|
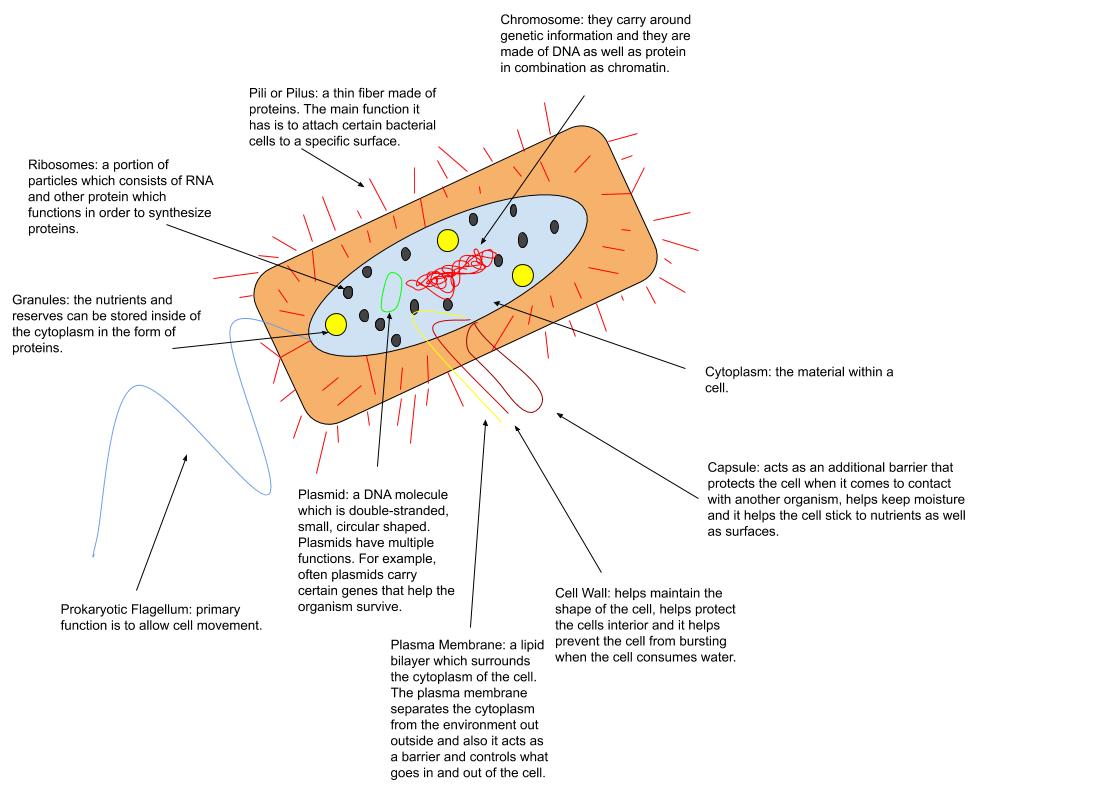
Deinococcus Deserti
''Deinococcus deserti'' is a Gram-negative bacteria, Gram-negative, rod-shaped Bacteria, bacterium that belongs to the Deinococcaceae, a group of extremely radiotolerant bacteria. ''D. deserti'' and other Deinococcaceae exhibit an extraordinary ability to withstand ionizing radiation. Description ''Deinococcus deserti'' has in common with other ''Deinococci'' a highly condensed nucleoid, a high cellular Mn/Fe ratio, and several of the ''Deinococcus'' specific radiation tolerance-associated genes, for example, ddrA to ddrD, pprA, and irrE. The genome of ''D. deserti'' VCD115 is composed of four replicons: a main chromosome (2.82 Mb) and three plasmids, P1 (325 kb), P2 (314 kb) and P3 (396 kb). History Two gamma- and UV-radiation-tolerant Strain (biology), strains were isolated from a mixture of sand samples collected in the Sahara Desert in Morocco and Tunisia, after exposure of the sand to 15 kGy gamma radiation. The strains did not grow on rich medium such as trypticase s ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
LPSN
List of Prokaryotic names with Standing in Nomenclature (LPSN) is an online database that maintains information on the naming and taxonomy of prokaryotes, following the taxonomy requirements and rulings of the International Code of Nomenclature of Prokaryotes. The database was curated from 1997 to June 2013 by Jean P. Euzéby. From July 2013 to January 2020, LPSN was curated by Aidan C. Parte. In February 2020, a new version of LPSN was published as a service of the Leibniz Institute DSMZ, thereby also integrating the Prokaryotic Nomenclature Up-to-date service. References External links List of Prokaryotic names with Standing in Nomenclature [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome
In the fields of molecular biology and genetics, a genome is all the genetic information of an organism. It consists of nucleotide sequences of DNA (or RNA in RNA viruses). The nuclear genome includes protein-coding genes and non-coding genes, other functional regions of the genome such as regulatory sequences (see non-coding DNA), and often a substantial fraction of 'junk' DNA with no evident function. Almost all eukaryotes have mitochondria and a small mitochondrial genome. Algae and plants also contain chloroplasts with a chloroplast genome. The study of the genome is called genomics. The genomes of many organisms have been sequenced and various regions have been annotated. The International Human Genome Project reported the sequence of the genome for ''Homo sapiens'' in 200The Human Genome Project although the initial "finished" sequence was missing 8% of the genome consisting mostly of repetitive sequences. With advancements in technology that could handle seq ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Hidden Markov Model
A hidden Markov model (HMM) is a statistical Markov model in which the system being modeled is assumed to be a Markov process — call it X — with unobservable ("''hidden''") states. As part of the definition, HMM requires that there be an observable process Y whose outcomes are "influenced" by the outcomes of X in a known way. Since X cannot be observed directly, the goal is to learn about X by observing Y. HMM has an additional requirement that the outcome of Y at time t=t_0 must be "influenced" exclusively by the outcome of X at t=t_0 and that the outcomes of X and Y at t handwriting recognition, handwriting, gesture recognition, part-of-speech tagging, musical score following, partial discharges and bioinformatics. Definition Let X_n and Y_n be discrete-time stochastic processes and n\geq 1. The pair (X_n,Y_n) is a ''hidden Markov model'' if * X_n is a Markov process whose behavior is not directly observable ("hidden"); * \operatorname\bigl(Y_n \ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Open Reading Frame
In molecular biology, open reading frames (ORFs) are defined as spans of DNA sequence between the start and stop codons. Usually, this is considered within a studied region of a Prokaryote, prokaryotic DNA sequence, where only one of the #Six-frame translation, six possible reading frames will be "open" (the "reading", however, refers to the RNA produced by Transcription (biology), transcription of the DNA and its subsequent interaction with the ribosome in Translation (biology), translation). Such an ORF may contain a start codon (usually AUG in terms of RNA) and by definition cannot extend beyond a stop codon (usually UAA, UAG or UGA in RNA). That start codon (not necessarily the first) indicates where translation may start. The transcription terminator, transcription termination site is located after the ORF, beyond the Translation (biology), translation stop codon. If transcription were to cease before the stop codon, an incomplete protein would be made during translation. In ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Chromatography
In chemical analysis, chromatography is a laboratory technique for the separation of a mixture into its components. The mixture is dissolved in a fluid solvent (gas or liquid) called the ''mobile phase'', which carries it through a system (a column, a capillary tube, a plate, or a sheet) on which a material called the ''stationary phase'' is fixed. Because the different constituents of the mixture tend to have different affinities for the stationary phase and are retained for different lengths of time depending on their interactions with its surface sites, the constituents travel at different apparent velocities in the mobile fluid, causing them to separate. The separation is based on the differential partitioning between the mobile and the stationary phases. Subtle differences in a compound's partition coefficient result in differential retention on the stationary phase and thus affect the separation. Chromatography may be preparative or analytical. The purpose of preparat ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Proteome
The proteome is the entire set of proteins that is, or can be, expressed by a genome, cell, tissue, or organism at a certain time. It is the set of expressed proteins in a given type of cell or organism, at a given time, under defined conditions. Proteomics is the study of the proteome. Types of proteomes While proteome generally refers to the proteome of an organism, multicellular organisms may have very different proteomes in different cells, hence it is important to distinguish proteomes in cells and organisms. A cellular proteome is the collection of proteins found in a particular cell type under a particular set of environmental conditions such as exposure to hormone stimulation. It can also be useful to consider an organism's complete proteome, which can be conceptualized as the complete set of proteins from all of the various cellular proteomes. This is very roughly the protein equivalent of the genome. The term ''proteome'' has also been used to refer to the collecti ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Genome Annotation
DNA annotation or genome annotation is the process of identifying the locations of genes and all of the coding regions in a genome and determining what those genes do. An annotation (irrespective of the context) is a note added by way of explanation or commentary. Once a genome is sequenced, it needs to be annotated to make sense of it. Genes in a eukaryotic genome can be annotated using various annotation tools such as FINDER. A modern annotation pipeline can support a user-friendly web interface and software containerization such as MOSGA. For DNA annotation, a previously unknown sequence representation of genetic material is enriched with information relating genomic position to intron-exon boundaries, regulatory sequences, repeats, gene names and protein products. This annotation is stored in genomic databases such as Mouse Genome Informatics, FlyBase, and WormBase. Educational materials on some aspects of biological annotation from the 2006 Gene Ontology annotation camp an ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Regulatory Sequence
A regulatory sequence is a segment of a nucleic acid molecule which is capable of increasing or decreasing the expression of specific genes within an organism. Regulation of gene expression is an essential feature of all living organisms and viruses. Description In DNA, regulation of gene expression normally happens at the level of RNA biosynthesis ( transcription). It is accomplished through the sequence-specific binding of proteins (transcription factors) that activate or inhibit transcription. Transcription factors may act as activators, repressors, or both. Repressors often act by preventing RNA polymerase from forming a productive complex with the transcriptional initiation region ( promoter), while activators facilitate formation of a productive complex. Furthermore, DNA motifs have been shown to be predictive of epigenomic modifications, suggesting that transcription factors play a role in regulating the epigenome. In RNA, regulation may occur at the level of prote ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Upregulation
In the biological context of organisms' production of gene products, downregulation is the process by which a cell decreases the quantity of a cellular component, such as RNA or protein, in response to an external stimulus. The complementary process that involves increases of such components is called upregulation. An example of downregulation is the cellular decrease in the expression of a specific receptor in response to its increased activation by a molecule, such as a hormone or neurotransmitter, which reduces the cell's sensitivity to the molecule. This is an example of a locally acting (negative feedback) mechanism. An example of upregulation is the response of liver cells exposed to such xenobiotic molecules as dioxin. In this situation, the cells increase their production of cytochrome P450 enzymes, which in turn increases degradation of these dioxin molecules. Downregulation or upregulation of an RNA or protein may also arise by an epigenetic alteration. Such an e ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Replication
In molecular biology, DNA replication is the biological process of producing two identical replicas of DNA from one original DNA molecule. DNA replication occurs in all living organisms acting as the most essential part for biological inheritance. This is essential for cell division during growth and repair of damaged tissues, while it also ensures that each of the new cells receives its own copy of the DNA. The cell possesses the distinctive property of division, which makes replication of DNA essential. DNA is made up of a double helix of two complementary strands. The double helix describes the appearance of a double-stranded DNA which is thus composed of two linear strands that run opposite to each other and twist together to form. During replication, these strands are separated. Each strand of the original DNA molecule then serves as a template for the production of its counterpart, a process referred to as semiconservative replication. As a result of semi-conservativ ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
Transcriptome
The transcriptome is the set of all RNA transcripts, including coding and non-coding, in an individual or a population of cells. The term can also sometimes be used to refer to all RNAs, or just mRNA, depending on the particular experiment. The term ''transcriptome'' is a portmanteau of the words ''transcript'' and ''genome''; it is associated with the process of transcript production during the biological process of transcription. The early stages of transcriptome annotations began with cDNA libraries published in the 1980s. Subsequently, the advent of high-throughput technology led to faster and more efficient ways of obtaining data about the transcriptome. Two biological techniques are used to study the transcriptome, namely DNA microarray, a hybridization-based technique and RNA-seq, a sequence-based approach. RNA-seq is the preferred method and has been the dominant transcriptomics technique since the 2010s. Single-cell transcriptomics allows tracking of transcript chang ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |
DNA Repair
DNA repair is a collection of processes by which a cell identifies and corrects damage to the DNA molecules that encode its genome. In human cells, both normal metabolic activities and environmental factors such as radiation can cause DNA damage, resulting in tens of thousands of individual molecular lesions per cell per day. Many of these lesions cause structural damage to the DNA molecule and can alter or eliminate the cell's ability to transcribe the gene that the affected DNA encodes. Other lesions induce potentially harmful mutations in the cell's genome, which affect the survival of its daughter cells after it undergoes mitosis. As a consequence, the DNA repair process is constantly active as it responds to damage in the DNA structure. When normal repair processes fail, and when cellular apoptosis does not occur, irreparable DNA damage may occur, including double-strand breaks and DNA crosslinkages (interstrand crosslinks or ICLs). This can eventually lead to malig ... [...More Info...] [...Related Items...] OR: [Wikipedia] [Google] [Baidu] |