|
C-Myc
''Myc'' is a family of regulator genes and proto-oncogenes that code for transcription factors. The ''Myc'' family consists of three related human genes: ''c-myc'' ( MYC), ''l-myc'' ( MYCL), and ''n-myc'' ( MYCN). ''c-myc'' (also sometimes referred to as ''MYC'') was the first gene to be discovered in this family, due to homology with the viral gene ''v-myc''. In cancer, ''c-myc'' is often constitutively (persistently) expressed. This leads to the increased expression of many genes, some of which are involved in cell proliferation, contributing to the formation of cancer. A common human translocation involving ''c-myc'' is critical to the development of most cases of Burkitt lymphoma. Constitutive upregulation of ''Myc'' genes have also been observed in carcinoma of the cervix, colon, breast, lung and stomach. Myc is thus viewed as a promising target for anti-cancer drugs. Unfortunately, Myc possesses several features that render it undruggable such that any anti-cancer dr ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Internal Ribosome Entry Site
An internal ribosome entry site, abbreviated IRES, is an RNA element that allows for translation initiation in a cap-independent manner, as part of the greater process of protein synthesis. In eukaryotic translation, initiation typically occurs at the 5' end of mRNA molecules, since 5' cap recognition is required for the assembly of the initiation complex. The location for IRES elements is often in the 5'UTR, but can also occur elsewhere in mRNAs. History IRES sequences were first discovered in 1988 in the poliovirus (PV) and encephalomyocarditis virus (EMCV) RNA genomes in the labs of Nahum Sonenberg and Eckard Wimmer, respectively. They are described as distinct regions of RNA molecules that are able to recruit the eukaryotic ribosome to the mRNA. This process is also known as cap-independent translation. It has been shown that IRES elements have a distinct secondary or even tertiary structure, but similar structural features at the levels of either primary or secondary structure ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Histone Acetyltransferase
Histone acetyltransferases (HATs) are enzymes that acetylate conserved lysine amino acids on histone proteins by transferring an acetyl group from acetyl-CoA to form ε-''N''-acetyllysine. DNA is wrapped around histones, and, by transferring an acetyl group to the histones, genes can be turned on and off. In general, histone acetylation increases gene expression. In general, histone acetylation is linked to transcriptional activation and associated with euchromatin. Euchromatin, which is less densely compact, allows transcription factors to bind more easily to regulatory sites on DNA, causing transcriptional activation. When it was first discovered, it was thought that acetylation of lysine neutralizes the positive charge normally present, thus reducing affinity between histone and (negatively charged) DNA, which renders DNA more accessible to transcription factors. Research has emerged, since, to show that lysine acetylation and other posttranslational modifications of h ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MYCL
L-myc-1 proto-oncogene protein is a protein that in humans is encoded by the ''MYCL1'' gene. MYCL1 is a bHLH (basic helix-loop-helix) transcription factor implicated in lung cancer. Interactions MYCL1 has been shown to interact Advocates for Informed Choice, doing business as, dba interACT or interACT Advocates for Intersex Youth, is a 501(c)(3) nonprofit organization using innovative strategies to advocate for the legal and human rights of children with intersex trai ... with MAX. References Further reading * * * * * * * * * * * * * * * External links * Transcription factors {{gene-1-stub ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Burkitt's Lymphoma
Burkitt lymphoma is a cancer of the lymphatic system, particularly B lymphocytes found in the germinal center. It is named after Denis Parsons Burkitt, the Irish surgeon who first described the disease in 1958 while working in equatorial Africa. The overall cure rate for Burkitt lymphoma in developed countries is about 90%, and it is worse in low-income countries. Burkitt lymphoma is uncommon in adults, in whom it has a worse prognosis. Classification Burkitt lymphoma can be divided into three main clinical variants: the endemic, the sporadic, and the immunodeficiency-associated variants. By morphology (i.e., microscopic appearance), immunophenotype, and genetics, the variants of Burkitt lymphoma are alike. * The endemic variant (also called "African variant") most commonly occurs in children living in malaria-endemic regions of the world (e.g., equatorial Africa, Brazil, and Papua New Guinea). Epstein–Barr virus (EBV) infection is found in nearly all patients. Chronic mal ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Wnt Signalling Pathway
The Wnt signaling pathways are a group of signal transduction pathways which begin with proteins that pass signals into a cell through cell surface receptors. The name Wnt is a portmanteau created from the names Wingless and Int-1. Wnt signaling pathways use either nearby cell-cell communication (paracrine) or same-cell communication (autocrine). They are highly evolutionarily conserved in animals, which means they are similar across animal species from fruit flies to humans. Three Wnt signaling pathways have been characterized: the canonical Wnt pathway, the noncanonical planar cell polarity pathway, and the noncanonical Wnt/calcium pathway. All three pathways are activated by the binding of a Wnt-protein ligand to a Frizzled family receptor, which passes the biological signal to the Dishevelled protein inside the cell. The canonical Wnt pathway leads to regulation of gene transcription, and is thought to be negatively regulated in part by the SPATS1 gene. The noncanonical plan ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Mitogen
A mitogen is a small bioactive protein or peptide that induces a cell to begin cell division, or enhances the rate of division (mitosis). Mitogenesis is the induction (triggering) of mitosis, typically via a mitogen. The mechanism of action of a mitogen is that it triggers signal transduction pathways involving mitogen-activated protein kinase (MAPK), leading to mitosis. The cell cycle Mitogens act primarily by influencing a set of proteins which are involved in the restriction of progression through the cell cycle. The G1 checkpoint is controlled most directly by mitogens: further cell cycle progression does not need mitogens to continue. The point where mitogens are no longer needed to move the cell cycle forward is called the "restriction point" and depends on cyclins to be passed.Bohmer et al. "Cytoskeletal Integrity Is Required throughout the Mitogen Stimulation Phase of the Cell Cycle and Mediates the Anchorage-dependent Expression of Cyclin DI". January 1996, Molecular B ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Co-activator
A coactivator is a type of transcriptional coregulator that binds to an activator (a transcription factor) to increase the rate of transcription of a gene or set of genes. The activator contains a DNA binding domain that binds either to a DNA promoter site or a specific DNA regulatory sequence called an enhancer. Binding of the activator-coactivator complex increases the speed of transcription by recruiting general transcription machinery to the promoter, therefore increasing gene expression. The use of activators and coactivators allows for highly specific expression of certain genes depending on cell type and developmental stage. Some coactivators also have histone acetyltransferase (HAT) activity. HATs form large multiprotein complexes that weaken the association of histones to DNA by acetylating the N-terminal histone tail. This provides more space for the transcription machinery to bind to the promoter, therefore increasing gene expression. Activators are found in all ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
EP300
Histone acetyltransferase p300 also known as p300 HAT or E1A-associated protein p300 (where E1A = adenovirus early region 1A) also known as EP300 or p300 is an enzyme that, in humans, is encoded by the ''EP300'' gene. It functions as histone acetyltransferase that regulates transcription of genes via chromatin remodeling by allowing histone proteins to wrap DNA less tightly. This enzyme plays an essential role in regulating cell growth and division, prompting cells to mature and assume specialized functions (differentiate), and preventing the growth of cancerous tumors. The p300 protein appears to be critical for normal development before and after birth. The EP300 gene is located on the long (q) arm of the human chromosome 22 at position 13.2. This gene encodes the adenovirus E1A-associated cellular p300 transcriptional co-activator protein. EP300 is closely related to another gene, CREB binding protein, which is found on human chromosome 16. Function p300 HAT functions ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
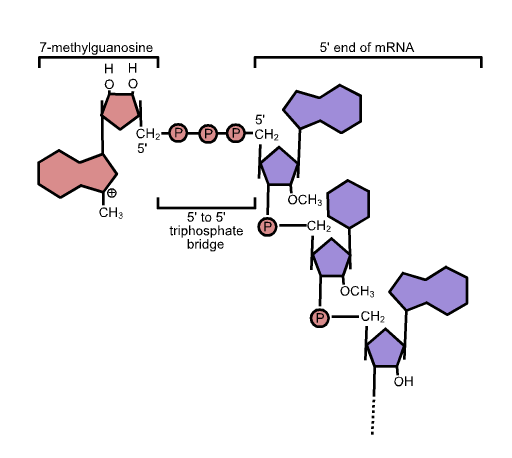
5' Cap
In molecular biology, the five-prime cap (5′ cap) is a specially altered nucleotide on the 5′ end of some primary transcripts such as precursor messenger RNA. This process, known as mRNA capping, is highly regulated and vital in the creation of stable and mature messenger RNA able to undergo translation during protein synthesis. Mitochondrial mRNA and chloroplastic mRNA are not capped. Structure In eukaryotes, the 5′ cap (cap-0), found on the 5′ end of an mRNA molecule, consists of a guanine nucleotide connected to mRNA via an unusual 5′ to 5′ triphosphate linkage. This guanosine is methylated on the 7 position directly after capping ''in vivo'' by a methyltransferase. It is referred to as a 7-methylguanylate cap, abbreviated m7G. In multicellular eukaryotes and some viruses, further modifications exist, including the methylation of the 2′ hydroxy-groups of the first 2 ribose sugars of the 5′ end of the mRNA. cap-1 has a methylated 2′-hydroxy grou ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MRNA
In molecular biology, messenger ribonucleic acid (mRNA) is a single-stranded molecule of RNA that corresponds to the genetic sequence of a gene, and is read by a ribosome in the process of synthesizing a protein. mRNA is created during the process of transcription, where an enzyme ( RNA polymerase) converts the gene into primary transcript mRNA (also known as pre-mRNA). This pre-mRNA usually still contains introns, regions that will not go on to code for the final amino acid sequence. These are removed in the process of RNA splicing, leaving only exons, regions that will encode the protein. This exon sequence constitutes mature mRNA. Mature mRNA is then read by the ribosome, and, utilising amino acids carried by transfer RNA (tRNA), the ribosome creates the protein. This process is known as translation. All of these processes form part of the central dogma of molecular biology, which describes the flow of genetic information in a biological system. As in DNA, ge ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
MAX (gene)
''MAX'' (also known as myc-associated factor X) is a gene that in humans encodes the MAX transcription factor. Function The protein product of ''MAX'' contains the basic helix-loop-helix and leucine zipper motifs. It is therefore included in the bHLHZ family of transcription factors. It is able to form homodimers with other MAX proteins and heterodimers with other transcription factors, including Mad, Mxl1 and Myc. The homodimers and heterodimers compete for a common DNA target site (the E-box) in a gene promoter zone. Rearrangement of dimers (e.g., Mad:Max, Max:Myc) provides a system of transcriptional regulation with greater diversity of gene targets. Max must dimerise in order to be biologically active. Transcriptionally active hetero- and homodimers involving Max can promote cell proliferation as well as apoptosis. Interactions The protein product of Max has been shown to interact with: * Myc, * MNT, * MSH2, * MXD1, * MXI1, * MYCL1, * N-Myc, * SPAG9, * TEAD1, an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Leucine Zipper
A leucine zipper (or leucine scissors) is a common three-dimensional structural motif in proteins. They were first described by Landschulz and collaborators in 1988 when they found that an enhancer binding protein had a very characteristic 30-amino acid segment and the display of these amino acid sequences on an idealized alpha helix revealed a periodic repetition of leucine residues at every seventh position over a distance covering eight helical turns. The polypeptide segments containing these periodic arrays of leucine residues were proposed to exist in an alpha-helical conformation and the leucine side chains from one alpha helix interdigitate with those from the alpha helix of a second polypeptide, facilitating dimerization. Leucine zippers are a dimerization motif of the bZIP (Basic-region leucine zipper) class of eukaryotic transcription factors. The bZIP domain is 60 to 80 amino acids in length with a highly conserved DNA binding basic region and a more diversified leuci ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |