hydrogen and carboxyl oxygen atoms in the peptide backbone chain, backbone. Secondary structure may alternatively be defined based on the regular pattern of backbone Dihedral angle#Dihedral angles of proteins, dihedral angles in a particular region of the Ramachandran plot regardless of whether it has the correct hydrogen bonds. The concept of secondary structure was first introduced by Kaj Ulrik ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Subunit
In structural biology, a protein subunit is a polypeptide chain or single protein molecule that assembles (or "''coassembles''") with others to form a protein complex. Large assemblies of proteins such as viruses often use a small number of types of protein subunits as building blocks. A subunit is often named with a Greek or Roman letter, and the numbers of this type of subunit in a protein is indicated by a subscript. For example, ATP synthase has a type of subunit called α. Three of these are present in the ATP synthase molecule, leading to the designation α3. Larger groups of subunits can also be specified, like α3β3-hexamer and c-ring. Naturally occurring proteins that have a relatively small number of subunits are referred to as oligomeric.Quote: ''Oligomer molecule: A molecule of intermediate relative molecular mass, the structure of which essentially comprises a small plurality of units derived, actually or conceptually, from molecules of lower relative molecula ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Beta Sheet
The beta sheet (β-sheet, also β-pleated sheet) is a common motif of the regular protein secondary structure. Beta sheets consist of beta strands (β-strands) connected laterally by at least two or three backbone hydrogen bonds, forming a generally twisted, pleated sheet. A β-strand is a stretch of polypeptide chain typically 3 to 10 amino acids long with backbone in an extended conformation. The supramolecular association of β-sheets has been implicated in the formation of the fibrils and protein aggregates observed in amyloidosis, Alzheimer's disease and other proteinopathies. History The first β-sheet structure was proposed by William Astbury in the 1930s. He proposed the idea of hydrogen bonding between the peptide bonds of parallel or antiparallel extended β-strands. However, Astbury did not have the necessary data on the bond geometry of the amino acids in order to build accurate models, especially since he did not then know that the peptide bond was planar. ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Protein Folding
Protein folding is the physical process by which a protein, after Protein biosynthesis, synthesis by a ribosome as a linear chain of Amino acid, amino acids, changes from an unstable random coil into a more ordered protein tertiary structure, three-dimensional structure. This structure permits the protein to become biologically functional or active. The folding of many proteins begins even during the translation of the polypeptide chain. The amino acids interact with each other to produce a well-defined three-dimensional structure, known as the protein's native state. This structure is determined by the amino-acid sequence or primary structure. The correct three-dimensional structure is essential to function, although some parts of functional proteins Intrinsically unstructured proteins, may remain unfolded, indicating that protein dynamics are important. Failure to fold into a native structure generally produces inactive proteins, but in some instances, misfolded proteins have ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Trimer (biochemistry)
thumbnail, 400px, Trimeric form of a TNF-α mutant In biochemistry, a protein trimer is a macromolecular complex formed by three, usually non-covalently bound, macromolecules like proteins or nucleic acids. A protein trimer often occurs from the assembly of a protein's quaternary structure. The non-covalent interactions between the hydrophobic and hydrophilic regions on the polypeptides units help to stabilize the quaternary structure. Since a protein trimer is composed of multiple polypeptide subunits, it is considered an oligomer. A homotrimer would be formed by three identical molecules. A heterotrimer would be formed by three different macromolecules. Type II Collagen is an example of homotrimeric protein, while Type I collagen is an AAB-type heterotrimeric protein. An example of viral protein homotrimeric protein is mammarenavirus of Z matrix protein. Porins usually arrange themselves in membranes as trimers. __TOC__ Bacteriophage T4 tail fiber Multiple copies of a poly ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |